

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
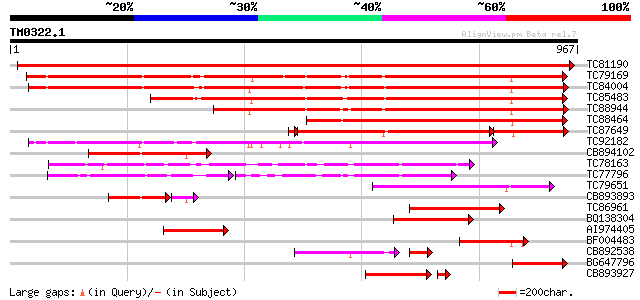
Query= TM0322.1
(967 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC81190 type IIB calcium ATPase [Medicago truncatula] 1601 0.0
TC79169 type IIB calcium ATPase [Medicago truncatula] 809 0.0
TC84004 type IIB calcium ATPase [Medicago truncatula] 785 0.0
TC85483 type IIB calcium ATPase MCA5 [Medicago truncatula] 634 0.0
TC88944 type IIB calcium ATPase [Medicago truncatula] 560 e-159
TC88464 similar to SP|Q9LU41|ACA9_ARATH Potential calcium-transp... 541 e-154
TC87649 similar to SP|Q9SZR1|ACAA_ARATH Potential calcium-transp... 400 e-152
TC92182 type IIA calcium ATPase [Medicago truncatula] 320 1e-87
CB894102 similar to GP|8843813|db Ca2+-transporting ATPase-like ... 189 6e-48
TC78163 H+-ATPase 170 2e-42
TC77796 H+-ATPase 110 2e-41
TC79651 similar to PIR|T52581|T52581 Ca2+-transporting ATPase (E... 147 2e-35
CB893893 similar to GP|8843813|db Ca2+-transporting ATPase-like ... 124 7e-33
TC86961 similar to GP|1742951|emb|CAA70946.1 Ca2+-ATPase {Arabid... 130 3e-30
BQ138304 homologue to GP|6688833|emb putative calcium P-type ATP... 125 6e-29
AI974405 similar to SP|Q9LF79|ACA Calcium-transporting ATPase 8 ... 124 1e-28
BF004483 GP|21314227| type IIB calcium ATPase MCA5 {Medicago tru... 124 1e-28
CB892538 similar to SP|Q9XES1|ECA Calcium-transporting ATPase 4 ... 83 1e-26
BG647796 GP|21314227|g type IIB calcium ATPase MCA5 {Medicago tr... 101 1e-21
CB893927 homologue to GP|17342714|g type IIA calcium ATPase {Med... 84 8e-19
>TC81190 type IIB calcium ATPase [Medicago truncatula]
Length = 3277
Score = 1601 bits (4146), Expect = 0.0
Identities = 829/961 (86%), Positives = 880/961 (91%), Gaps = 10/961 (1%)
Frame = +1
Query: 13 GTNHCSLVPHDV-DKARLSNMVKDKNLEAYSEFGGVEGVADVLGTIPAKGILGSDDDTAA 71
GTNH SLV V DK +L++MVKDKNL++ SEFGGVEGV VLGT P KGI+GSDDD +
Sbjct: 274 GTNHYSLVSDVVVDKTKLADMVKDKNLKSLSEFGGVEGVGHVLGTFPTKGIIGSDDDISR 453
Query: 72 RRELFGTNTYVRPPPKIFLHFVLEALNDTTILILLGCAGLSLGFGIKEHGPGEGWYEGGS 131
R ELFG+NTY +PPPK LHFVLEA NDTTI+ILL CAGLSLGFGIKEHGPGEGWYEGGS
Sbjct: 454 RLELFGSNTYKKPPPKGLLHFVLEAFNDTTIIILLVCAGLSLGFGIKEHGPGEGWYEGGS 633
Query: 132 IFLAVFLVVVVSALSNFRQDRQFDKLSKISNDIKVEVVRNGRPQQISIFDVLVGDVIYLK 191
IFLAVFLVVVVSALSNFRQ+RQF KLSKISN+IKVEVVRNGRPQQISIFDVLVGD++ LK
Sbjct: 634 IFLAVFLVVVVSALSNFRQERQFHKLSKISNNIKVEVVRNGRPQQISIFDVLVGDIVSLK 813
Query: 192 IGDQIPADGLFLGGHSLQVDESSMTGESDHVEIEPLKAPFLLSGAKVVDGYAQMLVTAVG 251
IGDQIPADG+FL G+SLQVDESSMTGESDHVEIEPL+APFLLSGAKVVDGYAQMLVT+VG
