

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0333b.7
(293 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
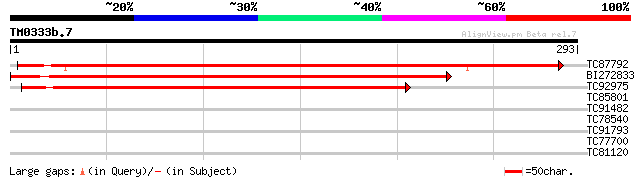
Score E
Sequences producing significant alignments: (bits) Value
TC87792 similar to GP|21593191|gb|AAM65140.1 dessication-related... 322 1e-88
BI272833 weakly similar to SP|P22242|DRPE Desiccation-related pr... 275 2e-74
TC92975 weakly similar to SP|P22242|DRPE_CRAPL Desiccation-relat... 238 3e-63
TC85801 weakly similar to GP|1311386|pdb|1CBG| Cyanogenic Beta-G... 32 0.29
TC91482 similar to GP|21703103|gb|AAM74494.1 AT3g54680/T5N23_40 ... 30 1.1
TC78540 similar to GP|15010680|gb|AAK73999.1 AT5g06580/F15M7_11 ... 30 1.1
TC91793 similar to GP|11275527|dbj|BAB18292. putative receptor-l... 30 1.4
TC77700 similar to PIR|D96779|D96779 probable 3-ketoacyl-ACP syn... 28 5.5
TC81120 similar to GP|7385215|gb|AAF61737.1| beta-ketoacyl-ACP s... 27 7.1
>TC87792 similar to GP|21593191|gb|AAM65140.1 dessication-related protein
putative {Arabidopsis thaliana}, partial (76%)
Length = 1202
Score = 322 bits (824), Expect = 1e-88
Identities = 160/294 (54%), Positives = 218/294 (73%), Gaps = 12/294 (4%)
Frame = +3
Query: 5 TSSIFFLLSSLVLSLFIPIFCSF-----EDVVDVDLVQFHLNLEFLEAEFYFYGTLGWGL 59
T+SI L + L+L+ I CSF ++ DVDL++F LNLE+LEAEF+ +G+LG GL
Sbjct: 57 TASIVVLHAFLILT---QISCSFVLTPPKNYSDVDLLEFPLNLEYLEAEFFLFGSLGHGL 227
Query: 60 DHADPKLAQGGPPPVGGQYAYLDPILRDIFSQFAFQQAGHLRAITEEVKGFPRPLLNISK 119
D P+LA+GGPPP+G + A L ++RD+ QF Q+ GHLRAI V+GFPRPLL++SK
Sbjct: 228 DVVAPELAEGGPPPIGAKVARLGDLVRDVILQFGVQEIGHLRAIKSTVRGFPRPLLDLSK 407
Query: 120 EAFAEVINSAFGKPLNPPFDPYANSLNYLLAAYMLPYVGLTGYVGTNPKLQGVRTRNLVS 179
+FA++++SAFG PL+PPFDPYAN +NYL+A+Y++PYVGLTGYVG NP L+ ++ LV+
Sbjct: 408 SSFAKIMDSAFGHPLHPPFDPYANDINYLIASYVIPYVGLTGYVGANPLLRNATSKKLVA 587
Query: 180 GLLGAKSGQDAMIRTLLYERRYLQVKPYQWTVYEVTHRLSLLRNDLGKKQAEGKVV---- 235
GLLG ++GQDA+IRTLLYERR +V PY TV E T+R+S LRN LG + + + +
Sbjct: 588 GLLGVEAGQDAVIRTLLYERRAWKVHPYGVTVAEFTNRISTLRNKLGNEGVKDEGLGFTS 767
Query: 236 ---GQVFGLNMYSLTYPRSPEEILRIVYGSGDEHVPGGFFPKGGNGNIAKSYLH 286
G + + SL+YPR+P+EILRI+YGSG+E VPGGF+PKG +G IA+ YLH
Sbjct: 768 PFSGNILSADNNSLSYPRTPQEILRIIYGSGNESVPGGFYPKGADGRIARYYLH 929
>BI272833 weakly similar to SP|P22242|DRPE Desiccation-related protein
PCC13-62 precursor. {Craterostigma plantagineum},
partial (63%)
Length = 687
Score = 275 bits (702), Expect = 2e-74
Identities = 140/229 (61%), Positives = 176/229 (76%), Gaps = 1/229 (0%)
Frame = +2
Query: 1 MSLHTSSIFFLLSSLVLSLFIPI-FCSFEDVVDVDLVQFHLNLEFLEAEFYFYGTLGWGL 59
M +H SSI FLL+SL+ IPI + S ++ DVDL++F LNLE+LEAEF+ +G G GL
