

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0334a.1
(1167 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
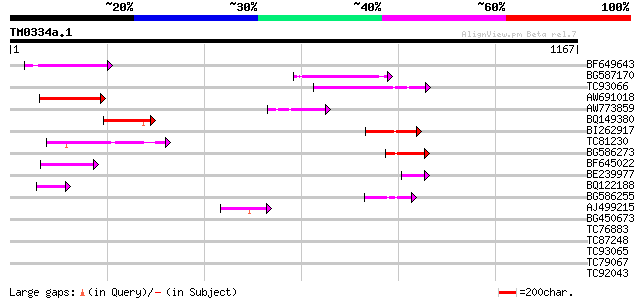
Score E
Sequences producing significant alignments: (bits) Value
BF649643 similar to PIR|H86486|H86 protein Ty1/copia-element pol... 146 4e-35
BG587170 similar to PIR|F86470|F8 probable retroelement polyprot... 138 1e-32
TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative ret... 135 1e-31
AW691018 weakly similar to GP|2462935|em open reading frame 1 {B... 120 2e-27
AW773859 99 1e-20
BQ149380 92 1e-18
BI262917 weakly similar to GP|19920130|g Putative retroelement {... 91 3e-18
TC81230 80 6e-15
BG586273 weakly similar to PIR|F86470|F8 probable retroelement p... 73 7e-13
BF645022 weakly similar to GP|8777581|dbj retroelement pol polyp... 71 3e-12
BE239977 weakly similar to GP|23237899|db polyprotein-like {Oryz... 58 2e-08
BQ122188 weakly similar to GP|9758772|dbj| contains similarity t... 57 3e-08
BG586255 similar to GP|7682800|gb Hypothetical protein T15F17.l ... 53 6e-07
AJ499215 weakly similar to GP|6642775|gb| gag-pol polyprotein {V... 51 2e-06
BG450673 homologue to GP|12055499|emb NADH dehydrogenase subunit... 41 0.002
TC76883 homologue to SP|Q9M4D8|DCAM_VICFA S-adenosylmethionine d... 34 0.38
TC87248 similar to GP|7594580|emb|CAB88075.1 hypothetical protei... 32 1.1
TC93065 32 1.1
TC79067 similar to PIR|T45816|T45816 hypothetical protein F28O9.... 32 1.4
TC92043 similar to GP|21554299|gb|AAM63374.1 apospory-associated... 31 2.4
>BF649643 similar to PIR|H86486|H86 protein Ty1/copia-element polyprotein
[imported] - Arabidopsis thaliana, partial (2%)
Length = 608
Score = 146 bits (369), Expect = 4e-35
Identities = 71/182 (39%), Positives = 104/182 (57%), Gaps = 2/182 (1%)
Frame = +1
Query: 31 QASNNNNNGVQQAHGVQQPHVVLEPAQNPISPYYIHSGENPSATVVSLPLNGRNYNAWA* 90
+ SNNN+N + + +PYYIH ++P +V LNG NY++W+
Sbjct: 55 EKSNNNSNKSNNKNNSDM---------DSSNPYYIHPSDHPGHLLVPTKLNGTNYSSWSR 207
Query: 91 SMKRVLVAKNKFKFINGEI--PIAVPGDANYEAWDRCNNLIHSWILNSVTSSIANSIVFV 148
SM L AKNK FING I P V A Y W+RCN++I SW+ +SV +A ++
Sbjct: 208 SMVHALTAKNKVGFINGSIKTPSEVDQPAEYALWNRCNSMILSWLTHSVEPDLAKGVIHA 387
Query: 149 ENACDAWRDLRDRFSQGVLVRIAELQNEIGNLKQNTLSVNDYYTEIKTLWEELEQYRPIP 208
+ AC W D +D+FSQ + I ++Q + +L Q T+S + Y+T+IK LW+ELE YR +P
Sbjct: 388 KTACQVWEDFKDQFSQKNIPAIYQIQKSLASLSQGTVSASTYFTKIKGLWDELETYRTLP 567
Query: 209 QC 210
C
Sbjct: 568 TC 573
>BG587170 similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 718
