

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0328.16
(738 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
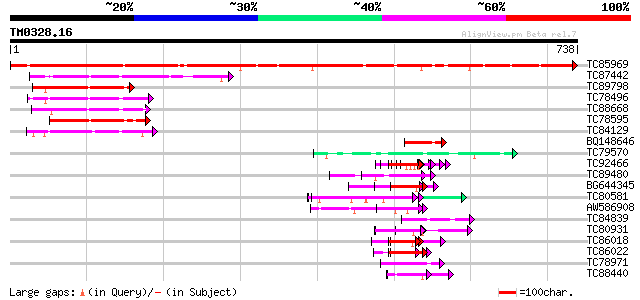
Score E
Sequences producing significant alignments: (bits) Value
TC85969 similar to GP|21553908|gb|AAM62991.1 unknown {Arabidopsi... 1033 0.0
TC87442 similar to GP|15809940|gb|AAL06897.1 AT4g05150/C17L7_70 ... 168 6e-42
TC89798 similar to PIR|T49936|T49936 hypothetical protein F17I14... 123 3e-28
TC78496 similar to PIR|T49936|T49936 hypothetical protein F17I14... 117 1e-26
TC88668 similar to GP|9293928|dbj|BAB01831.1 gb|AAF04880.1~gene_... 96 6e-20
TC78595 similar to PIR|T06718|T06718 hypothetical protein T29H11... 93 4e-19
TC84129 weakly similar to PIR|T51492|T51492 hypothetical protein... 91 2e-18
BQ148646 weakly similar to GP|11596186|gb| cystatin-like protein... 77 2e-14
TC79570 similar to PIR|F86427|F86427 auxin response factor 6 (AR... 73 5e-13
TC92466 similar to GP|11141605|gb|AAG32022.1 LEUNIG {Arabidopsis... 61 2e-09
TC89480 similar to GP|20260684|gb|AAM13240.1 unknown protein {Ar... 59 9e-09
BG644345 weakly similar to GP|19697333|gb putative protein poten... 59 9e-09
TC80581 similar to GP|19683021|gb|AAL92644.1 hypothetical protei... 58 1e-08
AW586908 weakly similar to GP|2104814|emb| pleiotropic effects o... 58 1e-08
TC84839 similar to GP|19386827|dbj|BAB86205. hypothetical protei... 54 2e-07
TC80931 similar to GP|14190783|gb|AAF65166.2 putative phloem tra... 54 2e-07
TC86018 similar to GP|19423939|gb|AAL87312.1 putative RNA helica... 54 3e-07
TC86022 homologue to GP|19423939|gb|AAL87312.1 putative RNA heli... 52 1e-06
TC78971 similar to GP|20260346|gb|AAM13071.1 unknown protein {Ar... 51 1e-06
TC88440 weakly similar to GP|7293965|gb|AAF49324.1| Eip74EF gene... 51 2e-06
>TC85969 similar to GP|21553908|gb|AAM62991.1 unknown {Arabidopsis
thaliana}, partial (31%)
Length = 2756
Score = 1033 bits (2670), Expect = 0.0
Identities = 565/775 (72%), Positives = 613/775 (78%), Gaps = 37/775 (4%)
Frame = +2
Query: 1 MEAPPQPPTPSAAPPLSSVAPPPQPPPHLNYADSVDSSPRSRNTDSWDDPFPP-TSKLRL 59
MEA PQPPTPS PPLSSVAPPP PHLNY DSVDSSPRSRN DSWD+PFPP TSKLRL
Sbjct: 110 MEAQPQPPTPSTGPPLSSVAPPP---PHLNYPDSVDSSPRSRNNDSWDEPFPPATSKLRL 280
Query: 60 MCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATSLADISTRLSKTFLDGRPFTLKYQLPN 119
MCSYGGHIVPRP DK+LCYVGGDTRIVVVERATSL D+STRLSKT L+GRPFT KYQLPN
Sbjct: 281 MCSYGGHIVPRPTDKALCYVGGDTRIVVVERATSLGDLSTRLSKTLLNGRPFTFKYQLPN 460
Query: 120 EDLDSLISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLFPTKPESAHSIPPQILDS 179
EDLDSLISV+TDEDLENMIDEYDRT +T SS+KPSRIRLFLFP+KPESA SIPPQILD
Sbjct: 461 EDLDSLISVSTDEDLENMIDEYDRTNSTN-SSLKPSRIRLFLFPSKPESAQSIPPQILDP 637
Query: 180 SAKSDDWFLDALNGA-GLLNRGFSDSA--SVNCLLGLDDVVAGNNNNIEQGGREGVEAGG 236
+ +S +WF D LNGA LNRGFSDSA SVN LLGL+D VAG NN+IEQG
Sbjct: 638 TVRSGEWFFDTLNGAVTSLNRGFSDSAASSVNHLLGLEDDVAG-NNHIEQG---STGEAP 805
Query: 237 GGASLPGSFANCKNSKQDVSSVPDSPMLETSSSFGSTSSSPALANLPPIRTHAEDGGGGG 296
G S PGSF NCKN KQDV SVPDSPM+ETSSSFGSTSSSP+LANLPPIR H EDG GG
Sbjct: 806 DGVSQPGSFGNCKNLKQDVHSVPDSPMMETSSSFGSTSSSPSLANLPPIRVHVEDGVSGG 985
Query: 297 VRV--------QQQDQKVLGIEEQFAQLGVAAGVGVGQK-QDEGFVVITSPPPPPVPTTL 347
VR QQQDQKVLGI+EQFAQ+ V G G+GQ+ QDEGF V++SPP PPVPTTL
Sbjct: 986 VRAQQQQQLQQQQQDQKVLGIDEQFAQMVVGVG-GIGQRLQDEGFAVMSSPPHPPVPTTL 1162
Query: 348 SAPAMGVPINSAVVSGDYQNRVFSDDERSDHGVPVGYRKPPTPTP-----------QQQA 396
+ A+GVPI SAVV GDYQNRVFSDDERSDHGVPVGYRKPPTP P Q Q
Sbjct: 1163A--AVGVPIGSAVVVGDYQNRVFSDDERSDHGVPVGYRKPPTPQPQVVQVQPQAQNQAQT 1336
Query: 397 QQQLHTQTQSQSHPAQFQQRSS-GGTDLPSPDSVSSDSSLANAMSRQKPVIYQEQVLIQS 455
Q Q +Q Q+QS QFQQ+SS GG+DLPSPDSVSSDSSL+NAMSRQKPV+YQEQ+ IQ
Sbjct: 1337QSQAQSQAQTQSQAPQFQQKSSGGGSDLPSPDSVSSDSSLSNAMSRQKPVVYQEQIQIQP 1516
Query: 456 GTTRVPTHSNPVDPKLNVSDPQSRIQVQQH-HEQGYLLQQQFDQQQHLLQQQFDQQQHQQ 514
GTTRV SNPVDPKLN+SDP SRIQVQQH + GYLLQQQF+ QQ QQQF+ QQ QQ
Sbjct: 1517GTTRV---SNPVDPKLNLSDPHSRIQVQQHVQDPGYLLQQQFELQQQ--QQQFELQQQQQ 1681
Query: 515 QQHQQQQHQQQQHQQQQQQQ----QQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQTHPQ 570
Q Q Q Q QHQ QQQQQ QQQQ+IHG +IHHNP AIP YYPVYPSQQQ
Sbjct: 1682QPQHQPQQSQPQHQPQQQQQLQHHQQQQFIHGGHYIHHNP--AIPTYYPVYPSQQQ---P 1846
Query: 571 HAQVYYVPARQ-AQAYNLSMQQANMGE------SGKPQTPPNPTTLVPQTAAYNNPMRNA 623
H QVYYVPARQ Q YN+S+QQANMGE S +PQ PPNPTTLV Q AAY NP+R+A
Sbjct: 1847HHQVYYVPARQPQQGYNISVQQANMGESATTIPSSRPQNPPNPTTLVQQNAAY-NPIRSA 2023
