

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0322.2
(856 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
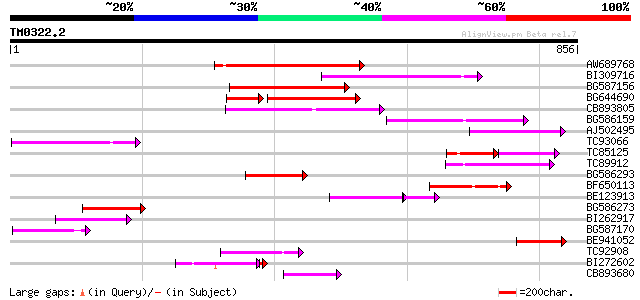
Score E
Sequences producing significant alignments: (bits) Value
AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsi... 287 1e-77
BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F2... 176 3e-44
BG587156 similar to PIR|G85055|G8 probable polyprotein [imported... 174 1e-43
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 137 2e-42
CB893805 similar to GP|10177935|d copia-type polyprotein {Arabid... 159 4e-39
BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2... 147 1e-35
AJ502495 weakly similar to GP|18071369|g putative gag-pol polypr... 111 1e-24
TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative ret... 105 6e-23
TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-relate... 62 7e-22
TC89912 weakly similar to PIR|B84512|B84512 probable retroelemen... 97 3e-20
BG586293 weakly similar to PIR|E84473|E84 probable retroelement ... 95 1e-19
BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vu... 94 2e-19
BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sati... 74 4e-18
BG586273 weakly similar to PIR|F86470|F8 probable retroelement p... 86 8e-17
BI262917 weakly similar to GP|19920130|g Putative retroelement {... 81 1e-15
BG587170 similar to PIR|F86470|F8 probable retroelement polyprot... 78 1e-14
BE941052 weakly similar to PIR|B85188|B85 retrotransposon like p... 78 2e-14
TC92908 similar to GP|10140704|gb|AAG13538.1 putative gag-pol po... 76 5e-14
BI272602 GP|11602812|db avenaIII {Gallus gallus}, partial (3%) 68 2e-13
CB893680 weakly similar to GP|1167523|db ORF(AA 1-1338) {Nicotia... 70 3e-12
>AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsis thaliana},
partial (9%)
Length = 675
Score = 287 bits (735), Expect = 1e-77
Identities = 133/226 (58%), Positives = 173/226 (75%)
Frame = +1
Query: 310 VTRAKDGIVKPRLVPTLLLSQAEPTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPS 369
+TR K G +KP++ L L L DP+W AM+ E+ AL N+TW LVPLP
Sbjct: 1 ITRTKTGKLKPKV---FLTEPCPQDCENLPLSDPRWLQAMKTEYKALIDNKTWDLVPLPP 171
Query: 370 HRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSL 429
H+KAIGCKWV+R+KEN DGSVN++KARLVAKG+SQ G DY ETFSPV+KP+T+R+IL++
Sbjct: 172 HKKAIGCKWVYRVKENPDGSVNKFKARLVAKGFSQTLGCDYTETFSPVIKPVTIRLILTI 351
Query: 430 AVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKDRSLVCRLNKALYGLKQAPRAWF 489
A++ W I Q+D+NNAFLNG L+EEVYM QP GFEA ++SLVC+LNK+LYGLKQAPRAW+
Sbjct: 352 AITYKWEIQQIDINNAFLNGFLQEEVYMSQPQGFEAANKSLVCKLNKSLYGLKQAPRAWY 531
Query: 490 ERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDIIITGNS 535