Score = 138 bits (347), Expect = 1e-32
Identities = 81/208 (38%), Positives = 124/208 (58%), Gaps = 3/208 (1%)
Frame = -3
Query: 584 ALWHFRLGHLSHDRILALNALYPSIDVSKHFVCDICHLAKLKRKMFPDSLHNAKCNFDLL 643
ALWH RLGH H R ALN + P + V ++ C+ C L K + +FP + + FDL+
Sbjct: 587 ALWHARLGH-PHGR--ALNLMLPGV-VFENKNCEACILGKHCKNVFPRTSTVYENCFDLI 420
Query: 644 HMDIWGPISVSSVHSHRYFLTVLDDHSRFVWVILLKRKGEVQQ*IKNFVALVKTQFSHHV 703
+ D+W S+S +H+YF+T +D+ S++ W+ L+ K V KNF A V + +
Sbjct: 419 YTDLWTAPSLSR-DNHKYFVTFIDEKSKYTWLTLIPSKDRVIDAFKNFQAYVTNHYHAKI 243
Query: 704 KVIRSDNGPEFMLHDFYS---AHGILHQRSCVNTPQQNGRVERKHQHILNIARALLFQSH 760
K++RSDNG E+ + F S HGILHQ SC TPQQNG +RK++H++ +AR+L+FQ++
Sbjct: 242 KILRSDNGGEYTSYAFKSHLDHHGILHQTSCPYTPQQNGVAKRKNKHLMEVARSLMFQAN 63
Query: 761 LPKKMWGYAILHAVFLMSRIQSKVLENK 788
+ A +L++ I +KVL K
Sbjct: 62 ---------VSTACYLINWIPTKVLRIK 6
>TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative retroelement
{Oryza sativa} [Oryza sativa (japonica cultivar-group)],
partial (10%)
Length = 823
Score = 135 bits (339), Expect = 1e-31
Identities = 87/249 (34%), Positives = 129/249 (50%), Gaps = 7/249 (2%)
Frame = +1
Query: 625 KRKMFPDSLHNAKCNFDLLHMDIWGPISVSSVHSHRYFLTVLDDHSRFVWVILLKRKGEV 684
K+ F + H K D +H D+WGP V+S RY +T++DD R VWV L+ K E
Sbjct: 52 KKVSFSTATHRTKGILDYIHSDLWGPSKVTSYGGRRYMMTIIDDFPRKVWVYFLRYKNET 231
Query: 685 QQ*IKNFVALVKTQFSHHVKVIRSDNGPEFM---LHDFYSAHGILHQRSCVNTPQQNGRV 741
K + LV+TQ +VK + +DN EF ++F + HGI ++ PQQNG
Sbjct: 232 FPTFKKWRILVETQTGKNVKKLITDN*LEFCSSDFNEFCTNHGIARHKTIPRNPQQNGVA 411
Query: 742 ERKHQHILNIARALLFQSHL--PKKMWGYAILHAVFLMSRIQSKVLENKSPYEILFGGKK 799
ER + +L AR +L + L + +W A A L++R L+ K P E ++ G
Sbjct: 412 ERMIRTLLERARCMLSNAGL*N*RDLWVEAASTACHLVNRSPHSALDFKVP-EDIWSGNL 588
Query: 800 IDLQELRVFGSLCFASTSSSSRSKLDSRARKCVFLGFKQGVKGFVL--LDLNSHEIFVSR 857
+D LR+FG +A + KL RA +C+FL + KG+ L D S ++ +SR
Sbjct: 589 VDYSNLRIFGCPAYALVND---GKLAPRAGECIFLSYASESKGYRLWCSDPKSQKLILSR 759
Query: 858 DVTFHEQIL 866
DVTF+E L
Sbjct: 760 DVTFNEDAL 786
>AW691018 weakly similar to GP|2462935|em open reading frame 1 {Brassica
oleracea}, partial (12%)
Length = 639
Score = 120 bits (302), Expect = 2e-27
Identities = 56/136 (41%), Positives = 82/136 (60%)
Frame = +2
Query: 62 PYYIHSGENPSATVVSLPLNGRNYNAWA*SMKRVLVAKNKFKFINGEIPIAVPGDANYEA 121
P +H + P ++V+ PLNG NY W+ SM L AKNK I+G + D
Sbjct: 155 PLLLHHSDTPGISLVNQPLNGGNYGEWSRSMLLSLSAKNKLGLIDGTVKAPSADDPKLPL 334
Query: 122 WDRCNNLIHSWILNSVTSSIANSIVFVENACDAWRDLRDRFSQGVLVRIAELQNEIGNLK 181
W RCN+L+ +WIL+S+ IA S++F + A W DL DRFSQG RI +++ EI +