Query: 624 PLPKTEMTAAGYRTATGGSPQLVQVPANQLQHQQQYVTYSQIHHPSQSIAPNSAPPANYA 683
PLPKTEMTAA YR ATGGSPQ VQVP + QHQQQYVTYSQIHHPSQS+APNSA PA+YA
Sbjct: 2024PLPKTEMTAAAYRAATGGSPQFVQVPTS--QHQQQYVTYSQIHHPSQSMAPNSAAPASYA 2197
Query: 684 YDYVDPAHAQIYYSQPLAPTMPSQYQTMTAAAMVLPEGSSAQLPSDGMKQQTRPS 738
Y+Y DP AQ+YYSQP+APTMPSQYQTM AA M+ PE Q PSDGMKQQ R S
Sbjct: 2198YEYADP--AQVYYSQPMAPTMPSQYQTMAAATMMQPE-VPGQHPSDGMKQQIRTS 2353
>TC87442 similar to GP|15809940|gb|AAL06897.1 AT4g05150/C17L7_70
{Arabidopsis thaliana}, partial (26%)
Length = 1572
Score = 168 bits (426), Expect = 6e-42
Identities = 118/273 (43%), Positives = 164/273 (59%), Gaps = 8/273 (2%)
Frame = +3
Query: 27 PHLNYADSVDSSPRSRNTDSWDDPFPPTSKLRLMCSYGGHIVPRPHDKSLCYVGGDTRIV 86
PHL+ DSV SPRS + + PP ++R MCS+GG I+PRP D L YVGGDTRIV
Sbjct: 99 PHLD-DDSVSPSPRSNHFND----APP--RVRFMCSFGGKILPRPSDNQLRYVGGDTRIV 257
Query: 87 VVERATSLADISTRLSKTFLDGRP-FTLKYQLPNEDLDSLISVTTDEDLENMIDEYDRTT 145
V R+ S + + +LSK L G T KYQLPNEDLD+LI+VTTDED++NMIDEYDR
Sbjct: 258 AVNRSISFSALVHKLSK--LSGMSNITAKYQLPNEDLDALITVTTDEDVDNMIDEYDR-- 425
Query: 146 TTTASSIKPSRIRLFLFPTKPESAHSIPPQILDSSAKSDDWFLDALN-GAGLLNRGFSDS 204
T + + + +R+RLFLFP +S + +L+ S+K ++WF+DALN G L RG S++
Sbjct: 426 VTQSENPRAARLRLFLFPEGEDSRTNSISSLLNGSSKRENWFMDALNGGVSGLERGRSEA 605
Query: 205 ASVNCLLGLDDVVAGNNNNIEQGGREGVEAGGGGASLPGSFANCKNSKQD--VSSVPDSP 262
+S+ + + D + G +NN E S P +QD +S P SP
Sbjct: 606 SSM--VSEVPDYLFGLDNNSEL-----------PESRPKEQQRHLLQQQDNVSNSDPGSP 746
Query: 263 M-LETSSSFGSTS---SSPALANLPPIRTHAED 291
+ +SS F STS S P++ NLPP++T ++
Sbjct: 747 APVVSSSPFCSTSSVLSVPSIPNLPPVKTKLDN 845
>TC89798 similar to PIR|T49936|T49936 hypothetical protein F17I14.190 -
Arabidopsis thaliana, partial (19%)
Length = 1225
Score = 123 bits (308), Expect = 3e-28
Identities = 67/142 (47%), Positives = 92/142 (64%), Gaps = 9/142 (6%)
Frame = +3
Query: 30 NYADSVDSSPRSRNTD--------SWDDPFPPTS-KLRLMCSYGGHIVPRPHDKSLCYVG 80
+Y DS DSSP SR D W+D TS K + MCSYGG I PR HD L YVG
Sbjct: 60 SYPDSGDSSPHSREIDFENPPTTTPWEDQQNQTSYKAKFMCSYGGKIQPRTHDNQLSYVG 239
Query: 81 GDTRIVVVERATSLADISTRLSKTFLDGRPFTLKYQLPNEDLDSLISVTTDEDLENMIDE 140
G+T+I+ V+R + + ++LS + ++ + KYQLP EDLD+LISVT D+DL++M++E
Sbjct: 240 GETKILAVDRNIKFSSMISKLS-SLIEAHDVSFKYQLPGEDLDALISVTNDDDLDHMMNE 416
Query: 141 YDRTTTTTASSIKPSRIRLFLF 162
YDR +A +P+R+RLFLF
Sbjct: 417 YDRLYRASA---RPARMRLFLF 473
>TC78496 similar to PIR|T49936|T49936 hypothetical protein F17I14.190 -
Arabidopsis thaliana, partial (23%)
Length = 1885
Score = 117 bits (294), Expect = 1e-26
Identities = 76/172 (44%), Positives = 100/172 (57%), Gaps = 8/172 (4%)
Frame = +1
Query: 24 QPPPHLNYADSVDSSPRSRNTD-----SWDDPFPPTSKLRLMCSYGGHIVPRPHDKSLCY 78
Q PP +Y DS +SSP SR D SW+D K + MCSYGG I PR HD L Y
Sbjct: 334 QYPP--SYPDSGNSSPLSREIDFENPSSWEDQ--QNYKAKFMCSYGGKIQPRSHDNQLSY 501
Query: 79 VGGDTRIVVVERATSLADISTRLSKTFLDG--RPFTLKYQLPNEDLDSLISVTTDEDLEN 136
+GGDT+I+ V+R+ ++LS T D + + KYQLP E+LD+LISVTT++DLE+
Sbjct: 502 IGGDTKILAVDRSIKFQAFLSKLS-TLCDALQQDISFKYQLPGEELDALISVTTEDDLEH 678
Query: 137 MIDEYDRTTTTTASSIKPSRIRLFLF-PTKPESAHSIPPQILDSSAKSDDWF 187
M+ EYDR S KP R+RLF+F S + P L ++K D F
Sbjct: 679 MMHEYDR---LYRPSSKPVRMRLFIFIAPNSGSVSASQPDPLKPNSKVDFLF 825
>TC88668 similar to GP|9293928|dbj|BAB01831.1
gb|AAF04880.1~gene_id:MFE16.2~similar to unknown protein
{Arabidopsis thaliana}, partial (39%)
Length = 802
Score = 95.5 bits (236), Expect = 6e-20
Identities = 61/167 (36%), Positives = 90/167 (53%), Gaps = 12/167 (7%)
Frame = +2
Query: 29 LNYADSVDSSPRSRNTDSWDDPFP------------PTSKLRLMCSYGGHIVPRPHDKSL 76
LN S P+ + S +P P P S ++ +CSYGG I+PR D L
Sbjct: 74 LNS*TQSSSKPKLNLSQSTSEPEPIMIGLSEPASGSPKSTIKFLCSYGGKILPRYPDGKL 253
Query: 77 CYVGGDTRIVVVERATSLADISTRLSKTFLDGRPFTLKYQLPNEDLDSLISVTTDEDLEN 136
Y+GG R++ V+R+ +++ +L K L+ QLP EDLD+L+S+T+DEDL N
Sbjct: 254 RYLGGHNRVLSVDRSIQFSELLLKL-KELCSSSVTQLRCQLPAEDLDALVSITSDEDLIN 430
Query: 137 MIDEYDRTTTTTASSIKPSRIRLFLFPTKPESAHSIPPQILDSSAKS 183
+I+EYDR TAS +I+ F+ P + + S PP L S +KS
Sbjct: 431 LIEEYDR----TASPQSQLKIKAFISPPRSSNKASKPP--LPSLSKS 553
>TC78595 similar to PIR|T06718|T06718 hypothetical protein T29H11.240 -
Arabidopsis thaliana, partial (46%)
Length = 1059
Score = 92.8 bits (229), Expect = 4e-19
Identities = 51/131 (38%), Positives = 83/131 (62%)
Frame = +1
Query: 53 PTSKLRLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATSLADISTRLSKTFLDGRPFT 112
P++ ++++ SYGG I R D L YVGG TR++ V+R+ S +++ +L + L G T