E L +A ++FGFT S C+PSL +G +Y+ +YVDDI+ITG+S
Sbjct: 532 EXLTSAQIQFGFTKSRCDPSLLIYNQNGACIYLXIYVDDILITGSS 669
>BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (10%)
Length = 744
Score = 176 bits (446), Expect = 3e-44
Identities = 97/244 (39%), Positives = 141/244 (57%)
Frame = +2
Query: 471 VCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDII 530
VC L K++YGLKQA R W+ +L +L+ FG+ S + SLFT + +LVYVDDI+
Sbjct: 20 VCELQKSIYGLKQASRQWYSKLSESLISFGYLQSSSDFSLFTKFKDSSFTTLLVYVDDIV 199
Query: 531 ITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLERAS 590
+ GN + +QH+ L F +K LG+L YFLG++V + +LL Q KY +LLE +
Sbjct: 200 LAGNDISEIQHVKCFLIDRFKIKDLGSLRYFLGLEVAR-SKQGILLNQRKYTLELLEDSG 376
Query: 591 MAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVNKVCQFLS 650
K TP + KL + D + YR ++G L Y+T TRP+IS++V ++ QF+S
Sbjct: 377 NLAVKSTLTPYDISLKLHNSDSPLYNDETQYRRLIGKLIYLTTTRPDISFAVQQLSQFVS 556
Query: 651 QPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSL 710
+P + H+ A R+L+YLK GL S L L +F D+DWA P +S +G
Sbjct: 557 KPQQVHYQAAIRVLQYLKTAPAKGLFYSATS---NLKLSSFADSDWATCPTTRKSVTGYW 727
Query: 711 VFLG 714
VFLG
Sbjct: 728 VFLG 739
>BG587156 similar to PIR|G85055|G8 probable polyprotein [imported] -
Arabidopsis thaliana, partial (17%)
Length = 618
Score = 174 bits (442), Expect = 1e-43
Identities = 90/182 (49%), Positives = 119/182 (64%), Gaps = 1/182 (0%)
Frame = -1
Query: 333 PTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPSHRKAIGCKWVFRIKENADGSVNR 392
P S + A++D +W ++ E A+ N TW LP +KA+ +W+F IK ADGS+ R
Sbjct: 558 PRSYEEAMEDKEWKESVGAEAGAMIKNDTWYESELPKGKKAVSSRWIFTIKYKADGSIER 379
Query: 393 YKARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLE 452
K RLVA+G++ G DY ETF+PV K T+RI+LSLAV+ GW + Q+DV NAFL G+LE
Sbjct: 378 KKTRLVARGFTLTYGEDYIETFAPVAKLHTIRIVLSLAVNLGWGLWQMDVKNAFLQGELE 199
Query: 453 EEVYMVQPPGFE-AKDRSLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLF 511
+EVYM PPG E R V RL KA+YGLKQ+PRAW+ +L L GF S + +LF
Sbjct: 198 DEVYMYPPPGLEHLVKRGNVLRLKKAIYGLKQSPRAWYNKLSTTLNGRGFRKSELDHTLF 19
Query: 512 TL 513
TL
Sbjct: 18 TL 13
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 137 bits (344), Expect(2) = 2e-42
Identities = 71/141 (50%), Positives = 97/141 (68%), Gaps = 1/141 (0%)
Frame = -2
Query: 390 VNRYKARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNG 449
+ R K++LV +GY+Q EG DY E FSPV + +RI+++ A G+ ++Q+DV +AF+NG
Sbjct: 424 ITRNKSKLVVQGYNQKEGIDYDEAFSPVARMEAIRILIAFAAFMGFKLYQMDVKSAFING 245
Query: 450 KLEEEVYMVQPPGFE-AKDRSLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNP 508
L+EEV++ QPPGFE A+ + V RLNK LYGLKQAPRAW+ERL LLK GF +
Sbjct: 244 DLKEEVFVKQPPGFEDAEVPNHVFRLNKTLYGLKQAPRAWYERLSKFLLKNGFKRGKIDN 65
