

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0322.5
(210 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
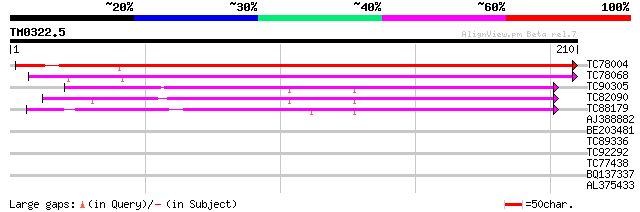
Score E
Sequences producing significant alignments: (bits) Value
TC78004 similar to GP|17979492|gb|AAL50082.1 AT3g22845/MWI23_22 ... 316 4e-87
TC78068 similar to GP|21595553|gb|AAM66112.1 putative coated ves... 127 3e-30
TC90305 similar to PIR|D96610|D96610 probable integral membrane ... 78 3e-15
TC82090 similar to GP|21387055|gb|AAM47931.1 transmembrane prote... 75 1e-14
TC88179 similar to GP|21387055|gb|AAM47931.1 transmembrane prote... 65 2e-11
AJ388882 similar to PIR|D96610|D966 probable integral membrane p... 31 0.30
BE203481 similar to GP|20259685|gb beta-D-glucan exohydrolase {T... 29 1.5
TC89336 weakly similar to GP|21554135|gb|AAM63215.1 unknown {Ara... 27 4.3
TC92292 homologue to PIR|T09572|T09572 cdc2-like protein kinase ... 27 5.7
TC77438 homologue to PIR|T06807|T06807 nucleosome assembly prote... 27 5.7
BQ137337 27 7.4
AL375433 27 7.4
>TC78004 similar to GP|17979492|gb|AAL50082.1 AT3g22845/MWI23_22
{Arabidopsis thaliana}, partial (88%)
Length = 1028
Score = 316 bits (810), Expect = 4e-87
Identities = 153/209 (73%), Positives = 176/209 (84%), Gaps = 1/209 (0%)
Frame = +3
Query: 3 WFSKICTLVIFFLGTLIYNVFSLSISLNDVECVSEHA-QSGDSVTGNFVVMDHDIFWSSD 61
W + +C + IF + + SLS+++ DVEC+ E+ GD+V+GNFVV+DHDIFWSSD
Sbjct: 204 WMALMCLVFIF-----VDRIESLSVTVQDVECLYEYVLYEGDTVSGNFVVVDHDIFWSSD 368
Query: 62 HPGIDFTVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFHNPMSTPETVSFYIHVGHI 121
HPGIDFTVT+PGGN V +KGTSGDKF+FKAP +G+YKFCFHNP STPETVSFYIHVGHI
Sbjct: 369 HPGIDFTVTSPGGNVVQDIKGTSGDKFQFKAPVHGMYKFCFHNPYSTPETVSFYIHVGHI 548
Query: 122 PSEHDLAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVL 181
PSEHDLAKDEHLDPINVKIAELREALES+ EQKYL+ARD RHR+TNEST RVIFYTV
Sbjct: 549 PSEHDLAKDEHLDPINVKIAELREALESVTAEQKYLRARDARHRHTNESTHKRVIFYTVG 728
Query: 182 EYILLAVTSILQVVYIRRLFSKSFAYNRV 210
EY+LLA S LQV+YIRRLFSKS AYNRV
Sbjct: 729 EYLLLAAVSALQVIYIRRLFSKSVAYNRV 815
>TC78068 similar to GP|21595553|gb|AAM66112.1 putative coated vesicle
membrane protein {Arabidopsis thaliana}, partial (97%)
Length = 958
Score = 127 bits (319), Expect = 3e-30
Identities = 72/206 (34%), Positives = 106/206 (50%), Gaps = 3/206 (1%)
Frame = +1
Query: 8 CTLVIFFLGTLIY--NVFSLSISLNDVECVSEHAQ-SGDSVTGNFVVMDHDIFWSSDHPG 64
C +VI +G L+ + ++ EC S + GD+V +FVV+ D W G
Sbjct: 142 CVMVIVLIGMLLNWKETEGIRFVIDRDECFSHDVKYEGDTVHVSFVVIKADSPWHYGDEG 321
Query: 65 IDFTVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFHNPMSTPETVSFYIHVGHIPSE 124
+D V P G + + + DKFEF A + G++KFCF N ET+ F +HVGH
Sbjct: 322 VDLVVKGPAGEQIQDFRDKTSDKFEFVAHKPGVHKFCFTNKSPYHETIDFDVHVGHFSYF 501
Query: 125 HDLAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEYI 184
AKDEH +P+ +IA+L EAL +I EQ +L+A+ +R N+ R + + E
Sbjct: 502 EQHAKDEHFNPLLEQIAKLEEALYNIQFEQHWLEAQTDRQAIVNDGMSRRAVHKAIFESA 681
Query: 185 LLAVTSILQVVYIRRLFSKSFAYNRV 210
L S LQV +RRLF + +RV