Sbjct: 145 PSNTIKILYSYGGKIRLRSTDGELRYVGGHTRVLAVDRSISFSELMVKLEE--LCGSSVT 318
Query: 113 LKYQLPNEDLDSLISVTTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLFPTKPESAHSI 172
L+ QLPN DL++LIS+T DEDL N+IDEY+R + P +I+ L ++P+S+ +
Sbjct: 319 LRCQLPNGDLETLISITNDEDLTNIIDEYERASLKLT---HPLKIKAIL--SQPKSSLKV 483
Query: 173 PPQILDSSAKS 183
P + SS+ +
Sbjct: 484 SPDLSSSSSNA 516
>TC84129 weakly similar to PIR|T51492|T51492 hypothetical protein T21H19_140
- Arabidopsis thaliana, partial (6%)
Length = 587
Score = 90.5 bits (223), Expect = 2e-18
Identities = 68/190 (35%), Positives = 105/190 (54%), Gaps = 20/190 (10%)
Frame = +1
Query: 23 PQPPPHL---NYADSVDSSPRSRNT-------DSWDDPFPPTS--KLRLMCSYGGHIVPR 70
P HL ++ +S +SSP SR S+D+P PP++ +++LM +GG I R
Sbjct: 19 PSSMEHLCFNSHPNSGNSSPTSRELLENDHHHRSFDEP-PPSNGPRVKLMIRFGGKIDLR 195
Query: 71 PHD-KSLCYVGGDTRIVVVERATSLADISTRLSK-TFLDGRPFTLKYQLPNEDLDSLISV 128
HD ++ Y+GGDT+I+ V+R+ + + +LS TF D LKYQ P +LDSLISV
Sbjct: 196 LHDDQNYSYIGGDTKILTVDRSIKFSSLIEKLSSMTFSD---LCLKYQQPGGELDSLISV 366
Query: 129 TTDEDLENMIDEYDRTTTTTASSIKPSRIRLFLFPTKPESAHS------IPPQILDSSAK 182
D+DL++M+ EYD S KP+R+++FLFP +A S +++ S
Sbjct: 367 FNDDDLDSMMFEYD---CMCRVSPKPARMKVFLFPLPVNNASSDSLDSAFNLAVVEDSKS 537
Query: 183 SDDWFLDALN 192
F++ LN
Sbjct: 538 DGQMFVNMLN 567
>BQ148646 weakly similar to GP|11596186|gb| cystatin-like protein {Citrus x
paradisi}, partial (34%)
Length = 645
Score = 77.0 bits (188), Expect = 2e-14
Identities = 38/55 (69%), Positives = 40/55 (72%)
Frame = +1
Query: 514 QQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQTH 568
Q Q Q QHQ QQ QQQ Q QQQQ+IHG +IHHNP AIP YYPVYPSQQQ H
Sbjct: 478 QPQQSQPQHQPQQ-QQQLQHHQQQQFIHGGHYIHHNP--AIPTYYPVYPSQQQPH 633
>TC79570 similar to PIR|F86427|F86427 auxin response factor 6 (ARF6)
[imported] - Arabidopsis thaliana, partial (15%)
Length = 1143
Score = 72.8 bits (177), Expect = 5e-13
Identities = 80/292 (27%), Positives = 112/292 (37%), Gaps = 26/292 (8%)
Frame = +2
Query: 396 AQQQLHTQTQSQSHPA---QFQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPVIYQEQVL 452
A Q + T S+ HPA QFQQ P + S+ ++ + Q Q+L
Sbjct: 320 ALQDMRTLDPSKQHPASLLQFQQ----------PQNFSNGNA----------ALMQNQML 439
Query: 453 IQSGTTRV-PTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQ 511
QS + P + P QS+ Q QQ QQ Q Q + QQ
Sbjct: 440 QQSQPHQAFPKNQESQHPS------QSQAQTQQ-------FQQLLQHQHSFTNQNYHMQQ 580
Query: 512 HQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFI-HHNPANAIP--AYYPVYPSQQQTH 568
QQQQ QQQQ QQQQHQQQQQQQ QQQ Q + H +NA+ + + P Q T
Sbjct: 581 QQQQQQQQQQQQQQQHQQQQQQQSQQQ----QQVVDHQQMSNAVSTMSQFVSAPPSQSTQ 748
Query: 569 PQHAQVYYVPARQAQAYNLSMQQANMGESGKPQTP-------------------PNPTTL 609
P A ++ QQ +G P +P P P +
Sbjct: 749 PMQA-----------ISSIGHQQNFSDSNGNPVSPLHNMLGSFSNDETSHLLNFPRPNSW 895
Query: 610 VPQTAAYNNPMRNAPLPKTEMTAAGYRTATGGSPQLVQVPANQLQHQQQYVT 661
VP ++ P + + + A + PQ+ Q+ +Q Q +T
Sbjct: 896 VPVQSSTARPSKRVAVDPLLSSGASHYVL----PQVEQLGQSQTTMSQNAIT 1039
Score = 31.2 bits (69), Expect = 1.5
Identities = 26/85 (30%), Positives = 35/85 (40%), Gaps = 4/85 (4%)
Frame = +2
Query: 393 QQQAQQQLHTQTQSQSHPAQFQQRSSGGTDL----PSPDSVSSDSSLANAMSRQKPVIYQ 448
QQQ QQQ Q Q Q H Q QQ+S + ++VS+ S +A Q Q
Sbjct: 578 QQQQQQQQQQQQQQQQHQQQQQQQSQQQQQVVDHQQMSNAVSTMSQFVSAPPSQSTQPMQ 757
Query: 449 EQVLIQSGTTRVPTHSNPVDPKLNV 473
I ++ NPV P N+
Sbjct: 758 AISSIGHQQNFSDSNGNPVSPLHNM 832
>TC92466 similar to GP|11141605|gb|AAG32022.1 LEUNIG {Arabidopsis thaliana},
partial (14%)
Length = 632
Score = 60.8 bits (146), Expect = 2e-09
Identities = 35/78 (44%), Positives = 42/78 (52%)
Frame = +1
Query: 477 QSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQ 536
Q + Q Q +Q QQQ QQQH+ QQ Q+H QQQ QQQQHQQQ Q QQ Q Q
Sbjct: 385 QQQQQQQPQPQQSQHAQQQ--QQQHMQMQQLLMQRHAQQQQQQQQHQQQPQSQPQQPQPQ 558
Query: 537 QQYIHGTQFIHHNPANAI 554
QQ + + AN +
Sbjct: 559 QQQNRDRTHLLNGSANGL 612
Score = 54.7 bits (130), Expect = 1e-07
Identities = 31/58 (53%), Positives = 33/58 (56%), Gaps = 3/58 (5%)
Frame = +1
Query: 483 QQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQ---QQHQQQQHQQQQHQQQQQQQQQQ 537
QQ +Q Q Q Q QH QQQ Q QQ Q+H QQQ QQQQHQQQ Q Q QQ
Sbjct: 373 QQQQQQQQQQQPQPQQSQHAQQQQQQHMQMQQLLMQRHAQQQQQQQQHQQQPQSQPQQ 546
Score = 52.0 bits (123), Expect = 8e-07
Identities = 35/78 (44%), Positives = 41/78 (51%), Gaps = 3/78 (3%)
Frame = +1
Query: 477 QSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQ---QHQQQQHQQQQQQ 533
Q + Q QH +Q QQQ Q Q LL Q+ QQQ QQQQHQQQ Q QQ Q QQQQ +