Query: 509 SLFTLYSSGCSVYVLVYVDDI 529
+LF L + + VYVDDI
Sbjct: 64 TLFLLKRE*ELLIIQVYVDDI 2
Score = 54.7 bits (130), Expect(2) = 2e-42
Identities = 23/56 (41%), Positives = 34/56 (60%)
Frame = -3
Query: 328 LSQAEPTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPSHRKAIGCKWVFRIK 383
+S EP + K AL+D W ++MQ+E + ++ W LVP P + IG +WVFR K
Sbjct: 600 ISSIEPKNVKEALRDADWINSMQEELHQFERSKVWYLVPRPEGKTVIGTRWVFRNK 433
>CB893805 similar to GP|10177935|d copia-type polyprotein {Arabidopsis
thaliana}, partial (14%)
Length = 778
Score = 159 bits (402), Expect = 4e-39
Identities = 90/246 (36%), Positives = 139/246 (55%), Gaps = 5/246 (2%)
Frame = +3
Query: 326 LLLSQAEPTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPSHRKAIGCKWVFRIKEN 385
+L ++PT+ + A+K KW ++M +E A N TW L L S K IG KW+F+ K N
Sbjct: 21 MLTMTSDPTTFEEAVKSEKWRASMNNEMEATERNNTWELTDLRSGAKTIGLKWIFKTKLN 200
Query: 386 ADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSLAVS---KGWHIHQLDV 442
+G + +YKARLVAKGYSQ G DY E F+PV + T+R++++LA G I +
Sbjct: 201 ENGEIEKYKARLVAKGYSQQYGVDYTEVFAPVARWDTIRMVIALAAQIKRDGVCIS*M*K 380
Query: 443 NNAFLNGKLEEEVYMVQPPGFEAKDRSLVC-RLNKALYGLKQAPRAWFERLKAALLKFGF 501
++ + + + + + R ++ R+ +ALYGLKQAPRAW+ R++A K GF
Sbjct: 381 AHSCMEN*MRKFLLI----NHRVM*RRVIS*RVKRALYGLKQAPRAWYSRIEAYFTKEGF 548
Query: 502 TASCCNPSLFTLYSSGCSVYVL-VYVDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDY 560
+LF S G + ++ +YVDD+I GN + + + EF++ LG + Y
Sbjct: 549 EKCPYEHTLFVKLSEGGKILIISLYVDDLIFIGNDENMFEEFKKSMKKEFNMSDLGKMHY 728
Query: 561 FLGVQV 566
FLGV+V
Sbjct: 729 FLGVEV 746
>BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2O9.150 -
Arabidopsis thaliana, partial (11%)
Length = 732
Score = 147 bits (372), Expect = 1e-35
Identities = 80/215 (37%), Positives = 124/215 (57%), Gaps = 1/215 (0%)
Frame = +1
Query: 570 HDKSLLLTQSKYIKDLLERASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQ 629
+++ + + Q KY+ DLLER M + P+ KL K D + Y+ IVG L
Sbjct: 16 NEEGIYICQRKYVTDLLERFGMEKSNLSRNPIAPRCKLIKDENGVKVDATKYKQIVGCLM 195
Query: 630 YVTITRPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLI 689
Y+ TRP++ Y ++ + +F++ P E H AVKR+LRYL GT++ G+ K + L
Sbjct: 196 YLAATRPDLMYVLSLISRFMNCPTELHMHAVKRVLRYLNGTINLGIMYKRNGSEK---LE 366
Query: 690 AFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLW 749
A+ D+D+A D DD +STSG + L VSW SKKQ +V S+T+AE+ A + +W
Sbjct: 367 AYTDSDYAGDLDDRKSTSGYVFMLSSGAVSWSSKKQPVVTLSTTKAEFIAAAFCACQSVW 546
Query: 750 VQSLLQELHIPHKTPLIL-CDNMSTVALTHNPVLH 783
++ +L++L + + CDN ST+ L+ NPVLH
Sbjct: 547 MRRVLEKLGYTQSGSITMYCDNNSTIKLSKNPVLH 651
>AJ502495 weakly similar to GP|18071369|g putative gag-pol polyprotein {Oryza