Sbjct: 814 IGDQIPADGVFLSGYSLQVDESSMTGESDHVEIEPLRAPFLLSGAKVVDGYAQMLVTSVG 993
Query: 252 ANTAWGQMMSSISGDNSERTPLQARLDKLTSSIGKIGLAVAFLVLLVLLIRYFTGNTEDE 311
NT+WGQMMSSIS D +ERTPLQARLDKLTSSIGK+GLAVAFLVLLVLLIRYFTGN+ DE
Sbjct: 994 KNTSWGQMMSSISRDTNERTPLQARLDKLTSSIGKVGLAVAFLVLLVLLIRYFTGNSHDE 1173
Query: 312 NGNKEYKGSKTDINDVCNAVVSIVAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMMADQAM 371
GNKE++GSKTDINDV N+VVSIVAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMMAD AM
Sbjct: 1174 KGNKEFRGSKTDINDVMNSVVSIVAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMMADHAM 1353
Query: 372 VRKLSACETMGSATVICTDKTGTLTLNQMRVTKFWLGLENVVENFSNAMAPTVLELFHQG 431
VRKLSACETMGSATVICTDKTGTLTLNQMRVTKF LG EN++ENFSNAM P VLELFHQG
Sbjct: 1354 VRKLSACETMGSATVICTDKTGTLTLNQMRVTKFCLGPENIIENFSNAMTPKVLELFHQG 1533
Query: 432 VGLNTTGSVYKPSAESEPEISGSPTEKAMLLWAVSDLGMDMDELKQKHKVLHVETFNSEK 491
VGLNTTGSVY P + SEPEISGSPTEKA+L+WAV DLGMDMDE+KQKHKVLHVETFNSEK
Sbjct: 1534 VGLNTTGSVYNPPSGSEPEISGSPTEKAILMWAVLDLGMDMDEMKQKHKVLHVETFNSEK 1713
Query: 492 KRSGVAVRKET-NNTVHVHWKGAAEMVLAMCSNYIDSNGTQKSLD-EERSKIEKIIQGMA 549
KRSGVA+RKE +N+VHVHWKGAAEM+LAMC+NYIDSNG +KSLD EERSKIE+IIQ MA
Sbjct: 1714 KRSGVAIRKENDDNSVHVHWKGAAEMILAMCTNYIDSNGARKSLDEEERSKIERIIQVMA 1893
Query: 550 ASSLRCIAFAYMEI--SEGGDYIEK--GKPRQVLREDGLTLLGIVGLKDPCRPNVKKAVE 605
ASSLRCIAFA+ EI SE DY+ K K Q+LREDGLTLLGIVGLKDPCRPN KKAVE
Sbjct: 1894 ASSLRCIAFAHTEISDSEDIDYMIKREKKSHQMLREDGLTLLGIVGLKDPCRPNTKKAVE 2073
Query: 606 TCKLAGVDIKMITGDNIFTAKAIATECGILDLND---AGGVVVEGVEFRNYTEEERMEKV 662
TCK AGV+IKMITGDNIFTAKAIA ECGILD N G VVEGVEFR+YTEEERMEKV
Sbjct: 2074 TCKAAGVEIKMITGDNIFTAKAIAIECGILDSNSDHAKAGEVVEGVEFRSYTEEERMEKV 2253
Query: 663 DKIRVMARSSPMDKLLMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKE 722
D IRVMARSSPMDKLLMVQCL+KKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKE
Sbjct: 2254 DNIRVMARSSPMDKLLMVQCLRKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKE 2433
Query: 723 SSDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTT 782
SSDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTT
Sbjct: 2434 SSDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTT 2613
Query: 783 VQLLWVNLIMDTLGALALATERPTKELMQKKPIGRTEPLITKIMWRNLLAQALYQIAVLL 842
VQLLWVNLIMDTLGALALATERPTKELM+KKPIGRT PLIT IMWRNLLAQA YQIAVLL
Sbjct: 2614 VQLLWVNLIMDTLGALALATERPTKELMKKKPIGRTAPLITNIMWRNLLAQASYQIAVLL 2793
Query: 843 VFQFYGKSIFNVSKEVKNTLIFNTFVLCQVFNEFNSRSMEKLNVFEGILKNHLFLGIVGI 902
+ QFYGKSIFNVSKEVK+TLIFNTFVLCQVFNEFNSRSMEKL VFEGILKNHLFLGI+GI