Sbjct: 403 QPQPQQSQHAQQQ---QQQHMQMQQLLMQRHAQQQQQQQQHQQQPQSQPQQPQPQQQQNR 573
Query: 534 QQQQQYIHGTQFIHHNPA 551
+ + NPA
Sbjct: 574 DRTHLLNGSANGLAGNPA 627
Score = 49.7 bits (117), Expect = 4e-06
Identities = 28/47 (59%), Positives = 29/47 (61%), Gaps = 1/47 (2%)
Frame = +1
Query: 493 QQQFDQQQHLLQQQFDQQQHQQQQHQQ-QQHQQQQHQQQQQQQQQQQ 538
QQQ QQQ QQ Q QQQQH Q QQ Q+H QQQQQQQQ Q
Sbjct: 379 QQQQQQQQQPQPQQSQHAQQQQQQHMQMQQLLMQRHAQQQQQQQQHQ 519
Score = 48.9 bits (115), Expect = 7e-06
Identities = 28/63 (44%), Positives = 28/63 (44%)
Frame = +1
Query: 504 QQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPS 563
QQQ QQQ QQ Q QQ QH QQQ QQ Q QQ H Q P P P
Sbjct: 373 QQQQQQQQQQQPQPQQSQHAQQQQQQHMQMQQLLMQRHAQQQQQQQQHQQQPQSQPQQPQ 552
Query: 564 QQQ 566
QQ
Sbjct: 553 PQQ 561
Score = 46.6 bits (109), Expect = 3e-05
Identities = 27/52 (51%), Positives = 28/52 (52%), Gaps = 7/52 (13%)
Frame = +1
Query: 493 QQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQ-------QHQQQQQQQQQQ 537
+QQ QQQ Q Q Q QH QQQ QQ QQ Q QQQQQQ QQQ
Sbjct: 370 EQQQQQQQQQQQPQPQQSQHAQQQQQQHMQMQQLLMQRHAQQQQQQQQHQQQ 525
Score = 46.2 bits (108), Expect = 5e-05
Identities = 26/49 (53%), Positives = 28/49 (57%), Gaps = 7/49 (14%)
Frame = +1
Query: 498 QQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQ-------QQQQQQQQY 539
+QQ QQQ Q Q QQ QH QQQ QQ QQ QQQQQQQQ+
Sbjct: 370 EQQQQQQQQQQQPQPQQSQHAQQQQQQHMQMQQLLMQRHAQQQQQQQQH 516
Score = 44.7 bits (104), Expect = 1e-04
Identities = 25/65 (38%), Positives = 29/65 (44%)
Frame = +1
Query: 509 QQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQTH 568
++Q QQQQ QQQQ Q QQ Q QQQQQQ + H + P Q
Sbjct: 367 REQQQQQQQQQQQPQPQQSQHAQQQQQQHMQMQQLLMQRHAQQQQQQQQHQQQPQSQPQQ 546
Query: 569 PQHAQ 573
PQ Q
Sbjct: 547 PQPQQ 561
Score = 33.9 bits (76), Expect = 0.23
Identities = 26/80 (32%), Positives = 35/80 (43%)
Frame = +1
Query: 448 QEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQF 507
Q+Q Q P S + ++ +Q+H QQQ QQQH QQ
Sbjct: 373 QQQQQQQQQQQPQPQQSQHAQQQQQQHMQMQQLLMQRH------AQQQQQQQQH---QQQ 525
Query: 508 DQQQHQQQQHQQQQHQQQQH 527
Q Q QQ Q QQQQ++ + H
Sbjct: 526 PQSQPQQPQPQQQQNRDRTH 585
>TC89480 similar to GP|20260684|gb|AAM13240.1 unknown protein {Arabidopsis
thaliana}, partial (43%)
Length = 1068
Score = 58.5 bits (140), Expect = 9e-09
Identities = 45/105 (42%), Positives = 55/105 (51%), Gaps = 8/105 (7%)
Frame = +1
Query: 458 TRVPT-HSNPVDPKLNVS----DPQSRIQVQQH--HEQGYLLQQQFDQQQHLLQQQFDQQ 510
T +PT +SNP + S DP +Q QQ H+Q QQQ QQ Q QF QQ
Sbjct: 82 TMIPTPNSNPQIQSQSQSQSNQDPNLYLQQQQQFLHQQ----QQQPFQQSQTTQSQFQQQ 249
Query: 511 QHQQQQHQQQQHQQQQHQQQQQQQ-QQQQYIHGTQFIHHNPANAI 554
Q QQ QQQQ + Q QQQQ QQ QQQQ +H + H + N +
Sbjct: 250 QQQQLYQQQQQQRILQQQQQQLQQPQQQQNLHQSLASHFHLLNLV 384
Score = 44.3 bits (103), Expect = 2e-04
Identities = 42/129 (32%), Positives = 56/129 (42%), Gaps = 3/129 (2%)
Frame = +1
Query: 417 SSGGT--DLPSPDSVSSDSSLANAMSRQKPVIYQEQVLIQSGTTRVPTHSNPVDPKLNVS 474
S GGT +P+P+S S + + S Q P +Y +Q + H P
Sbjct: 64 SPGGTWTMIPTPNSNPQIQSQSQSQSNQDPNLYLQQ-------QQQFLHQQQQQPFQQSQ 222
Query: 475 DPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQ-QQHQQQQHQQQQHQQQQHQQQQQQ 533
QS+ Q QQ + L QQQ QQQ +LQQQ Q QQ QQQQ+ Q H +
Sbjct: 223 TTQSQFQQQQQQQ---LYQQQ--QQQRILQQQQQQLQQPQQQQNLHQSLASHFHLLNLVE 387
Query: 534 QQQQQYIHG 542
+ HG
Sbjct: 388 NLAEVVEHG 414
Score = 29.6 bits (65), Expect = 4.4
Identities = 22/82 (26%), Positives = 31/82 (36%)
Frame = +1
Query: 516 QHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQTHPQHAQVY 575
Q Q Q Q QQQQQ++H Q + + + QQQ Q Q
Sbjct: 112 QIQSQSQSQSNQDPNLYLQQQQQFLHQQQQQPFQQSQTTQSQF--QQQQQQQLYQQQQQQ 285
Query: 576 YVPARQAQAYNLSMQQANMGES 597
+ +Q Q QQ N+ +S
Sbjct: 286 RILQQQQQQLQQPQQQQNLHQS 351
>BG644345 weakly similar to GP|19697333|gb putative protein potential
transcriptional repressor Not4hp - Mus musculus, partial
(7%)
Length = 635
Score = 58.5 bits (140), Expect = 9e-09
Identities = 34/65 (52%), Positives = 37/65 (56%)
Frame = +1
Query: 476 PQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQ 535
P S+I +Q QQQ QQQ LQQQ QQ QQQQ Q QQ QQQ QQ Q QQ
Sbjct: 289 PFSQITPFSQQQQQLQFQQQQQQQQQQLQQQQQQQLQQQQQLQLQQQQQQLLQQHQSSQQ 468
Query: 536 QQQYI 540
QQ Y+
Sbjct: 469 QQLYL 483
Score = 55.1 bits (131), Expect = 1e-07
Identities = 30/49 (61%), Positives = 31/49 (63%)
Frame = +1
Query: 496 FDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQ 544