Sbjct: 335 WKRCNDLVLTWILHSIEPDIARSVIFSDTAAAVWSDLHDRFSQGDESRIYQIRQEISECR 514
Query: 182 QNTLSVNDYYTEIKTL 197
Q +L ++DYYT++K+L
Sbjct: 515 QGSLLISDYYTKLKSL 562
>AW773859
Length = 538
Score = 99.0 bits (245), Expect = 1e-20
Identities = 52/129 (40%), Positives = 75/129 (57%)
Frame = -3
Query: 532 VQEKHTKKRIGLGSLKEDDLYHLVVTSPSSTCFAIPNVVERISDSDQLIPPGALWHFRLG 591
+QEK + K IG G L E LY L S A V + + +P ALWHFRLG
Sbjct: 380 IQEKRSLKMIGSGELFEG-LYFLTTQDTPSATIASSQVQPQPQPT--FLPQEALWHFRLG 210
Query: 592 HLSHDRILALNALYPSIDVSKHFVCDICHLAKLKRKMFPDSLHNAKCNFDLLHMDIWGPI 651
HLS+ ++L+L++ +P I + ++ VCDICH ++ K+ F S + A ++L H DIWGP
Sbjct: 209 HLSNRKLLSLHSNFPFITIDQNSVCDICHYSRHKKLPFQLSTNRASKCYELFHFDIWGPF 30
Query: 652 SVSSVHSHR 660
S S+H+ R
Sbjct: 29 STQSIHNQR 3
>BQ149380
Length = 419
Score = 92.0 bits (227), Expect = 1e-18
Identities = 43/114 (37%), Positives = 70/114 (60%), Gaps = 7/114 (6%)
Frame = +3
Query: 193 EIKTLWEELEQYRPIPQCRCVVPCRCEAIEHAKMFREQDNAIRFLLGLNETFSVVNSQIL 252
+++ LW+EL+QYRP PQC C + C C A+ + K +D I+FL+GLN+ + V S++L
Sbjct: 69 DLRGLWDELDQYRPKPQCTCPIQCTCFAMRNNKALTAEDRIIKFLMGLNDDYQGVTSKVL 248
Query: 253 MSNPLPPIAKVVSLAMQHERQ-------SETGENEESKSLVKMAEGKKTYGKGK 299
+ PLP I V S+ MQ ER+ S T ++ + LV +G++ +G+G+
Sbjct: 249 LMYPLPQINMVFSMVMQQERKMQ*NVVSSPTPIDDTTCGLVNAMDGQRQFGRGR 410
>BI262917 weakly similar to GP|19920130|g Putative retroelement {Oryza
sativa} [Oryza sativa (japonica cultivar-group)],
partial (8%)
Length = 426
Score = 90.5 bits (223), Expect = 3e-18
Identities = 45/115 (39%), Positives = 71/115 (61%)
Frame = +1
Query: 733 NTPQQNGRVERKHQHILNIARALLFQSHLPKKMWGYAILHAVFLMSRIQSKVLENKSPYE 792
+TPQQNG ER ++ +L RA+L + + K W A+ A ++++R S V++ K+P E
Sbjct: 82 HTPQQNGVAERMNRTLLERTRAMLKTAGMAKSFWAEAVKTACYVINRSPSTVIDLKTPME 261
Query: 793 ILFGGKKIDLQELRVFGSLCFASTSSSSRSKLDSRARKCVFLGFKQGVKGFVLLD 847
++ GK +D L VFG + +S R+KLD ++RKC+FLG+ VKG+ L D
Sbjct: 262 -MWKGKPVDYSSLHVFGCPVYVMYNSQERTKLDPKSRKCIFLGYADNVKGYXLWD 423
>TC81230
Length = 958
Score = 79.7 bits (195), Expect = 6e-15
Identities = 62/264 (23%), Positives = 108/264 (40%), Gaps = 9/264 (3%)
Frame = +1
Query: 76 VSLPLNGRNYNAWA*SMKRVLVAKNKFKFINGEIPIAVPG-----DA---NYEAWDRCNN 127
+S+ LNG NYN WA SM L + ++++ G+ G DA E WD N+
Sbjct: 226 ISIILNGSNYNHWAESMCGFLKGRRLWRYVTGDKKCPTKGKDDTADAFADKLEEWDSKNH 405
Query: 128 LIHSWILNSVTSSIANSIVFVENACDAWRDLRDRFSQGVLVRIAELQNEIGNLKQNT-LS 186
I +W N+ SI ENA + W L+ R++ L +L ++ NLKQ +
Sbjct: 406 QIITWFRNTSIPSIHMQFGRFENAKEVWDHLKQRYTISDLSHQYQLLKDLSNLKQQSGQP 585