sativa}, partial (9%)
Length = 542
Score = 111 bits (278), Expect = 1e-24
Identities = 55/146 (37%), Positives = 86/146 (58%), Gaps = 1/146 (0%)
Frame = +2
Query: 694 ADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSL 753
+DWA D + +STSG LG +SW SKKQ +VA S+ EAEY + + +W++ +
Sbjct: 2 SDWAGDTETRKSTSGYAFHLGTGAISWSSKKQPVVAFSTAEAEYIASTSCATQTVWLRRI 181
Query: 754 LQELHIPHKTPL-ILCDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPS 812
L+ +H TP I CDN S +AL+ NPV H R+KH+++ +RE + K ++I++ P+
Sbjct: 182 LEVMHHEQNTPTKIYCDNKSAIALSKNPVFHGRSKHIDIQFHKIRELIAEKEVVIEYCPT 361
Query: 813 LDQRDDILTKALSPLRFEDLRTKLNV 838
++ DI TK L F L+ L +
Sbjct: 362 EEKIADIFTKPLKIESFYKLKKMLGM 439
>TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative retroelement
{Oryza sativa} [Oryza sativa (japonica cultivar-group)],
partial (10%)
Length = 823
Score = 105 bits (263), Expect = 6e-23
Identities = 70/201 (34%), Positives = 103/201 (50%), Gaps = 7/201 (3%)
Frame = +1
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEF--KPLTPYFTNLGI 61
+ R WV+ L+ K++T+ TF + ++E Q K VK + TD EF + TN GI
Sbjct: 184 FPRKVWVYFLRYKNETFPTFKKWRILVETQTGKNVKKLITDN*LEFCSSDFNEFCTNHGI 363
Query: 62 VHRFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHIS--LEYWDHAFLTVVYLINRLPTV 119
T P QNG+ ER R ++E +L++A + + W A T +L+NR P
Sbjct: 364 ARHKTIPRNPQQNGVAERMIRTLLERARCMLSNAGL*N*RDLWVEAASTACHLVNRSPHS 543
Query: 120 VLDNESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYK 179
LD + P D LRIFGC Y + N KL+ + EC+FL Y++ K Y+
Sbjct: 544 ALDFKVPEDIWSGNLVDYSNLRIFGCPAYALV---NDGKLAPRAGECIFLSYASESKGYR 714
Query: 180 --CLDP-TGRLYISKDVLFNE 197
C DP + +L +S+DV FNE
Sbjct: 715 LWCSDPKSQKLILSRDVTFNE 777
>TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
(EC 3.4.23.-);, partial (7%)
Length = 705
Score = 61.6 bits (148), Expect(2) = 7e-22
Identities = 34/78 (43%), Positives = 48/78 (60%)
Frame = +1
Query: 660 VKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSLVFLGPNLVS 719
VKRI+RY+KGT + C L++ + D+D+A D D +ST+G + L VS
Sbjct: 1 VKRIMRYIKGTSGVAV----CFGGSELTVRGYVDSDFAGDHDKRKSTTGYVFTLAGGAVS 168
Query: 720 WCSKKQSLVARSSTEAEY 737
W SK Q++VA S+TEAEY
Sbjct: 169 WLSKLQTVVALSTTEAEY 222
Score = 61.2 bits (147), Expect(2) = 7e-22
Identities = 29/91 (31%), Positives = 51/91 (55%)
Frame = +3
Query: 739 NLANTVAELLWVQSLLQELHIPHKTPLILCDNMSTVALTHNPVLHTRTKHMELDIFFVRE 798
+L E +W+Q L++EL + + CD+ S + + NP H+RTKH+ + FVRE
Sbjct: 228 SLPQACKEAIWMQRLMEELGHKQEQITVYCDSQSALHIARNPAFHSRTKHIGIQYHFVRE 407
Query: 799 KVLNKSLIIQHVPSLDQRDDILTKALSPLRF 829
V S+ +Q + + D D +TK+++ +F
Sbjct: 408 VVEEGSVDMQKIHTNDNLADAMTKSINTDKF 500
>TC89912 weakly similar to PIR|B84512|B84512 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(10%)