F QQQ LQ Q QQQ QQQ QQQQ Q QQ QQ Q QQQQQQ + Q
Sbjct: 310 FSQQQQQLQFQQQQQQQQQQLQQQQQQQLQQQQQLQLQQQQQQLLQQHQ 456
Score = 48.9 bits (115), Expect = 7e-06
Identities = 43/123 (34%), Positives = 54/123 (42%), Gaps = 6/123 (4%)
Frame = +1
Query: 442 QKPVIYQEQVLIQSGTTRVP--THSNPVD-PKLNVSDPQSRIQVQQHHEQGYLLQQQFDQ 498
Q+P ++Q Q+ + P S+P P + P QQ + Q
Sbjct: 145 QQPSLFQTPGQQQASPFQTPGQQQSSPFQTPGQQQASPFQTPGQQQPSPFSQITPFSQQQ 324
Query: 499 QQHLLQQQFDQQQHQQQQHQQQQHQQQQH---QQQQQQQQQQQYIHGTQFIHHNPANAIP 555
QQ QQQ QQQ Q QQ QQQQ QQQQ QQQQQQ QQ Q ++ + P
Sbjct: 325 QQLQFQQQQQQQQQQLQQQQQQQLQQQQQLQLQQQQQQLLQQHQSSQQQQLYLFTNDKTP 504
Query: 556 AYY 558
A Y
Sbjct: 505 ATY 513
>TC80581 similar to GP|19683021|gb|AAL92644.1 hypothetical protein
{Dictyostelium discoideum}, partial (5%)
Length = 850
Score = 58.2 bits (139), Expect = 1e-08
Identities = 52/165 (31%), Positives = 71/165 (42%), Gaps = 14/165 (8%)
Frame = +1
Query: 388 PTPTPQQQAQQQLH------TQTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAMSR 441
P P Q Q QQQL Q Q Q + QQ+ + L + L M +
Sbjct: 283 PKPMGQPQLQQQLQHQQLLQQQLQLQQQYQRLQQQLASSAQLQQNSLTLNQQQLPLLMQQ 462
Query: 442 QK----PVIYQEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQH----HEQGYLLQ 493
K P++ Q+Q + ++ + +L S Q+QQ+ ++ L+Q
Sbjct: 463 PKSMGQPLLQQQQQQLLQQQLQLQQQYQRLQQQL-----ASSAQLQQNSSTLNQLPQLMQ 627
Query: 494 QQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQ 538
QQ Q LLQQQ QQQ QQ Q QQQQ QQQQ+ Q QQ
Sbjct: 628 QQKSMGQPLLQQQLQQQQLLLQQQQLVLQQQQQQQQQQRTQLLQQ 762
Score = 54.7 bits (130), Expect = 1e-07
Identities = 46/142 (32%), Positives = 63/142 (43%), Gaps = 4/142 (2%)
Frame = +1
Query: 393 QQQAQQQLHTQTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAM-SRQKPVIYQEQV 451
Q Q QQQ Q + AQ QQ S P + S+ + +Q+ + Q+Q+
Sbjct: 346 QLQLQQQYQRLQQQLASSAQLQQNSLTLNQQQLPLLMQQPKSMGQPLLQQQQQQLLQQQL 525
Query: 452 LIQSGTTRVPTH---SNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFD 508
+Q R+ S + + + ++ QQ LLQQQ QQQ LLQQQ
Sbjct: 526 QLQQQYQRLQQQLASSAQLQQNSSTLNQLPQLMQQQKSMGQPLLQQQLQQQQLLLQQQQL 705
Query: 509 QQQHQQQQHQQQQHQQQQHQQQ 530
Q QQQQ QQQ+ Q Q Q Q
Sbjct: 706 VLQQQQQQQQQQRTQLLQQQMQ 771
Score = 43.5 bits (101), Expect = 3e-04
Identities = 53/209 (25%), Positives = 74/209 (35%), Gaps = 4/209 (1%)
Frame = +1
Query: 390 PTPQQQAQQQLHTQTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPVIYQE 449
P Q Q+QQQL + Q Q Q+ L L + Q+ + Q+
Sbjct: 178 PLQQPQSQQQLASSAQLQQSSLTLNQQQL--PQLMQQPKPMGQPQLQQQLQHQQ--LLQQ 345
Query: 450 QVLIQSGTTRVP----THSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQ 505
Q+ +Q R+ + + L ++ Q + +QQ G L QQ QQQ LLQQ
Sbjct: 346 QLQLQQQYQRLQQQLASSAQLQQNSLTLNQQQLPLLMQQPKSMGQPLLQQ--QQQQLLQQ 519
Query: 506 QFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQ 565
Q Q Q Q Q+ QQQ Q QQ Q + + P QQ
Sbjct: 520 QL------QLQQQYQRLQQQLASSAQLQQNSSTLNQLPQLMQQQKSMGQPLLQQQLQQQQ 681
Query: 566 QTHPQHAQVYYVPARQAQAYNLSMQQANM 594
Q V +Q Q + Q M
Sbjct: 682 LLLQQQQLVLQQQQQQQQQQRTQLLQQQM 768
Score = 40.8 bits (94), Expect = 0.002
Identities = 58/234 (24%), Positives = 84/234 (35%), Gaps = 19/234 (8%)
Frame = +1
Query: 443 KPVIYQEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQV----QQHHEQGYLLQQQFDQ 498
+P + Q Q+L Q +P S L + PQS+ Q+ Q L QQQ Q
Sbjct: 109 QPAMIQNQLLKQ-----IPAVSGSAS-LLPLQQPQSQQQLASSAQLQQSSLTLNQQQLPQ 270
Query: 499 QQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYY 558
+ Q QQ QHQQ QQ Q QQQ Q+ QQQ + + + N
Sbjct: 271 LMQQPKPMGQPQLQQQLQHQQLLQQQLQLQQQYQRLQQQ--LASSAQLQQNSLTLNQQQL 444
Query: 559 PVYPSQQQTHPQHAQVYYVPARQAQAYNLSMQQANMGE---------SGKPQTPPNPTT- 608
P+ Q ++ Q P Q Q L QQ + + + Q N +T
Sbjct: 445 PLLMQQPKSMGQ-------PLLQQQQQQLLQQQLQLQQQYQRLQQQLASSAQLQQNSSTL 603
Query: 609 -----LVPQTAAYNNPMRNAPLPKTEMTAAGYRTATGGSPQLVQVPANQLQHQQ 657
L+ Q + P+ L + ++ + Q Q QL QQ
Sbjct: 604 NQLPQLMQQQKSMGQPLLQQQLQQQQLLLQQQQLVLQQQQQQQQQQRTQLLQQQ 765
Score = 30.8 bits (68), Expect = 2.0
Identities = 44/168 (26%), Positives = 55/168 (32%)
Frame = +1
Query: 510 QQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQTHP 569
QQ Q QQ Q QQ QQQ Q + + P+ Q Q
Sbjct: 184 QQPQSQQQLASSAQLQQSSLTLNQQQLPQLMQQPK--------------PMGQPQLQQQL 321
Query: 570 QHAQVYYVPARQAQAYNLSMQQANMGESGKPQTPPNPTTLVPQTAAYNNPMRNAPLPKTE 629
QH Q+ + Q Y QQ + Q N TL Q PL +
Sbjct: 322 QHQQLLQQQLQLQQQYQRLQQQL----ASSAQLQQNSLTLNQQ---------QLPLLMQQ 462
Query: 630 MTAAGYRTATGGSPQLVQVPANQLQHQQQYVTYSQIHHPSQSIAPNSA 677
+ G QL+Q QLQ QQQY Q S + NS+