Query: 187 VNDYYTEIKTLWEELEQYRPIPQCRCVVPCRCEAIEHAKMFREQDNAIRFLLGLNETFSV 246
V ++ +++ +W +L P + ++ + R + I+FL+ L + +
Sbjct: 586 VYEFLAQMEVIWNQLTSCEPSLK-------DATDMKTYETHRNRVRLIQFLMALTDEYEP 744
Query: 247 VNSQILMSNPLPPIAKVVSLAMQHERQSETGENEESKSLVKMAEGKKTYGKGKAPGSSSS 306
V + L NPLP + + E + + P + +
Sbjct: 745 VRASSLHQNPLPTLENALPCLKSEETRLQL----------------------VPPKADLA 858
Query: 307 GSGYKSTGKYCTHCKKPGHTVDVC 330
+ + K C HC+K GH+ C
Sbjct: 859 FAVTNNATKPCRHCQKSGHSFSDC 930
>BG586273 weakly similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 705
Score = 72.8 bits (177), Expect = 7e-13
Identities = 35/92 (38%), Positives = 60/92 (65%)
Frame = -2
Query: 773 AVFLMSRIQSKVLENKSPYEILFGGKKIDLQELRVFGSLCFASTSSSSRSKLDSRARKCV 832
A +L++RI ++VL++++P+E+L +K L +RVFG LC+ R+KL++R+RK +
Sbjct: 704 ACYLINRIPTRVLKDQAPFEVL-NQRKPSLTYMRVFGCLCYVLVPGELRNKLEARSRKAM 528
Query: 833 FLGFKQGVKGFVLLDLNSHEIFVSRDVTFHEQ 864
F+G+ KG+ D + + VSRDV F E+
Sbjct: 527 FIGYSTTQKGYKCYDPEARRVLVSRDVKFIEE 432
>BF645022 weakly similar to GP|8777581|dbj retroelement pol polyprotein-like
{Arabidopsis thaliana}, partial (8%)
Length = 685
Score = 70.9 bits (172), Expect = 3e-12
Identities = 32/120 (26%), Positives = 65/120 (53%)
Frame = -2
Query: 63 YYIHSGENPSATVVSLPLNGRNYNAWA*SMKRVLVAKNKFKFINGEIPIAVPGDANYEAW 122
+++ S +NP + + L G NY+ W+ +++ AK K+ F+ G+IP + E W
Sbjct: 375 FFLGSNDNPGNVITPIQLRGLNYDEWSRAIRTSFQAKRKYGFVEGKIPKPTTPE-KLEDW 199
Query: 123 DRCNNLIHSWILNSVTSSIANSIVFVENACDAWRDLRDRFSQGVLVRIAELQNEIGNLKQ 182
+++ +W+LN++ S+ +++ + ++A W L+ RF RI +L+ +G KQ
Sbjct: 198 KAVQSMLIAWLLNTIEPSLRSTLSYYDDAESLWTHLKQRFCVVNGARICQLKASLGECKQ 19
>BE239977 weakly similar to GP|23237899|db polyprotein-like {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 514
Score = 57.8 bits (138), Expect = 2e-08
Identities = 29/59 (49%), Positives = 34/59 (57%)
Frame = -2
Query: 806 RVFGSLCFASTSSSSRSKLDSRARKCVFLGFKQGVKGFVLLDLNSHEIFVSRDVTFHEQ 864
R+FG F S RSK D RA KCVF+ + KG+ S + FVSRDVTFHEQ
Sbjct: 456 RIFGCTSFVHIHSDGRSKFDHRALKCVFIRYSSTQKGYRCYHPPSRKYFVSRDVTFHEQ 280
>BQ122188 weakly similar to GP|9758772|dbj| contains similarity to
retroelement pol polyprotein~gene_id:MNJ7.3 {Arabidopsis
thaliana}, partial (7%)
Length = 708
Score = 57.4 bits (137), Expect = 3e-08
Identities = 26/70 (37%), Positives = 40/70 (57%), Gaps = 1/70 (1%)
Frame = -3
Query: 56 AQNPISPYYIHSGENPSATVVSLPLNG-RNYNAWA*SMKRVLVAKNKFKFINGEIPIAVP 114
A N +P+++H ENP+ +V+ PL G +N+ +W SM+ L +KNK F++G
Sbjct: 211 ATNASNPFFLHPSENPAVELVTPPLEGNKNFQSWIRSMRLALTSKNKIAFVDGTFLTPAK 32
Query: 115 GDANYEAWDR 124