Sbjct: 8 MLIHISSIAFLLTSLL----IPISYSSNINITDVDLLEFPLNLEYLEAEFFLFGATGHGL 175
Query: 60 DHADPKLAQGGPPPVGGQYAYLDPILRDIFSQFAFQQAGHLRAITEEVKGFPRPLLNISK 119
D P+LAQGGPPP+G + A LD RD+ QFA Q+ GH+RAI VKGFPRPLLNISK
Sbjct: 176 DVVAPELAQGGPPPIGAKMALLDTFTRDVIFQFALQEVGHIRAIKSTVKGFPRPLLNISK 355
Query: 120 EAFAEVINSAFGKPLNPPFDPYANSLNYLLAAYMLPYVGLTGYVGTNPKLQGVRTRNLVS 179
E+FA+V+N+AF KPL P FDPYANS+NYLLA+Y++PYVGLTGYVG P+LQ ++ LV+
Sbjct: 356 ESFAQVMNNAFEKPLYPTFDPYANSINYLLASYIIPYVGLTGYVGAIPELQESTSKKLVA 535
Query: 180 GLLGAKSGQDAMIRTLLYERRYLQVKPYQWTVYEVTHRLSLLRNDLGKK 228
LL +SGQDA+IRTLLYERR L+V PY TV E T+R+S+L+N LG K
Sbjct: 536 SLLAVESGQDAVIRTLLYERRKLKVLPYPITVEEFTNRISMLKNKLGNK 682
>TC92975 weakly similar to SP|P22242|DRPE_CRAPL Desiccation-related protein
PCC13-62 precursor. {Craterostigma plantagineum},
partial (53%)
Length = 662
Score = 238 bits (606), Expect = 3e-63
Identities = 118/202 (58%), Positives = 154/202 (75%), Gaps = 1/202 (0%)
Frame = +1
Query: 7 SIFFLLSSLVLSLFIPIFCSFEDVVDVDLVQFHLNLEFLEAEFYFYGTLGWGLDHADPKL 66
SI LL+SL+ ++ + + DVDL++F LNLE+LEAEF+ +G+ G GLD P+L
Sbjct: 49 SIVVLLASLITNV---VATKETTLSDVDLLEFPLNLEYLEAEFFLFGSFGHGLDAVAPEL 219
Query: 67 AQGGPPPVGGQYAYL-DPILRDIFSQFAFQQAGHLRAITEEVKGFPRPLLNISKEAFAEV 125
A GGP P+G + A L D ++ I +F Q+ GHLRAI VKGF RPLLN+SK FA+V
Sbjct: 220 ADGGPSPIGAKVAKLKDRKIKQIIFEFGLQEVGHLRAIKSTVKGFSRPLLNLSKSTFAKV 399
Query: 126 INSAFGKPLNPPFDPYANSLNYLLAAYMLPYVGLTGYVGTNPKLQGVRTRNLVSGLLGAK 185
I++AFGKPL+PPFDPYAN +N+LLA+Y++PYVGLTGYVGTNP LQ +R LV+GLLG +
Sbjct: 400 IDNAFGKPLHPPFDPYANDINFLLASYLIPYVGLTGYVGTNPHLQNAASRQLVAGLLGVE 579
Query: 186 SGQDAMIRTLLYERRYLQVKPY 207
+GQDA+IRTLL+ERR L+VKPY
Sbjct: 580 AGQDAVIRTLLFERRELKVKPY 645
>TC85801 weakly similar to GP|1311386|pdb|1CBG| Cyanogenic Beta-Glucosidase
Mol_id: 1; Molecule: Cyanogenic Beta-Glucosidase; Chain:
Null; Ec: 3.2., partial (72%)
Length = 1732
Score = 32.0 bits (71), Expect = 0.29
Identities = 40/171 (23%), Positives = 70/171 (40%), Gaps = 12/171 (7%)
Frame = +1
Query: 101 RAITEEVKGFPRPLLNISKE--AFAEVINSAFGKPLNPPF---DPYANSLNYLLAAYMLP 155
+A+ +E KGF NI+K+ +A+ FG + +PY+ ++N P
Sbjct: 502 QALEDEYKGFLNK--NIAKDFAVYADFCFKTFGDRVKHWVTLNEPYSYTINGYNGGTFAP 675
Query: 156 -----YVG--LTGYVGTNPKLQGVRTRNLVSGLLGAKSGQDAMIRTLLYERRYLQVKPYQ 208
YV TG T P + NL+ A +Y+R+Y + +
Sbjct: 676 GRCSKYVANCTTGDSSTEPYIVA---HNLILSHAAA---------VRVYKRKYQAHQKGK 819
Query: 209 WTVYEVTHRLSLLRNDLGKKQAEGKVVGQVFGLNMYSLTYPRSPEEILRIV 259
V VTH N + K+A G+ + FG + +TY P+ ++ ++
Sbjct: 820 IGVTLVTHFFEPYSNSVADKKAAGRALDFFFGWFAHPITYGHYPQSMISLL 972
>TC91482 similar to GP|21703103|gb|AAM74494.1 AT3g54680/T5N23_40
{Arabidopsis thaliana}, partial (29%)
Length = 771