Sbjct: 463 PKSMGQPLLQQQQQQLLQ---QQLQLQQQYQRLQQQLASSAQLQQNSS 597
>AW586908 weakly similar to GP|2104814|emb| pleiotropic effects on cellular
differentiation and slug behaviour {Dictyostelium
discoideum}, partial (8%)
Length = 668
Score = 57.8 bits (138), Expect = 1e-08
Identities = 60/167 (35%), Positives = 75/167 (43%), Gaps = 20/167 (11%)
Frame = +2
Query: 392 PQQQAQQQLHTQTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPVIY---Q 448
PQ Q QQ TQ QS P Q QQ + + + + N +Q PV Y Q
Sbjct: 134 PQPQQQQPQLTQ-QSPQQPQQQQQHQQPCQNTTMNNGTIGSNQIPNQCVQQ-PVTYSQLQ 307
Query: 449 EQVLIQSGTTRVPTHSNPVDPKLNVSDPQ-SRIQVQQHHEQGYLLQQQFDQQQ------H 501
+Q+L S ++ S + + S PQ S+ Q Q +Q LLQ+Q Q Q
Sbjct: 308 QQLLSGSMQSQQNLQSAGKNGLMMTSLPQDSQFQQQIDQQQAGLLQRQQQQNQLQQSSLQ 487
Query: 502 LLQQQFDQQQHQQQQ----------HQQQQHQQQQHQQQQQQQQQQQ 538
LLQQ Q+ QQ Q QQQQ Q Q QQQQQQQQQQ
Sbjct: 488 LLQQSMLQRGPQQPQMSQTVPQNISDQQQQLQLLQKLQQQQQQQQQQ 628
Score = 42.4 bits (98), Expect = 7e-04
Identities = 30/68 (44%), Positives = 31/68 (45%), Gaps = 2/68 (2%)
Frame = +2
Query: 478 SRIQVQQHHEQGYLLQ--QQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQ 535
S+ Q QQ Q L Q QQQ QQ Q Q Q QQ Q QQ QQ QQQQQ
Sbjct: 23 SQQQQQQQKMQSLLPMPLDQLQQQQQRQQQLAGSQNLPQPQQQQPQLTQQSPQQPQQQQQ 202
Query: 536 QQQYIHGT 543
QQ T
Sbjct: 203 HQQPCQNT 226
Score = 39.7 bits (91), Expect = 0.004
Identities = 49/163 (30%), Positives = 63/163 (38%), Gaps = 26/163 (15%)
Frame = +2
Query: 476 PQSRIQVQQHHEQGYLLQQ---QFDQQQHLLQQQFDQQQHQQQQHQQ------------- 519
P ++Q QQ +Q Q Q QQQ L QQ QQ QQQQHQQ
Sbjct: 71 PLDQLQQQQQRQQQLAGSQNLPQPQQQQPQLTQQSPQQPQQQQQHQQPCQNTTMNNGTIG 250
Query: 520 --QQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANA------IPAYYPVYPSQQQTHPQH 571
Q Q Q Q QQQ + G+ N +A + + QQQ Q
Sbjct: 251 SNQIPNQCVQQPVTYSQLQQQLLSGSMQSQQNLQSAGKNGLMMTSLPQDSQFQQQIDQQQ 430
Query: 572 AQVYYVPARQAQAYNLSMQ--QANMGESGKPQTPPNPTTLVPQ 612
A + +Q Q S+Q Q +M + G Q P + VPQ
Sbjct: 431 AGLLQRQQQQNQLQQSSLQLLQQSMLQRGPQQ--PQMSQTVPQ 553
Score = 38.1 bits (87), Expect = 0.012
Identities = 24/63 (38%), Positives = 29/63 (45%)
Frame = +2
Query: 511 QHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQTHPQ 570
Q QQQQ + Q Q QQQQQ+QQ + G+Q + P P P Q Q Q
Sbjct: 26 QQQQQQQKMQSLLPMPLDQLQQQQQRQQQLAGSQNL-PQPQQQQPQLTQQSPQQPQQQQQ 202
Query: 571 HAQ 573
H Q
Sbjct: 203 HQQ 211
Score = 37.7 bits (86), Expect = 0.016
Identities = 31/81 (38%), Positives = 32/81 (39%)
Frame = +2
Query: 493 QQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPAN 552
QQQ + Q LL DQ Q QQQQ QQQ Q Q QQQ TQ
Sbjct: 32 QQQQQKMQSLLPMPLDQLQ------QQQQRQQQLAGSQNLPQPQQQQPQLTQ-------- 169
Query: 553 AIPAYYPVYPSQQQTHPQHAQ 573
P P QQQ H Q Q
Sbjct: 170 ----QSPQQPQQQQQHQQPCQ 220
Score = 36.2 bits (82), Expect = 0.047
Identities = 52/203 (25%), Positives = 72/203 (34%), Gaps = 11/203 (5%)
Frame = +2
Query: 476 PQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQ 535
P Q+QQ ++ QQQ Q+L Q Q Q Q QQ QQ Q Q QQHQQ Q
Sbjct: 65 PMPLDQLQQQQQR----QQQLAGSQNLPQPQQQQPQLTQQSPQQPQQQ-QQHQQPCQNTT 229
Query: 536 QQQYIHGTQFIHHNPANAIPAYYPVYPS------QQQTHPQHA-----QVYYVPARQAQA 584
G+ I + Y + Q Q + Q A + +P
Sbjct: 230 MNNGTIGSNQIPNQCVQQPVTYSQLQQQLLSGSMQSQQNLQSAGKNGLMMTSLPQDSQFQ 409
Query: 585 YNLSMQQANMGESGKPQTPPNPTTLVPQTAAYNNPMRNAPLPKTEMTAAGYRTATGGSPQ 644
+ QQA + + + Q ++L Q + R P+ T + Q
Sbjct: 410 QQIDQQQAGLLQRQQQQNQLQQSSL--QLLQQSMLQRGPQQPQMSQTVPQNISDQQQQLQ 583
Query: 645 LVQVPANQLQHQQQYVTYSQIHH 667
L+Q Q Q QQQ S H
Sbjct: 584 LLQKLQQQQQQQQQQQPLSTSSH 652
>TC84839 similar to GP|19386827|dbj|BAB86205. hypothetical protein~similar
to Arabidopsis thaliana chromosome 5 T6I14_10, partial
(20%)
Length = 819
Score = 54.3 bits (129), Expect = 2e-07
Identities = 39/98 (39%), Positives = 46/98 (46%), Gaps = 3/98 (3%)
Frame = +1
Query: 511 QHQQ-QQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYY--PVYPSQQQT 567
QHQQ QQHQQQQ QQQQ QQ QQQQQ + H Q + P ++ P P QQ
Sbjct: 175 QHQQHQQHQQQQQQQQQQYQQHQQQQQPPHFHHQQ--QQQQSGPGPDFHRGPPPPMPQQP 348
Query: 568 HPQHAQVYYVPARQAQAYNLSMQQANMGESGKPQTPPN 605
P Q A + NL + G G P PP+
Sbjct: 349 PPMMRQ------PSASSTNLGSEFLPGGPPGPPGPPPH 444
Score = 35.4 bits (80), Expect = 0.080
Identities = 22/44 (50%), Positives = 26/44 (59%)
Frame = +1
Query: 475 DPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQ 518
+PQ + Q QQH +Q QQQ+ QQH QQQ HQQQQ Q
Sbjct: 169 NPQHQ-QHQQHQQQQQQQQQQY--QQHQQQQQPPHFHHQQQQQQ 291
Score = 29.6 bits (65), Expect = 4.4