DA Y W R
Sbjct: 31 SDALYNQWIR 2
>BG586255 similar to GP|7682800|gb Hypothetical protein T15F17.l {Arabidopsis
thaliana}, partial (12%)
Length = 436
Score = 53.1 bits (126), Expect = 6e-07
Identities = 34/107 (31%), Positives = 54/107 (49%)
Frame = -1
Query: 730 SCVNTPQQNGRVERKHQHILNIARALLFQSHLPKKMWGYAILHAVFLMSRIQSKVLENKS 789
S V+ QNG E + I I R LL +S P WG+A+LHA L+ RI+ S
Sbjct: 397 SVVHVHTQNGLAESFIKRIQLITRPLLMRSKQPVSAWGHAVLHAAELI-RIRPSSEHKYS 221
Query: 790 PYEILFGGKKIDLQELRVFGSLCFASTSSSSRSKLDSRARKCVFLGF 836
P ++L G D+ ++ FG + + R+K+ + R +++GF
Sbjct: 220 PSQLL-SGHVPDVSHIKTFGYPVYVPIAPPHRTKMGPQRRMGIYVGF 83
>AJ499215 weakly similar to GP|6642775|gb| gag-pol polyprotein {Vitis
vinifera}, partial (18%)
Length = 567
Score = 51.2 bits (121), Expect = 2e-06
Identities = 32/111 (28%), Positives = 62/111 (55%), Gaps = 5/111 (4%)
Frame = +3
Query: 434 SYSSQYDSRDWIIDSGASDHICFNKLCFDTLNRIKPVSVRLPNGISIQTCYAGVVSIT-- 491
+++++ S+ W+IDSG + H+ ++ F LN+ VR+ NG I+ G V +
Sbjct: 78 TFATKQPSKYWLIDSGCTHHMTHDRDLFKELNKSTISKVRMLNGAHIEVEGIGTVLVKSH 257
Query: 492 ---KQILLTHVLYVPEFTYNLISVHKLAKRNRCEVVFGEFTCFVQEKHTKK 539
KQI ++VLY P+ +L+SV +L + +V+F C +++++ K+
Sbjct: 258 SGYKQI--SNVLYAPKLNQSLLSVPQLLTKG-YKVLFEHEKCVIKDQNNKE 401
>BG450673 homologue to GP|12055499|emb NADH dehydrogenase subunit 2 {Sphagnum
fallax}, partial (2%)
Length = 678
Score = 41.2 bits (95), Expect = 0.002
Identities = 17/62 (27%), Positives = 31/62 (49%)
Frame = +3
Query: 99 KNKFKFINGEIPIAVPGDANYEAWDRCNNLIHSWILNSVTSSIANSIVFVENACDAWRDL 158
K+K +ING++P D + W N ++ WI+NS+ SS+ ++ + W +
Sbjct: 231 KDKLGYINGDLPPPPQTDPTFRRWGTDNAIVKGWIINSMDSSLISNFIRFSTTKLVWDAI 410
Query: 159 RD 160
D
Sbjct: 411 AD 416
>TC76883 homologue to SP|Q9M4D8|DCAM_VICFA S-adenosylmethionine
decarboxylase proenzyme (EC 4.1.1.50) (AdoMetDC)
(SamDC), complete
Length = 1819
Score = 33.9 bits (76), Expect = 0.38
Identities = 16/41 (39%), Positives = 22/41 (53%)
Frame = -1
Query: 252 LMSNPLPPIAKVVSLAMQHERQSETGENEESKSLVKMAEGK 292
L SNP PIA+ +Q ER +E NE+ ++ K GK
Sbjct: 604 LFSNPSKPIAETAMATVQTERHTEETRNEKKSTIAKRGFGK 482
>TC87248 similar to GP|7594580|emb|CAB88075.1 hypothetical protein
{Arabidopsis thaliana}, partial (82%)
Length = 1271
Score = 32.3 bits (72), Expect = 1.1
Identities = 16/48 (33%), Positives = 24/48 (49%), Gaps = 1/48 (2%)
Frame = -2
Query: 183 NTLSVNDYYTEIKTLWEEL-EQYRPIPQCRCVVPCRCEAIEHAKMFRE 229
N LS++ + LW L E++ PIP+ PCR +EH + E
Sbjct: 787 NNLSISPSCPQCSQLWPLLHEKHPPIPEAASTSPCRSAVLEHRWLIGE 644
>TC93065
Length = 783