Sbjct: 2794 IMQFYGKSIFNVSKEVKDTLIFNTFVLCQVFNEFNSRSMEKLYVFEGILKNHLFLGIIGI 2973
Query: 903 TIVLQVLMVELLRKFADTERLNWEQWGICIGIAAVSWPIAWLTKLTPVPSKLFFTNAKWV 962
TIVLQ+LMVELLRKFADTERL WEQWGICIGIA VSWP+A L KL PV K F+ KWV
Sbjct: 2974 TIVLQILMVELLRKFADTERLTWEQWGICIGIAVVSWPLACLVKLIPVSDKPSFSYTKWV 3153
Query: 963 K 963
K
Sbjct: 3154 K 3156
>TC79169 type IIB calcium ATPase [Medicago truncatula]
Length = 3565
Score = 809 bits (2090), Expect = 0.0
Identities = 445/935 (47%), Positives = 618/935 (65%), Gaps = 13/935 (1%)
Frame = +2
Query: 29 LSNMVKDKNLEAYSEFGGVEGVADVLGTIPAKGILGSDDDTAARRELFGTNTYVRPPPKI 88
++++V+ + + Y + G V+G+ L +G+ S D +R+E++G N Y P K
Sbjct: 470 IASVVRSHDFKNYKKVGEVQGITSKLSVSVDEGV--SQDSIHSRQEIYGLNRYTEKPSKS 643
Query: 89 FLHFVLEALNDTTILILLGCAGLSLGFGIKEHGPGEGWYEGGSIFLAVFLVVVVSALSNF 148
FL FV +AL+D T++IL+ CA +S+G G+ G +G Y+G I L++FLVV V+A+S++
Sbjct: 644 FLMFVWDALHDLTLIILIVCALVSIGIGLPTEGWPKGVYDGVGILLSIFLVVTVTAVSDY 823
Query: 149 RQDRQFDKLSKISNDIKVEVVRNGRPQQISIFDVLVGDVIYLKIGDQIPADGLFLGGHSL 208
+Q QF L K I + V R+G+ Q++SI+D++VGD+++L GDQ+PADG+F+ G+SL
Sbjct: 824 QQSLQFLDLDKEKKKISIHVTRDGKRQKVSIYDLVVGDIVHLSTGDQVPADGIFIQGYSL 1003
Query: 209 QVDESSMTGESDHVEIEPLKAPFLLSGAKVVDGYAQMLVTAVGANTAWGQMMSSISGDNS 268
+DESS++GES+ V+I+ + PFLLSG KV DG A+M+VT VG T WG++M ++S
Sbjct: 1004 LIDESSLSGESEPVDIDN-RRPFLLSGTKVQDGQAKMIVTTVGMRTEWGKLMETLSEGGE 1180
Query: 269 ERTPLQARLDKLTSSIGKIGLAVAFLVLLVLLIRYFTGNTEDENGNKEYKGSKTDINDVC 328
+ TPLQ +L+ + + IGKIGL A L LVL R+ NG+ S+ +
Sbjct: 1181 DETPLQVKLNGVATVIGKIGLTFAVLTFLVLTARFVIEKAI--NGDFTSWSSEDALK--- 1345
Query: 329 NAVVSIVAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMMADQAMVRKLSACETMGSATVIC 388
++ A AVTI+VVAIPEGLPLAVTL+LA++MK++M D+A+VR LSACETMGSA+ IC
Sbjct: 1346 --LLDYFAIAVTIIVVAIPEGLPLAVTLSLAFAMKKLMNDRALVRHLSACETMGSASCIC 1519
Query: 389 TDKTGTLTLNQMRVTKFWLGLENV-------VENFSNAMAPTVLELFHQGVGLNTTGSVY 441
TDKTGTLT N M V K W+ + V + + ++ VL + Q + NT+ V
Sbjct: 1520 TDKTGTLTTNHMVVDKIWICEKTVEMKGDESTDKLKSEISDEVLSILLQAIFQNTSSEVV 1699
Query: 442 KPSAESEPEISGSPTEKAMLLWAVSDLGMDMDELKQKHKVLHVETFNSEKKRSGVAVRKE 501
K + E + I G+PTE A+L + + G D D ++ KVL VE FNS++K+ V V
Sbjct: 1700 KDN-EGKQTILGTPTESALLEFGLVS-GGDFDAQRRSCKVLKVEPFNSDRKKMSVLVGLP 1873
Query: 502 TNNTVHVHWKGAAEMVLAMCSNYIDSNGTQKSLDEERSKI-EKIIQGMAASSLRCIAFAY 560
V KGA+E+VL MC IDSNGT L EE+++I II G A +LR + A
Sbjct: 1874 DGG-VRAFCKGASEIVLKMCDKIIDSNGTTIDLPEEKARIVSDIIDGFANEALRTLCLAV 2050
Query: 561 MEISEGGDYIEKGKPRQVLREDGLTLLGIVGLKDPCRPNVKKAVETCKLAGVDIKMITGD 620
+I E +G+ + E+G TL+ IVG+KDP RP VK+AV+ C AG+ ++M+TGD