Identities = 14/35 (40%), Positives = 16/35 (45%)
Frame = +1
Query: 393 QQQAQQQLHTQTQSQSHPAQFQQRSSGGTDLPSPD 427
QQQ QQQ + Q Q Q P F + P PD
Sbjct: 205 QQQQQQQQYQQHQQQQQPPHFHHQQQQQQSGPGPD 309
>TC80931 similar to GP|14190783|gb|AAF65166.2 putative phloem transcription
factor M1 {Apium graveolens}, partial (67%)
Length = 1193
Score = 53.9 bits (128), Expect = 2e-07
Identities = 34/68 (50%), Positives = 39/68 (57%), Gaps = 2/68 (2%)
Frame = +2
Query: 476 PQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQ- 534
PQ + Q+QQ +Q L Q Q +QQ +QQQ Q Q QQQQ Q Q QQ QQQQQQ
Sbjct: 560 PQLQQQMQQQQQQQQLPQLQQNQQLSQIQQQIPQLQQQQQQLPQLQQQQLSQLQQQQQQL 739
Query: 535 -QQQQYIH 541
Q QQ H
Sbjct: 740 PQLQQLQH 763
Score = 46.2 bits (108), Expect = 5e-05
Identities = 33/72 (45%), Positives = 36/72 (49%), Gaps = 6/72 (8%)
Frame = +2
Query: 477 QSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQ 536
Q Q+ Q +Q LQQQ Q L QQQ Q QQQQ Q Q QQ QHQQ QQQQ
Sbjct: 617 QQNQQLSQIQQQIPQLQQQQQQLPQLQQQQLSQL--QQQQQQLPQLQQLQHQQLPQQQQM 790
Query: 537 ------QQYIHG 542
Q Y+ G
Sbjct: 791 VGAGMGQTYVQG 826
Score = 42.0 bits (97), Expect = 9e-04
Identities = 38/104 (36%), Positives = 43/104 (40%), Gaps = 4/104 (3%)
Frame = +2
Query: 503 LQQQFDQQQHQQQQHQQQQHQQ----QQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYY 558
LQQQ QQQ QQQ Q QQ+QQ QQ Q QQQQQQ
Sbjct: 566 LQQQMQQQQQQQQLPQLQQNQQLSQIQQQIPQLQQQQQQ--------------------L 685
Query: 559 PVYPSQQQTHPQHAQVYYVPARQAQAYNLSMQQANMGESGKPQT 602
P QQ + Q Q +Q Q L QQ +G +G QT
Sbjct: 686 PQLQQQQLSQLQQQQQQLPQLQQLQHQQLPQQQQMVG-AGMGQT 814
Score = 40.8 bits (94), Expect = 0.002
Identities = 42/108 (38%), Positives = 47/108 (42%), Gaps = 2/108 (1%)
Frame = +2
Query: 492 LQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQ--QQQQQYIHGTQFIHHN 549
LQQQ QQQ QQQ Q Q QQ Q QQ Q Q QQQQQQ Q QQQ + Q
Sbjct: 566 LQQQMQQQQQ--QQQLPQLQQNQQLSQIQQ-QIPQLQQQQQQLPQLQQQQLSQLQ----- 721
Query: 550 PANAIPAYYPVYPSQQQTHPQHAQVYYVPARQAQAYNLSMQQANMGES 597
QQQ PQ Q+ + Q Q M A MG++
Sbjct: 722 -------------QQQQQLPQLQQLQHQQLPQQQ----QMVGAGMGQT 814
Score = 40.4 bits (93), Expect = 0.002
Identities = 23/48 (47%), Positives = 23/48 (47%)
Frame = +1
Query: 509 QQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPA 556
Q QQQQ Q QQ Q QQHQQ Q QQQ I T A A A
Sbjct: 472 QLSGQQQQQQMQQQQMQQHQQMQSQQQHLSTIAATDATTTTTATASTA 615
Score = 29.6 bits (65), Expect = 4.4
Identities = 29/106 (27%), Positives = 39/106 (36%)
Frame = +2
Query: 432 DSSLANAMSRQKPVIYQEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYL 491
+SS+ + +Q Q+Q L Q + + P+L Q QQ Q
Sbjct: 545 NSSIFPQLQQQMQQQQQQQQLPQLQQNQQLSQIQQQIPQLQQQQQQLPQLQQQQLSQLQQ 724
Query: 492 LQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQ 537
QQQ Q Q L QQ QQQ Q + Q + Q Q Q
Sbjct: 725 QQQQLPQLQQLQHQQLPQQQQMVGAGMGQTYVQGPGGRSQMVSQGQ 862
>TC86018 similar to GP|19423939|gb|AAL87312.1 putative RNA helicase
{Arabidopsis thaliana}, partial (12%)
Length = 885
Score = 53.5 bits (127), Expect = 3e-07
Identities = 30/78 (38%), Positives = 39/78 (49%), Gaps = 6/78 (7%)
Frame = +1
Query: 496 FDQQQHLLQQQFDQQQHQQQQHQQQQH------QQQQHQQQQQQQQQQQYIHGTQFIHHN 549
F Q+ H QQQ+ Q+ Q Q+Q QQH QQQ+QQQ QQQQQQQ++ Q
Sbjct: 553 FQQRPHYQQQQYVQRHLMQNQNQHQQHYQHHQQNQQQYQQQNQQQQQQQWLRRNQLGGGT 732
Query: 550 PANAIPAYYPVYPSQQQT 567
N + S+ T
Sbjct: 733 DTNVVEEVEKTVQSETMT 786
Score = 51.2 bits (121), Expect = 1e-06
Identities = 25/42 (59%), Positives = 28/42 (66%)
Frame = +1
Query: 493 QQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQ 534
QQQ Q+HL+Q Q QQH Q Q QQ QQQ+QQQQQQQ
Sbjct: 574 QQQQYVQRHLMQNQNQHQQHYQHHQQNQQQYQQQNQQQQQQQ 699
Score = 49.3 bits (116), Expect = 5e-06
Identities = 24/46 (52%), Positives = 28/46 (60%)
Frame = +1
Query: 493 QQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQ 538
Q+ QQQ +Q+ Q Q+Q QQH Q Q QQ QQQ QQQQQQ
Sbjct: 559 QRPHYQQQQYVQRHLMQNQNQHQQHYQHHQQNQQQYQQQNQQQQQQ 696
Score = 46.2 bits (108), Expect = 5e-05
Identities = 22/63 (34%), Positives = 35/63 (54%)
Frame = +1
Query: 472 NVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQ 531
N ++ + Q + H++Q +Q+ Q Q+ QQ + Q QQQ+QQQ QQQQ Q +
Sbjct: 529 NPNNANAGFQQRPHYQQQQYVQRHLMQNQNQHQQHYQHHQQNQQQYQQQNQQQQQQQWLR 708
Query: 532 QQQ 534
+ Q
Sbjct: 709 RNQ 717
>TC86022 homologue to GP|19423939|gb|AAL87312.1 putative RNA helicase
{Arabidopsis thaliana}, partial (89%)
Length = 1982
Score = 51.6 bits (122), Expect = 1e-06