Sbjct: 682 ALIGASALQVYLLRRLFERKLGTSRV 759
>TC90305 similar to PIR|D96610|D96610 probable integral membrane protein
T8L23.9 [imported] - Arabidopsis thaliana, partial (65%)
Length = 1072
Score = 77.8 bits (190), Expect = 3e-15
Identities = 52/186 (27%), Positives = 96/186 (50%), Gaps = 3/186 (1%)
Frame = +2
Query: 21 NVFSLSISLNDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHPGIDFTVTNPGGNAVHSL 80
N LSI + +CVS+ Q+ V N+ V+ +I P I VT+P GN +H+
Sbjct: 245 NAIWLSIPSSGTKCVSQDIQTHVVVLANYYVVADNI-QGHPLPTISAKVTSPYGNNLHNN 421
Query: 81 KGTSGDKFEFKAPQNGIYKFCF--HNPMSTPETVSFYIHVGHIPSEHD-LAKDEHLDPIN 137
+ + +F F ++G Y CF ++ ++S G + D +AK E ++ +
Sbjct: 422 ENVTQGQFAFTTTESGSYVACFWMNSKNLAGSSISLEWKTGISAKDWDSVAKKEKIEGVE 601
Query: 138 VKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEYILLAVTSILQVVYI 197
+++ +L A+++I YLK R+ R +E+T RV +++++ + S+LQ++Y+
Sbjct: 602 LELRKLDGAVQAIHENLLYLKNREAEMREVSEATNGRVAWFSIMSLGVCVSVSVLQLLYL 781
Query: 198 RRLFSK 203
RR F K
Sbjct: 782 RRYFRK 799
>TC82090 similar to GP|21387055|gb|AAM47931.1 transmembrane protein-like
protein {Arabidopsis thaliana}, partial (62%)
Length = 829
Score = 75.5 bits (184), Expect = 1e-14
Identities = 55/198 (27%), Positives = 98/198 (48%), Gaps = 7/198 (3%)
Frame = +3
Query: 13 FFLGTLIYNVFSLSISL--NDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHPGIDFTVT 70
FFL L + F++ ++L +CVSE Q+ V ++VV+ D + P + VT
Sbjct: 132 FFLINLPPSTFAIWLTLPTTGTKCVSEEIQNNVVVLADYVVVPDD---HTRSPTLAVKVT 302
Query: 71 NPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCF----HNPMSTPETVSFYIHVGHIPSEHD 126
+P GN +H + T+ F F + G Y CF N +V+ +G + D
Sbjct: 303 SPYGNNLHHKENTTHGSFAFTTQETGNYLACFWVDGGNQGGHEVSVNLDWKIGIAAKDWD 482
Query: 127 -LAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEYIL 185
+AK E ++ + +++ +L A+E+I YLK R+ + R +E T RV + +++ +
Sbjct: 483 SVAKKEKIEGVELELRKLEGAVEAIHENLLYLKGREAKMRIVSERTNGRVAWLSIMSLGI 662
Query: 186 LAVTSILQVVYIRRLFSK 203
S LQ+ +++R F K
Sbjct: 663 CIGVSALQLWHLKRFFQK 716
>TC88179 similar to GP|21387055|gb|AAM47931.1 transmembrane protein-like
protein {Arabidopsis thaliana}, partial (75%)
Length = 1126
Score = 64.7 bits (156), Expect = 2e-11
Identities = 53/201 (26%), Positives = 95/201 (46%), Gaps = 4/201 (1%)
Frame = +3
Query: 7 ICTLVIFFLGTLIYNVFSLSISLNDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHPGID 66
+C +F I+ L+I + +CVSE Q+ V ++ V+ + + I
Sbjct: 165 LCLTTLFLKTEAIW----LTIPSSGTKCVSEDIQTHVVVLADYYVVADEGLHT-----IS 317
Query: 67 FTVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFHNPMSTPE---TVSFYIHVGHIPS 123
VT+P GN VH + + +F F ++G Y CF PE +VS G
Sbjct: 318 AKVTSPFGNNVHHNENVTQGQFAFTTTESGNYVACFWMDGKHPEGSVSVSLDWKTGISAK 497
Query: 124 EHD-LAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLE 182
+ D +AK E ++ + ++I +L +++I YLK ++ R +E T RV +++++
Sbjct: 498 DWDSVAKKEKIEGVELEIRKLEGIVDAIHDYLIYLKEKEATMREVSEKTNARVAWFSIMS 677
Query: 183 YILLAVTSILQVVYIRRLFSK 203
L + S LQ+ Y++ F K
Sbjct: 678 LGLCILVSGLQLWYLQCYFRK 740
>AJ388882 similar to PIR|D96610|D966 probable integral membrane protein
