

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0323a.10
(175 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
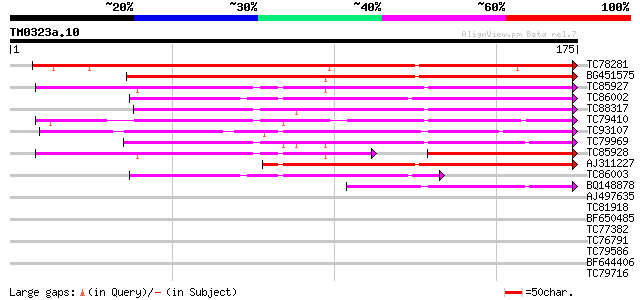
Score E
Sequences producing significant alignments: (bits) Value
TC78281 weakly similar to GP|14423500|gb|AAK62432.1 Unknown prot... 180 2e-46
BG451575 similar to SP|Q41495|ST14 STS14 protein precursor. [Pot... 158 1e-39
TC85927 PR-1 122 9e-29
TC86002 weakly similar to PIR|S10205|S10205 pathogenesis-related... 108 8e-25
TC88317 weakly similar to PIR|T04989|T04989 pathogenesis-related... 102 6e-23
TC79410 similar to PIR|H84518|H84518 pathogenesis-related PR-1-l... 102 1e-22
TC93107 weakly similar to GP|13560653|gb|AAK30143.1 pathogenesis... 101 1e-22
TC79969 weakly similar to PIR|T04232|T04232 pathogenesis-related... 100 4e-22
TC85928 SP|Q40374|PR1_MEDTR Pathogenesis-related protein PR-1 pr... 62 5e-21
AJ311227 similar to PIR|H84518|H845 pathogenesis-related PR-1-li... 94 3e-20
TC86003 weakly similar to GP|7407641|gb|AAF62171.1| pathogenesis... 65 1e-11
BQ148878 weakly similar to GP|4928711|gb|A PR1a precursor {Glyci... 58 2e-09
AJ497635 similar to GP|688429|dbj|B tumor-related protein {Nicot... 28 0.071
TC81918 similar to GP|21594031|gb|AAM65949.1 unknown {Arabidopsi... 32 0.099
BF650485 similar to GP|21594012|gb|A unknown {Arabidopsis thalia... 30 0.64
TC77382 similar to GP|14517512|gb|AAK62646.1 AT4g37300/C7A10_60 ... 30 0.64
TC76791 similar to GP|21311555|gb|AAM46778.1 15.8 kDa oleosin {T... 29 0.83
TC79586 similar to SP|P52425|GPDA_CUPLA Glycerol-3-phosphate deh... 29 1.1
BF644406 similar to GP|13560653|gb| pathogenesis-related protein... 29 1.1
TC79716 weakly similar to SP|O24495|GL2M_ARATH Hydroxyacylglutat... 28 1.9
>TC78281 weakly similar to GP|14423500|gb|AAK62432.1 Unknown protein
{Arabidopsis thaliana}, partial (71%)
Length = 760
Score = 180 bits (457), Expect = 2e-46
Identities = 98/177 (55%), Positives = 121/177 (67%), Gaps = 9/177 (5%)
Frame = +3
Query: 8 FFSLA-LALFLLSAEAA-----TVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEA 61
FFSLA L+L L+S+ AA T + P L+ AKEFL++HN ARAEVGVEPL+WSE
Sbjct: 72 FFSLATLSLLLVSSTAARPAATTPEPTAPPPLTPAAKEFLESHNKARAEVGVEPLQWSEK 251
Query: 62 LAKDAGLFARYQRDRLGCKLANFSHNKYGSNQDLGG--MGWTPSVAVQDWVNEKEFYSYR 119
LAKD L RYQR+++ C AN + +KYG NQ G TPS AV++WV EKEFY +
Sbjct: 252 LAKDTSLLVRYQRNKMACDFANLTASKYGGNQLWAGSAAAVTPSKAVEEWVKEKEFYIHV 431
Query: 120 TNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDK-LILTICFYDPPGNINGESPY 175
N+CV + C YTQVVW+KS Q+GC+Q TC K LTICFYDPPGN+ GESP+
Sbjct: 432 NNTCVV-NHECGVYTQVVWKKSAQLGCSQATCTGKKEASLTICFYDPPGNVIGESPF 599
>BG451575 similar to SP|Q41495|ST14 STS14 protein precursor. [Potato]