Score = 32.3 bits (72), Expect = 1.1
Identities = 50/224 (22%), Positives = 85/224 (37%), Gaps = 10/224 (4%)
Frame = +2
Query: 171 AELQNEIGNLK-QNTLSVNDY-------YTEIKTLWEELEQYRPIPQCRCVVPCRCEAIE 222
A+L+ E LK + T +V ++ T+I+ L E+L R + + +P
Sbjct: 209 AKLRREFEALKMKETETVREFSDRISKVVTQIRLLGEDLSDQRVVEKILVCLP------- 367
Query: 223 HAKMFREQDNAIRFLLGLNETFSVVNSQILMSNPLPPIAKVVSLAMQHERQSETGENEES 282
+MF + +++ N+ FS + L+ N L + SL M+ + N +
Sbjct: 368 --EMFEAKISSLEE----NKNFSEITVAELV-NALQASEQRRSLRMEENVEGAFLANNKG 526
Query: 283 KSLVKMAEGKKTYGKGKAPGSSSSGSGYKSTGKYCTHCKKPGHTVDVCYRLHGYPTTSKS 342
K+ + K++GK K P C HCKK H C+ G
Sbjct: 527 KN-----QSFKSFGKKKFPP--------------CPHCKKDTHLDKFCWYRPGVKCR--- 640
Query: 343 KPAFNNVSHINNIT*GYDSEDDQEES--SKSQRGNGDLFTADQY 384
A N + H+ + ++ +QE Q LF A Y
Sbjct: 641 --ACNQLGHVEKVCKNKTNQQEQEARVVEHHQEDEEPLFKASCY 766
>TC79067 similar to PIR|T45816|T45816 hypothetical protein F28O9.230 -
Arabidopsis thaliana, partial (75%)
Length = 1979
Score = 32.0 bits (71), Expect = 1.4
Identities = 18/61 (29%), Positives = 30/61 (48%), Gaps = 2/61 (3%)
Frame = +3
Query: 312 STGKYCTHCKKPGHTVDVCYRLHGYPTTSKSKPAF--NNVSHINNIT*GYDSEDDQEESS 369
STG C C + G+ D+C T S S F N++SH NN++ ++ ++E
Sbjct: 459 STGTIC--CDRSGYRSDICVMKGDIRTHSSSSSIFLYNSISHGNNVSRTIEARKGEDEED 632
Query: 370 K 370
+
Sbjct: 633 Q 635
>TC92043 similar to GP|21554299|gb|AAM63374.1 apospory-associated protein C
{Arabidopsis thaliana}, partial (37%)
Length = 1163
Score = 31.2 bits (69), Expect = 2.4
Identities = 20/70 (28%), Positives = 29/70 (40%)
Frame = -3
Query: 277 GENEESKSLVKMAEGKKTYGKGKAPGSSSSGSGYKSTGKYCTHCKKPGHTVDVCYRLHGY 336
G E SK V + G K G A S+ S C + P T+D + G+
Sbjct: 849 GTTESSKRPVHSSPGFKVIGFPTAAPSTQS---------ICLYSSSPKSTIDFDFFSQGF 697
Query: 337 PTTSKSKPAF 346
TT+ +P+F
Sbjct: 696 QTTTSGRPSF 667
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.331 0.141 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,290,916
Number of Sequences: 36976
Number of extensions: 607701
Number of successful extensions: 3821
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 2834
Number of HSP's successfully gapped in prelim test: 120
Number of HSP's that attempted gapping in prelim test: 900
Number of HSP's gapped (non-prelim): 3059
length of query: 1167
length of database: 9,014,727
effective HSP length: 107
effective length of query: 1060
effective length of database: 5,058,295
effective search space: 5361792700
effective search space used: 5361792700
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0334a.1