Length = 814
Score = 96.7 bits (239), Expect = 3e-20
Identities = 49/164 (29%), Positives = 94/164 (56%)
Frame = +1
Query: 659 AVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSLVFLGPNLV 718
A+K +L+YL ++ L + + +L + DAD+A + D +S SG + L +
Sbjct: 4 ALKWVLKYLNESLKSSLKYTKAAQEED-ALEGYVDADYAGNVDTRKSLSGFVFTLYGTTI 180
Query: 719 SWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIPHKTPLILCDNMSTVALTH 778
SW + +QS+V S+T+AEY V + +W++ ++ EL I + I CD+ S + L +
Sbjct: 181 SWKANQQSVVTLSTTQAEYIAFVEGVKDAIWLKGMIGELGITQEYVKIHCDSQSAIHLAN 360
Query: 779 NPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTK 822
+ V H RTKH+++ + F+R+ + +K ++++ + S + D+ TK
Sbjct: 361 HQVYHERTKHIDIRLHFIRDMIESKEIVVEKMASEENPADVFTK 492
>BG586293 weakly similar to PIR|E84473|E84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(7%)
Length = 763
Score = 94.7 bits (234), Expect = 1e-19
Identities = 45/93 (48%), Positives = 65/93 (69%)
Frame = +2
Query: 357 HSNQTWTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFSP 416
+ QT LV P+ K IG +W+++IK N DG++ +YKARLVAKGY + +G D+ E F+P
Sbjct: 44 YQKQTLKLVKKPTGVKPIGLRWIYKIKRNEDGTLIKYKARLVAKGYVKQQGIDFDEVFAP 223
Query: 417 VVKPITVRIILSLAVSKGWHIHQLDVNNAFLNG 449
VV+ T+ ++L+LA + G IH +DV AFLNG
Sbjct: 224 VVRIETI*LLLALAATNGC*IHHIDVKIAFLNG 322
>BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vulgaris},
partial (13%)
Length = 494
Score = 94.4 bits (233), Expect = 2e-19
Identities = 51/125 (40%), Positives = 76/125 (60%), Gaps = 2/125 (1%)
Frame = +1
Query: 635 RPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDA 694
RP+I YSV+ + +F+ P + H A RILRY++GT+++GL + + LI + D+
Sbjct: 121 RPDICYSVSVISKFMHDPRKPHLIAANRILRYVRGTMEYGLLFPYGAKSEVYELICYSDS 300
Query: 695 DWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVA--ELLWVQS 752
DW D RSTSG + +SWC+KKQ + A SS EAEY +A T A + LW+ S
Sbjct: 301 DWCG---DRRSTSGYVFKFNDAAISWCTKKQPITALSSYEAEY--IAGTFATFQALWLDS 465
Query: 753 LLQEL 757
+++EL
Sbjct: 466 VIKEL 480
>BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 503
Score = 73.6 bits (179), Expect(2) = 4e-18
Identities = 45/119 (37%), Positives = 65/119 (53%), Gaps = 1/119 (0%)
Frame = +1
Query: 483 QAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVL-VYVDDIIITGNSLPVLQH 541
Q+PR WF+R + KFG+ + ++F +SS +L VYVDDI +TG+ ++
Sbjct: 1 QSPRDWFDRFT*VVKKFGYIQCQTDHAMFIKHSSTVKKAILIVYVDDIFLTGDHGK*IKR 180
Query: 542 LIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYIKDLLERASMAYAKGISTP 600
L L EF +K LG L YFLG++V K ++Q KY+ DLL+ M K I P
Sbjct: 181 LKNLLAEEFEIKDLGNLKYFLGMEVAR-WKKGSSISQRKYVLDLLKETRMIGCKTIRDP 354
Score = 36.6 bits (83), Expect(2) = 4e-18