{Solanum tuberosum}, partial (48%)
Length = 427
Score = 158 bits (399), Expect = 1e-39
Identities = 77/140 (55%), Positives = 94/140 (67%), Gaps = 1/140 (0%)
Frame = +1
Query: 37 AKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYGSNQDLG 96
A+EFL HN ARA VGVEPL WSE LA RYQRD+L C+ AN + KYG+NQ +
Sbjct: 4 AREFLQTHNQARASVGVEPLTWSEQLANTTSKLVRYQRDKLSCQFANLTAGKYGANQLMA 183
Query: 97 -GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDK 155
G TP + V++WV EKEFY++ N+CV + C YTQVVWRKS ++GCAQ TC +
Sbjct: 184 RGAAVTPRMVVEEWVKEKEFYNHSDNTCVV-NHRCGVYTQVVWRKSVELGCAQTTCGKED 360
Query: 156 LILTICFYDPPGNINGESPY 175
L+ICFY PPGN G SPY
Sbjct: 361 TXLSICFYYPPGNYVGXSPY 420
>TC85927 PR-1
Length = 949
Score = 122 bits (305), Expect = 9e-29
Identities = 73/172 (42%), Positives = 96/172 (55%), Gaps = 5/172 (2%)
Frame = +2
Query: 9 FSLALALFLLSAEAATVLDSVVPQLSAEAK----EFLDAHNWARAEVGVEPLEWSEALAK 64
FS AL L LL+ + + + S +++ +FL N ARA VG+ PL W + L
Sbjct: 62 FSFALFLLLLTFISHVSASYIPNKKSFKSRSFKNQFLIPQNIARAAVGLRPLVWDDKLTH 241
Query: 65 DAGLFARYQRDRLGCKLANFSHNKYGSNQDLG-GMGWTPSVAVQDWVNEKEFYSYRTNSC 123
A +A +R+ C L + S+ YG N G G+GW P+ AV WV+EK+FY+Y NSC
Sbjct: 242 YAQWYANQRRN--DCALEH-SNGPYGENIFWGSGVGWNPAQAVSAWVDEKQFYNYWHNSC 412
Query: 124 VAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
V C HYTQVVW + +VGCA + C DK C YDPPGN GE PY
Sbjct: 413 VDGEM-CGHYTQVVWGSTTKVGCASVVCSDDKGTFMTCNYDPPGNYYGERPY 565
>TC86002 weakly similar to PIR|S10205|S10205 pathogenesis-related protein 1
- common tobacco, partial (72%)
Length = 830
Score = 108 bits (271), Expect = 8e-25
Identities = 54/138 (39%), Positives = 80/138 (57%)
Frame = +1
Query: 38 KEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYGSNQDLGG 97
+++L HN AR++VGV P+ W +A A + + + CK+ + S YG N
Sbjct: 127 QDYLKIHNKARSDVGVGPISWDAKVASYAETYVN--KLKANCKMVH-SKGPYGENLAWSS 297
Query: 98 MGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLI 157
T + AV W+ EK++Y+Y +NSC A + C HYTQVVWR S +VGCA++ C +
Sbjct: 298 GDMTGTAAVTMWIGEKKYYNYNSNSC-AVGYQCGHYTQVVWRDSVRVGCAKVKCNDGRST 474
Query: 158 LTICFYDPPGNINGESPY 175
+ C YDPPGN G+ P+
Sbjct: 475 IISCNYDPPGNYIGQRPF 528
>TC88317 weakly similar to PIR|T04989|T04989 pathogenesis-related protein 1
precursor 19.3K - Arabidopsis thaliana, partial (84%)
Length = 658
Score = 102 bits (255), Expect = 6e-23
Identities = 55/138 (39%), Positives = 79/138 (56%), Gaps = 1/138 (0%)
Frame = +3
Query: 39 EFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHN-KYGSNQDLGG 97
++++AHN AR +VGV + W +A A +A ++ C+L + +YG N
Sbjct: 120 DYVNAHNDARRQVGVANIVWDNTVASFAQDYANQRKG--DCQLIHSGGGGRYGENLAWSS 293
Query: 98 MGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLI 157
+ S AV+ WVNEK Y Y +N+C + C HYTQVVWR S++VGCA++ C ++