Identities = 27/52 (51%), Positives = 33/52 (62%), Gaps = 1/52 (1%)
Frame = +2
Query: 494 QQFDQQQHLLQQQFDQQQHQQQQHQQQ-QHQQQQHQQQQQQQQQQQYIHGTQ 544
QQ Q+ QQQ+ Q+ Q QHQQ Q+QQQ QQ QQQQQQQQ++ Q
Sbjct: 182 QQRPHHQYQQQQQYVQRHMMQNQHQQHYQNQQQNQQQNQQQQQQQQWLRRNQ 337
Score = 51.6 bits (122), Expect = 1e-06
Identities = 27/61 (44%), Positives = 34/61 (55%)
Frame = +2
Query: 474 SDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQ 533
S+ + Q + HH Q+ QQQ +Q+ Q QHQQ QQQ+QQQ QQQQQQ
Sbjct: 161 SNSNTGFQQRPHH--------QYQQQQQYVQRHMMQNQHQQHYQNQQQNQQQNQQQQQQQ 316
Query: 534 Q 534
Q
Sbjct: 317 Q 319
Score = 44.3 bits (103), Expect = 2e-04
Identities = 24/54 (44%), Positives = 29/54 (53%)
Frame = +2
Query: 496 FDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHN 549
F Q+ H QQ QQQ+ Q+ Q QHQQ QQQ QQQ QQ Q++ N
Sbjct: 179 FQQRPHHQYQQ--QQQYVQRHMMQNQHQQHYQNQQQNQQQNQQQQQQQQWLRRN 334
>TC78971 similar to GP|20260346|gb|AAM13071.1 unknown protein {Arabidopsis
thaliana}, partial (48%)
Length = 2260
Score = 51.2 bits (121), Expect = 1e-06
Identities = 34/84 (40%), Positives = 41/84 (48%)
Frame = +1
Query: 483 QQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHG 542
QQ Q L Q Q QQQ QQQ Q QQ Q Q QQ QQQ Q QQQQQQQ ++
Sbjct: 253 QQSQPQQTLFQPQQQQQQSPFQQQSSLFQPQQPQFQPQQQFQQQQFQAQQQQQQQLFLFT 432
Query: 543 TQFIHHNPANAIPAYYPVYPSQQQ 566
PA+ + ++P Q+
Sbjct: 433 ND---KTPASYSTNWADLHPDSQK 495
Score = 40.8 bits (94), Expect = 0.002
Identities = 26/57 (45%), Positives = 28/57 (48%)
Frame = +1
Query: 474 SDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQ 530
S PQ + Q +Q Q F QQ L Q Q Q Q QQQ QQQ QQQ QQQ
Sbjct: 259 SQPQQTLFQPQQQQQ----QSPFQQQSSLFQPQQPQFQPQQQFQQQQFQAQQQQQQQ 417
Score = 36.6 bits (83), Expect = 0.036
Identities = 21/51 (41%), Positives = 25/51 (48%), Gaps = 3/51 (5%)
Frame = +1
Query: 469 PKLNVSDPQSRIQVQQHHEQGYLLQQQ---FDQQQHLLQQQFDQQQHQQQQ 516
P+ + PQ + Q +Q L Q Q F QQ QQQF QQ QQQQ
Sbjct: 265 PQQTLFQPQQQQQQSPFQQQSSLFQPQQPQFQPQQQFQQQQFQAQQQQQQQ 417
Score = 29.3 bits (64), Expect = 5.7
Identities = 21/60 (35%), Positives = 23/60 (38%)
Frame = +1
Query: 507 FDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQ 566
F+ QQ Q QQ Q QQ QQQ QQQ Q P + P QQQ
Sbjct: 235 FNFSTPQQSQPQQTLFQPQQQQQQSPFQQQSSLFQPQQ----------PQFQPQQQFQQQ 384
Score = 28.5 bits (62), Expect = 9.7
Identities = 14/28 (50%), Positives = 16/28 (57%)
Frame = +1
Query: 391 TPQQQAQQQLHTQTQSQSHPAQFQQRSS 418
TPQQ QQ Q Q Q + FQQ+SS
Sbjct: 247 TPQQSQPQQTLFQPQQQQQQSPFQQQSS 330
>TC88440 weakly similar to GP|7293965|gb|AAF49324.1| Eip74EF gene product
{Drosophila melanogaster}, partial (5%)
Length = 711
Score = 50.8 bits (120), Expect = 2e-06
Identities = 27/62 (43%), Positives = 37/62 (59%), Gaps = 3/62 (4%)
Frame = +2
Query: 491 LLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQ---QQYIHGTQFIH 547
LL F + Q + Q Q ++ QHQ+Q HQQ+ HQQ+ H QQ+Q QQ QQ + Q H
Sbjct: 179 LLNLLFVKHQEM-QHQLEEMQHQEQPHQQEDHQQELHHQQEQPHQQEDHQQELPHQQEDH 355
Query: 548 HN 549
H+
Sbjct: 356 HH 361
Score = 45.4 bits (106), Expect = 8e-05
Identities = 25/86 (29%), Positives = 38/86 (44%)
Frame = +2
Query: 492 LQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPA 551
+Q Q ++ QH +Q Q+ HQQ+ H QQ+ QQ QQ+ QQ+ H + A
Sbjct: 212 MQHQLEEMQHQ-EQPHQQEDHQQELHHQQEQPHQQEDHQQELPHQQEDHHHLHLVQQQQA 388
Query: 552 NAIPAYYPVYPSQQQTHPQHAQVYYV 577
+ V Q P +V Y+
Sbjct: 389 LQLQLPVLVVAQLPQALPTKLKVVYL 466
Score = 41.6 bits (96), Expect = 0.001
Identities = 26/65 (40%), Positives = 33/65 (50%), Gaps = 3/65 (4%)
Frame = +2
Query: 477 QSRIQVQQHHEQGYLLQQQFDQQQHLLQQQ---FDQQQHQQQQHQQQQHQQQQHQQQQQQ 533
Q +++ QH EQ + QQ D QQ L QQ Q+ HQQ+ QQ+ H QQQQ
Sbjct: 215 QHQLEEMQHQEQPH---QQEDHQQELHHQQEQPHQQEDHQQELPHQQEDHHHLHLVQQQQ 385
Query: 534 QQQQQ 538
Q Q
Sbjct: 386 ALQLQ 400
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.311 0.127 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,868,630
Number of Sequences: 36976
Number of extensions: 471075
Number of successful extensions: 16874
Number of sequences better than 10.0: 571
Number of HSP's better than 10.0 without gapping: 5723
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 11667
length of query: 738
length of database: 9,014,727
effective HSP length: 103
effective length of query: 635
effective length of database: 5,206,199
effective search space: 3305936365
effective search space used: 3305936365
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0328.16