T8L23.9 [imported] - Arabidopsis thaliana, partial (40%)
Length = 588
Score = 31.2 bits (69), Expect = 0.30
Identities = 17/62 (27%), Positives = 33/62 (52%)
Frame = -1
Query: 142 ELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEYILLAVTSILQVVYIRRLF 201
+L ++E+I LK R+ +EST RV +++ + + S+LQ+ +++R F
Sbjct: 411 KLEGSVEAIHENLIRLKXREAGMWXVSESTNARVAWFSFMSLGICIAVSVLQLWHLKRYF 232
Query: 202 SK 203
K
Sbjct: 231 QK 226
>BE203481 similar to GP|20259685|gb beta-D-glucan exohydrolase {Triticum
aestivum}, partial (21%)
Length = 399
Score = 28.9 bits (63), Expect = 1.5
Identities = 22/83 (26%), Positives = 38/83 (45%), Gaps = 1/83 (1%)
Frame = +3
Query: 2 DWFSKICTLVIFFLGTLIYNVFSLSISLNDVECVSEHAQSGDSVTGNFV-VMDHDIFWSS 60
DW + + + ++ ++I V ++ S + V HA D +TG + F S
Sbjct: 3 DWHTLMSLHMPAYIDSIIKGVSTVMASYSSWNGVKMHANR-DLITGYLKNTLKFKGFVIS 179
Query: 61 DHPGIDFTVTNPGGNAVHSLKGT 83
D GID T PG N +S++ +
Sbjct: 180DWQGIDKITTPPGSNYTYSVQAS 248
>TC89336 weakly similar to GP|21554135|gb|AAM63215.1 unknown {Arabidopsis
thaliana}, partial (7%)
Length = 1241
Score = 27.3 bits (59), Expect = 4.3
Identities = 15/42 (35%), Positives = 27/42 (63%)
Frame = +1
Query: 117 HVGHIPSEHDLAKDEHLDPINVKIAELREALESIITEQKYLK 158
HV ++ + ++A +EHL N ++A LR+ LES + E + L+
Sbjct: 691 HVPNLVEQLEVA-NEHLKISNDEVARLRKELESNLAETRQLQ 813
>TC92292 homologue to PIR|T09572|T09572 cdc2-like protein kinase cdc2MsC -
alfalfa, partial (38%)
Length = 711
Score = 26.9 bits (58), Expect = 5.7
Identities = 11/21 (52%), Positives = 14/21 (66%)
Frame = +1
Query: 49 FVVMDHDIFWSSDHPGIDFTV 69
F MDHD+ +D PG+ FTV
Sbjct: 451 FEYMDHDLTGLADRPGMRFTV 513
>TC77438 homologue to PIR|T06807|T06807 nucleosome assembly protein 1 -
garden pea, partial (93%)
Length = 1607
Score = 26.9 bits (58), Expect = 5.7
Identities = 11/23 (47%), Positives = 15/23 (64%)
Frame = -2
Query: 2 DWFSKICTLVIFFLGTLIYNVFS 24
+WFS +CTL+ FL L+ FS
Sbjct: 820 NWFSILCTLLGLFLQNLLCQAFS 752
>BQ137337
Length = 951
Score = 26.6 bits (57), Expect = 7.4
Identities = 11/28 (39%), Positives = 19/28 (67%)
Frame = -2
Query: 1 MDWFSKICTLVIFFLGTLIYNVFSLSIS 28
+D++ I LV+FF+ L Y +++ SIS
Sbjct: 773 LDYYLSILVLVLFFVCDLTYYMYAASIS 690
>AL375433
Length = 522
Score = 26.6 bits (57), Expect = 7.4
Identities = 9/16 (56%), Positives = 11/16 (68%)
Frame = +3
Query: 101 CFHNPMSTPETVSFYI 116
CF NP +P T +FYI
Sbjct: 348 CFRNPACSPSTATFYI 395
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.138 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,798,646
Number of Sequences: 36976
Number of extensions: 93188
Number of successful extensions: 510
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 508
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 508
length of query: 210
length of database: 9,014,727
effective HSP length: 92
effective length of query: 118
effective length of database: 5,612,935
effective search space: 662326330
effective search space used: 662326330
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0322.5