Sbjct: 294 GDMSGSDAVKLWVNEKADYDYNSNTCASGKV-CGHYTQVVWRNSQRVGCAKVRCDNNRGT 470
Query: 158 LTICFYDPPGNINGESPY 175
C YDPPGN GE PY
Sbjct: 471 FITCNYDPPGNYVGEKPY 524
>TC79410 similar to PIR|H84518|H84518 pathogenesis-related PR-1-like protein
[imported] - Arabidopsis thaliana, partial (84%)
Length = 1002
Score = 102 bits (253), Expect = 1e-22
Identities = 65/173 (37%), Positives = 92/173 (52%), Gaps = 6/173 (3%)
Frame = +2
Query: 9 FSL--ALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDA 66
FSL L LF++ A +S +++++HN AR +VGV + W LA A
Sbjct: 41 FSLICVLGLFIIIGHVAHAQNSQA--------DYVNSHNEARRQVGVANVVWDNNLATVA 196
Query: 67 GLFARYQRDRLGCKLAN----FSHNKYGSNQDLGGMGWTPSVAVQDWVNEKEFYSYRTNS 122
+A +R C+L + + N GS DL G AV+ WVNEK Y+Y +N+
Sbjct: 197 QNYANSRRG--DCRLTHSGGRYGENLAGSTGDLSGTD-----AVRLWVNEKNDYNYNSNT 355
Query: 123 CVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
C + C HYTQVVWR +K++GCA++ C + IC YDPPGN G+ PY
Sbjct: 356 CASGKV-CGHYTQVVWRNTKRIGCAKVRCNNGGTFI-ICNYDPPGNYVGQKPY 508
>TC93107 weakly similar to GP|13560653|gb|AAK30143.1 pathogenesis-related
protein PR-1 precursor {Capsicum annuum}, partial (68%)
Length = 560
Score = 101 bits (252), Expect = 1e-22
Identities = 61/167 (36%), Positives = 89/167 (52%), Gaps = 1/167 (0%)
Frame = +1
Query: 10 SLALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLF 69
++ LA+F + + + + S+ ++FL+ HN AR EVGV PL W + L A
Sbjct: 22 NMILAIFFICSTLSCMNISLAQN---SPQDFLEVHNQARDEVGVGPLYWEQTLEAYA--- 183
Query: 70 ARYQRDRL-GCKLANFSHNKYGSNQDLGGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSF 128
Y R+ C+L + S YG N G + +V+ W++EK Y Y +NSCV
Sbjct: 184 QNYANKRIKNCELEH-SMGPYGENLAEGYGEVNGTDSVKFWLSEKPNYDYNSNSCVNDE- 357
Query: 129 PCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
C HYTQ++WR S +GCA+ C + + IC Y PPGN+ GE PY
Sbjct: 358 -CGHYTQIIWRDSVHLGCAKSKC-KNGWVFVICSYSPPGNVEGERPY 492
>TC79969 weakly similar to PIR|T04232|T04232 pathogenesis-related protein
homolog F14M19.60 - Arabidopsis thaliana, partial (78%)
Length = 1035
Score = 100 bits (248), Expect = 4e-22
Identities = 58/144 (40%), Positives = 84/144 (58%), Gaps = 4/144 (2%)
Frame = +2
Query: 36 EAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLAN-FSHN--KYGSN 92
E+ EFL HN RA PL W L + A +A ++ CK+ + F + K G N
Sbjct: 251 ESLEFLFRHNLVRASKWELPLMWDYQLEQYARWWASQRKP--DCKVEHSFPEDGFKLGEN 424
Query: 93 QDLG-GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTC 151
G G WTP+ AV+ W +E+++Y+Y TNSCV+ C HYTQ+VW+ ++++GCA++ C
Sbjct: 425 IYWGSGSDWTPTDAVKAWADEEKYYTYVTNSCVSGQM-CGHYTQIVWKSTRRIGCARVVC 601
Query: 152 VVDKLILTICFYDPPGNINGESPY 175
+ +T C YDP GN GE PY
Sbjct: 602 DDGDVFMT-CNYDPVGNYVGERPY 670
>TC85928 SP|Q40374|PR1_MEDTR Pathogenesis-related protein PR-1 precursor.