Identities = 20/55 (36%), Positives = 30/55 (54%)
Frame = +2
Query: 595 KGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEISYSVNKVCQFL 649
K TPM + KL + D Y+ +VG L Y++ TRP+IS+ V + QF+
Sbjct: 338 KPSETPMDATVKLGTLDNGTLVDKGRYQRLVGKLIYLSHTRPDISFVVCTMSQFM 502
>BG586273 weakly similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 705
Score = 85.5 bits (210), Expect = 8e-17
Identities = 43/95 (45%), Positives = 61/95 (63%), Gaps = 1/95 (1%)
Frame = -2
Query: 111 YLINRLPTVVLDNESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLG 170
YLINR+PT VL +++PF L Q + ++R+FGC CY + +KL S++ +F+G
Sbjct: 698 YLINRIPTRVLKDQAPFEVLNQRKPSLTYMRVFGCLCYVLVPGELRNKLEARSRKAMFIG 519
Query: 171 YSTTHKSYKCLDPTG-RLYISKDVLFNEFRFPYKE 204
YSTT K YKC DP R+ +S+DV F E R Y+E
Sbjct: 518 YSTTQKGYKCYDPEARRVLVSRDVKFIEERGYYEE 414
>BI262917 weakly similar to GP|19920130|g Putative retroelement {Oryza
sativa} [Oryza sativa (japonica cultivar-group)],
partial (8%)
Length = 426
Score = 81.3 bits (199), Expect = 1e-15
Identities = 41/115 (35%), Positives = 59/115 (50%)
Frame = +1
Query: 69 HTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDNESPFT 128
HT QNG+ ER +R ++E A+L A ++ +W A T Y+INR P+ V+D ++P
Sbjct: 82 HTPQQNGVAERMNRTLLERTRAMLKTAGMAKSFWAEAVKTACYVINRSPSTVIDLKTPME 261
Query: 129 KLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLDP 183
D L +FGC Y KL S++C+FLGY+ K Y DP
Sbjct: 262 MWKGKPVDYSSLHVFGCPVYVMYNSQERTKLDPKSRKCIFLGYADNVKGYXLWDP 426
>BG587170 similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 718
Score = 78.2 bits (191), Expect = 1e-14
Identities = 38/119 (31%), Positives = 68/119 (56%), Gaps = 2/119 (1%)
Frame = -3
Query: 5 TRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEFKPLT--PYFTNLGIV 62
++YTW+ + K F NF + + K+K++++D GGE+ + + GI+
Sbjct: 344 SKYTWLTLIPSKDRVIDAFKNFQAYVTNHYHAKIKILRSDNGGEYTSYAFKSHLDHHGIL 165
Query: 63 HRFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVL 121
H+ +CP+T QNG+ +RK++H++E+ +L+ A++S T YLIN +PT VL
Sbjct: 164 HQTSCPYTPQQNGVAKRKNKHLMEVARSLMFQANVS---------TACYLINWIPTKVL 15
>BE941052 weakly similar to PIR|B85188|B85 retrotransposon like protein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 480
Score = 77.8 bits (190), Expect = 2e-14
Identities = 39/76 (51%), Positives = 50/76 (65%)
Frame = +2
Query: 765 LILCDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKAL 824
L+ CD +S LTHNPV H+R KH+ +DI FVR+ V L +QHV ++DQ D LTK L
Sbjct: 20 LLRCDYLSATYLTHNPVYHSRMKHISIDIHFVRDLVQQGKLKVQHVCTVDQLADCLTKPL 199
Query: 825 SPLRFEDLRTKLNVTD 840
S R + LR K+ VTD
Sbjct: 200 SKSRHQLLRNKIGVTD 247
>TC92908 similar to GP|10140704|gb|AAG13538.1 putative gag-pol polyprotein