Score = 30.0 bits (66), Expect = 1.1
Identities = 19/60 (31%), Positives = 36/60 (59%), Gaps = 1/60 (1%)
Frame = -2
Query: 163 VGTNPKLQGVRTRNLVSGLLGAKSGQDAMIRTLLYERR-YLQVKPYQWTVYEVTHRLSLL 221
+G+ PK G+R RNL L+ A+ G ++TLL + + +++ +++T + HR +LL
Sbjct: 572 IGSIPKT-GIRPRNLPHPLITARLGSQVFLKTLLPRHKPWPKMRRHKFTTF---HRQTLL 405
>TC78540 similar to GP|15010680|gb|AAK73999.1 AT5g06580/F15M7_11
{Arabidopsis thaliana}, partial (77%)
Length = 2163
Score = 30.0 bits (66), Expect = 1.1
Identities = 16/43 (37%), Positives = 22/43 (50%)
Frame = +1
Query: 246 LTYPRSPEEILRIVYGSGDEHVPGGFFPKGGNGNIAKSYLHPN 288
+ YPRS EE+ +IV + +P P GG +I L PN
Sbjct: 493 IVYPRSEEEVSKIVKSCNNHKIP--IVPYGGATSIEGHTLSPN 615
>TC91793 similar to GP|11275527|dbj|BAB18292. putative receptor-like protein
kinase {Oryza sativa (japonica cultivar-group)}, partial
(28%)
Length = 781
Score = 29.6 bits (65), Expect = 1.4
Identities = 27/91 (29%), Positives = 37/91 (39%), Gaps = 2/91 (2%)
Frame = +3
Query: 139 DPYANSLNYLLAAYMLPYVGLTGY--VGTNPKLQGVRTRNLVSGLLGAKSGQDAMIRTLL 196
+PYA L+ YM P L GY V T+ GV +VSG DA LL
Sbjct: 81 NPYAFLCLMNLSGYMAPEYALRGYLSVKTDVFSYGVLVLEIVSGRKNHDLKLDAEKADLL 260
Query: 197 YERRYLQVKPYQWTVYEVTHRLSLLRNDLGK 227
Y W +Y+ + L+ ++GK
Sbjct: 261 ---------SYAWKLYQGGKIMDLIDQNIGK 326
>TC77700 similar to PIR|D96779|D96779 probable 3-ketoacyl-ACP synthase
F9E10.19 [imported] - Arabidopsis thaliana, partial
(36%)
Length = 1059
Score = 27.7 bits (60), Expect = 5.5
Identities = 13/36 (36%), Positives = 21/36 (58%)
Frame = +1
Query: 156 YVGLTGYVGTNPKLQGVRTRNLVSGLLGAKSGQDAM 191
Y L G NP+L+ T++++ LLGA G +A+
Sbjct: 298 YQALIHCFGKNPELRVNSTKSMIGHLLGASGGVEAV 405
>TC81120 similar to GP|7385215|gb|AAF61737.1| beta-ketoacyl-ACP synthetase 2
{Glycine max}, partial (90%)
Length = 1569
Score = 27.3 bits (59), Expect = 7.1
Identities = 13/36 (36%), Positives = 21/36 (58%)
Frame = +2
Query: 156 YVGLTGYVGTNPKLQGVRTRNLVSGLLGAKSGQDAM 191
Y L G NP+L+ T++++ LLGA G +A+
Sbjct: 1244 YQALIHCFGQNPELKVNSTKSMIGHLLGAAGGVEAV 1351
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.142 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,306,275
Number of Sequences: 36976
Number of extensions: 133139
Number of successful extensions: 680
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 672
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 677
length of query: 293
length of database: 9,014,727
effective HSP length: 95
effective length of query: 198
effective length of database: 5,502,007
effective search space: 1089397386
effective search space used: 1089397386
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0333b.7