[Barrel medic] {Medicago truncatula}, partial (92%)
Length = 795
Score = 61.6 bits (148), Expect(2) = 5e-21
Identities = 26/46 (56%), Positives = 29/46 (62%)
Frame = +1
Query: 130 CWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
C HYTQVVW + +VGCA + C DK C YDPPGN GE PY
Sbjct: 361 CGHYTQVVWGSTTKVGCASVVCSDDKGTFMTCNYDPPGNYYGERPY 498
Score = 55.5 bits (132), Expect(2) = 5e-21
Identities = 40/110 (36%), Positives = 58/110 (52%), Gaps = 5/110 (4%)
Frame = +2
Query: 9 FSLALALFLLSAEAATVLDSVVPQLSAEAK----EFLDAHNWARAEVGVEPLEWSEALAK 64
FS AL L LL+ + + + S +++ +FL N ARA VG+ PL W + L
Sbjct: 38 FSFALFLLLLTFISHVSASYIPNKKSFKSRSFKNQFLIPQNIARAAVGLRPLVWDDKLTH 217
Query: 65 DAGLFARYQRDRLGCKLANFSHNKYGSNQDLG-GMGWTPSVAVQDWVNEK 113
A +A +R+ C L + S+ YG N G G+GW P+ AV WV+EK
Sbjct: 218 YAQWYANQRRN--DCALEH-SNGPYGENIFWGSGVGWNPAQAVSAWVDEK 358
>AJ311227 similar to PIR|H84518|H845 pathogenesis-related PR-1-like protein
[imported] - Arabidopsis thaliana, partial (52%)
Length = 379
Score = 94.0 bits (232), Expect = 3e-20
Identities = 44/97 (45%), Positives = 59/97 (60%)
Frame = +2
Query: 79 CKLANFSHNKYGSNQDLGGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVW 138
C++ + S YG N G T + AV W+ EK +Y+Y +NSC A + C HYTQVVW
Sbjct: 5 CQMVH-SRGPYGENLAWGSGDITGTGAVNMWIGEKRYYNYNSNSCAA-GYQCGHYTQVVW 178
Query: 139 RKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
R S +VGCA++ C + + C YDPPGN NG+ PY
Sbjct: 179 RNSVRVGCAKVKCNNGRSTIISCNYDPPGNYNGQRPY 289
>TC86003 weakly similar to GP|7407641|gb|AAF62171.1| pathogenesis-related
protein 1 {Betula pendula}, partial (95%)
Length = 356
Score = 65.5 bits (158), Expect = 1e-11
Identities = 34/97 (35%), Positives = 53/97 (54%)
Frame = +1
Query: 38 KEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYGSNQDLGG 97
+++L HN AR++VGV P+ W +A A + + + CK+ + S YG N
Sbjct: 55 QDYLKIHNKARSDVGVGPISWDAKVASYAETYVN--KLKANCKMVH-SKGPYGENLAWSS 225
Query: 98 MGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYT 134
T + AV W+ EK++Y+Y +NSC A + C HYT
Sbjct: 226 GDMTGTAAVTMWIGEKKYYNYNSNSC-AVGYQCGHYT 333
>BQ148878 weakly similar to GP|4928711|gb|A PR1a precursor {Glycine max},
partial (44%)
Length = 450
Score = 57.8 bits (138), Expect = 2e-09
Identities = 29/71 (40%), Positives = 39/71 (54%)
Frame = +2
Query: 105 AVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYD 164
AV+ W +EK + N C C H+TQVVW S ++GC ++ C +T C Y
Sbjct: 119 AVKGWADEKPHFDNYLNKCFDGE--CHHFTQVVWSGSLRLGCGKVKCNNGGTFVT-CNYY 289
Query: 165 PPGNINGESPY 175
PPGNI G+ PY
Sbjct: 290 PPGNIPGQLPY 322
>AJ497635 similar to GP|688429|dbj|B tumor-related protein {Nicotiana glauca
x Nicotiana langsdorffii}, partial (22%)
Length = 445
Score = 28.5 bits (62), Expect(2) = 0.071
Identities = 11/19 (57%), Positives = 12/19 (62%)
Frame = +2
Query: 130 CWHYTQVVWRKSKQVGCAQ 148
C HYTQVVW +KQ Q
Sbjct: 5 CRHYTQVVWSNTKQTWLCQ 61
Score = 23.1 bits (48), Expect(2) = 0.071
Identities = 8/13 (61%), Positives = 9/13 (68%)
Frame = +1
Query: 163 YDPPGNINGESPY 175
Y PPGN G+ PY
Sbjct: 103 YYPPGNYVGDKPY 141
>TC81918 similar to GP|21594031|gb|AAM65949.1 unknown {Arabidopsis
thaliana}, partial (78%)
Length = 1051
Score = 32.3 bits (72), Expect = 0.099
Identities = 16/38 (42%), Positives = 19/38 (49%)
Frame = +3
Query: 109 WVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGC 146