{Oryza sativa}, partial (1%)
Length = 638
Score = 76.3 bits (186), Expect = 5e-14
Identities = 44/125 (35%), Positives = 69/125 (55%)
Frame = +2
Query: 319 KPRLVPTLLLSQAEPTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPSHRKAIGCKW 378
K R + S EP + A + +W +AM+ E L N T TL PL +K + C+
Sbjct: 239 KHRASLAAITSNIEPKNYVQAAQ*QEWLAAMEQEIQVLEENNTSTLEPLREGKKWVDCRP 418
Query: 379 VFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSLAVSKGWHIH 438
V++I A+G + +YKA+LVAK + QVEG D+ + K R +L++A +KG +H
Sbjct: 419 VYKIIHKANGEIEKYKAQLVAKDFVQVEGEDF*D-LCLSNKDDNCRCLLTIAAAKG*QLH 595
Query: 439 QLDVN 443
+DV+
Sbjct: 596 LMDVS 610
>BI272602 GP|11602812|db avenaIII {Gallus gallus}, partial (3%)
Length = 488
Score = 68.2 bits (165), Expect(2) = 2e-13
Identities = 48/138 (34%), Positives = 67/138 (47%), Gaps = 8/138 (5%)
Frame = +1
Query: 251 SSPASQSSTPILPEPTSPVPQSSPAPISPALSSPAPQSSSTDSSSSVSPIVNPHNTHS-- 308
S+ S S+P E + SSPA SP + A ++S P PHN H+
Sbjct: 7 STQTSSPSSPNSNEVAQNIQISSPA--SPTSNELANITTSPTPPPPPPPTARPHNNHTTN 180
Query: 309 ---MVTRAKDGIVKPRLVPTLL---LSQAEPTSAKLALKDPKWYSAMQDEFNALHSNQTW 362
++TR+++ IVK L L EP++ AL+D W S MQDEF+AL +N TW
Sbjct: 181 VGGIITRSQNNIVKTIKKLNLHVRPLCPIEPSNITQALRDHDWRSIMQDEFDALQNNNTW 360
Query: 363 TLVPLPSHRKAIGCKWVF 380
LV + +GCK F
Sbjct: 361 DLVSRSFAQNLVGCKLGF 414
Score = 26.2 bits (56), Expect(2) = 2e-13
Identities = 10/12 (83%), Positives = 10/12 (83%)
Frame = +2
Query: 378 WVFRIKENADGS 389
WVFRIK N DGS
Sbjct: 407 WVFRIKRNPDGS 442
>CB893680 weakly similar to GP|1167523|db ORF(AA 1-1338) {Nicotiana tabacum},
partial (7%)
Length = 780
Score = 70.5 bits (171), Expect = 3e-12
Identities = 37/88 (42%), Positives = 52/88 (59%)
Frame = -2
Query: 414 FSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKDRSLVCR 473
F P+VK T+ +LS+ + ++ LDV AFL G L E++YM QP GF + +V +
Sbjct: 554 FVPIVKLNTIMFLLSIVAIENLYLE*LDVKTAFLRGDLVEDIYMHQPEGFS*EVGKMVGK 375
Query: 474 LNKALYGLKQAPRAWFERLKAALLKFGF 501
L K++YGLKQ PR LKA + GF
Sbjct: 374 LKKSMYGLKQGPRQCI*SLKALCTRKGF 291
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.135 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,722,354
Number of Sequences: 36976
Number of extensions: 522003
Number of successful extensions: 7543
Number of sequences better than 10.0: 560
Number of HSP's better than 10.0 without gapping: 4943
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 6246
length of query: 856
length of database: 9,014,727
effective HSP length: 104
effective length of query: 752
effective length of database: 5,169,223
effective search space: 3887255696
effective search space used: 3887255696
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0322.2