W E+ F+ Y C S PC HY+ V R K VGC
Sbjct: 588 WTQERCFFEYYQKDC*V-SMPCRHYSPVY*RGFK*VGC 698
>BF650485 similar to GP|21594012|gb|A unknown {Arabidopsis thaliana}, partial
(76%)
Length = 525
Score = 29.6 bits (65), Expect = 0.64
Identities = 13/31 (41%), Positives = 16/31 (50%)
Frame = +1
Query: 116 YSYRTNSCVAPSFPCWHYTQVVWRKSKQVGC 146
Y +TNSC+ S PCW Y+ V GC
Sbjct: 91 YGSKTNSCIK-SHPCWRYSGVC*NSRCNEGC 180
>TC77382 similar to GP|14517512|gb|AAK62646.1 AT4g37300/C7A10_60
{Arabidopsis thaliana}, partial (17%)
Length = 1044
Score = 29.6 bits (65), Expect = 0.64
Identities = 28/113 (24%), Positives = 43/113 (37%), Gaps = 4/113 (3%)
Frame = +3
Query: 55 PLEWSEALAKDA-GLFARYQRDRLGCKLANFSHNKYGSNQDLGGMGWTPSVAVQDWVNEK 113
P S +L K L +Q D AN HNK TP + + K
Sbjct: 66 PRNKSRSLIKSKLNLML*HQNDPSNPTEANLKHNK---------TSLTPLFLHTTYKSLK 218
Query: 114 EFYSYRTNSCVAPSFPC---WHYTQVVWRKSKQVGCAQLTCVVDKLILTICFY 163
F + N + FP W + +W++S C QL + + ++ +CFY
Sbjct: 219 NFNLF--NQILKLYFPQRD*WIHKMSLWKQSITKSCCQLINNITRFVVLLCFY 371
>TC76791 similar to GP|21311555|gb|AAM46778.1 15.8 kDa oleosin {Theobroma
cacao}, partial (64%)
Length = 933
Score = 29.3 bits (64), Expect = 0.83
Identities = 15/50 (30%), Positives = 28/50 (56%)
Frame = +3
Query: 117 SYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPP 166
SY ++SC +P CW + V+W + C+ +C+V + + +C + PP
Sbjct: 342 SYPSSSCYSPITHCW-WLYVLW----WLWCSCYSCIV--MDIQLCLW*PP 470
>TC79586 similar to SP|P52425|GPDA_CUPLA Glycerol-3-phosphate dehydrogenase
[NAD+] (EC 1.1.1.8). {Cuphea lanceolata}, partial (92%)
Length = 1521
Score = 28.9 bits (63), Expect = 1.1
Identities = 15/39 (38%), Positives = 19/39 (48%)
Frame = -3
Query: 7 PFFSLALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHN 45
PFF+ A A FL V+ S PQLS + D +N
Sbjct: 1093 PFFA*ASATFLFLPPKQVVIRSATPQLSKKVLSLTDGNN 977
>BF644406 similar to GP|13560653|gb| pathogenesis-related protein PR-1
precursor {Capsicum annuum}, partial (25%)
Length = 293
Score = 28.9 bits (63), Expect = 1.1
Identities = 18/62 (29%), Positives = 30/62 (48%)
Frame = +1
Query: 13 LALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARY 72
LAL L+ A +S ++++AHN AR +VGV + W +A A +A
Sbjct: 118 LALLLVIGHVANAQNS--------RADYVNAHNDARRQVGVGDIVWDNTVASFAQDYANQ 273
Query: 73 QR 74
++
Sbjct: 274 RK 279
>TC79716 weakly similar to SP|O24495|GL2M_ARATH Hydroxyacylglutathione
hydrolase mitochondrial precursor (EC 3.1.2.6)
(Glyoxalase II) (Glx II)., partial (94%)
Length = 1241
Score = 28.1 bits (61), Expect = 1.9
Identities = 8/30 (26%), Positives = 17/30 (56%)
Frame = -3
Query: 137 VWRKSKQVGCAQLTCVVDKLILTICFYDPP 166
+W++ +++GC C+V+ + C PP
Sbjct: 93 IWKEREEMGCH*FNCLVENEVAFFCLPPPP 4
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.135 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,623,306
Number of Sequences: 36976
Number of extensions: 102997
Number of successful extensions: 536
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 511
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 512
length of query: 175
length of database: 9,014,727
effective HSP length: 90
effective length of query: 85
effective length of database: 5,686,887
effective search space: 483385395
effective search space used: 483385395
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0323a.10