Sbjct: 2051 KDIDE-----TQGETN--IPENGYTLITIVGIKDPVRPGVKEAVQKCLAAGISVRMVTGD 2209
Query: 621 NIFTAKAIATECGILDLNDAGGVVVEGVEFRNYTEEERMEKVDKIRVMARSSPMDKLLMV 680
NI TAKAIA ECGIL GGV +EG EFRN +EE+ + + +I+VMARS P+DK +V
Sbjct: 2210 NINTAKAIAKECGILT---EGGVAIEGPEFRNLSEEQMKDIIPRIQVMARSLPLDKHTLV 2380
Query: 681 QCLKKK-GHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFNSVATV 739
L+ G VVAVTGDGTNDAPAL E+DIGL+MGI GTEVAKE++D++I+DDNF ++ V
Sbjct: 2381 TRLRNMFGEVVAVTGDGTNDAPALHESDIGLAMGIAGTEVAKENADVIIMDDNFTTIVKV 2560
Query: 740 LRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTTVQLLWVNLIMDTLGALA 799
+WGR +Y NIQKF+QFQLTVNV AL+ NF++A +G PLT VQLLWVNLIMDTLGALA
Sbjct: 2561 AKWGRAIYINIQKFVQFQLTVNVVALITNFVSACITGAAPLTAVQLLWVNLIMDTLGALA 2740
Query: 800 LATERPTKELMQKKPIGRTEPLITKIMWRNLLAQALYQIAVLLVFQFYGKSIFNV----S 855
LATE P LM+++P+GR ITK MWRN+ Q+LYQ+ VL V F GK + + S
Sbjct: 2741 LATEPPNDGLMERQPVGRKASFITKPMWRNIFGQSLYQLIVLGVLNFEGKRLLGLSGPDS 2920
Query: 856 KEVKNTLIFNTFVLCQVFNEFNSRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMVELLR 915
V NTLIFN+FV CQVFNE NSR +EK+N+F G+ + +FL ++ T V QV++VE L
Sbjct: 2921 TAVLNTLIFNSFVFCQVFNEINSREIEKINIFRGMFDSWIFLSVILATAVFQVIIVEFLG 3100
Query: 916 KFADTERLNWEQWGICIGIAAVSWPIAWLTKLTPV 950
FA T L W+ W + + +S P+A + K PV
Sbjct: 3101 TFASTVPLTWQFWLLSLLFGVLSMPLAAILKCIPV 3205
>TC84004 type IIB calcium ATPase [Medicago truncatula]
Length = 3655
Score = 785 bits (2028), Expect = 0.0
Identities = 440/939 (46%), Positives = 611/939 (64%), Gaps = 17/939 (1%)
Frame = +1
Query: 32 MVKDKNLEAYSEFGGVEGVADVLGTIPAKGILGSDDDTAARRELFGTNTYVRPPPKIFLH 91
+ + ++ + S GGVE VA L +G+ +D R+++FG N Y P + FL
Sbjct: 547 LFRSQDYKNLSNNGGVEAVARKLSVSIDEGV--NDTSVDCRQQIFGANRYTEKPSRTFLM 720
Query: 92 FVLEALNDTTILILLGCAGLSLGFGIKEHGPGEGWYEGGSIFLAVFLVVVVSALSNFRQD 151
FV +AL D T+ IL+ CA +S+G G+ G +G Y+G I L++FLVV+V+A+S++RQ
Sbjct: 721 FVWDALQDLTLTILMVCAVVSIGIGLATEGWPKGTYDGVGIILSIFLVVIVTAVSDYRQS 900
Query: 152 RQFDKLSKISNDIKVEVVRNGRPQQISIFDVLVGDVIYLKIGDQIPADGLFLGGHSLQVD 211
QF L + I V+V R+G+ ++ISI+DV+VGD+I+L GDQ+PADG+++ G+SL +D
Sbjct: 901 LQFMDLDREKKKIFVQVNRDGKRKKISIYDVVVGDIIHLSTGDQVPADGIYISGYSLLID 1080
Query: 212 ESSMTGESDHVEIEPLKAPFLLSGAKVVDGYAQMLVTAVGANTAWGQMMSSISGDNSERT 271
ESS++GES+ V I + PFLLSG KV DG +MLVT VG T WG++M +++ + T
Sbjct: 1081 ESSLSGESEPVFITE-EHPFLLSGTKVQDGQGKMLVTTVGMRTEWGKLMETLNEGGEDET 1257
Query: 272 PLQARLDKLTSSIGKIGLAVAFLVLLVLLIRYFTGNT-EDENGNKEYKGSKTDINDVCNA 330
PLQ +L+ + + IGKIGL A + LVL +R+ E GN ND
Sbjct: 1258 PLQVKLNGVATIIGKIGLFFAIVTFLVLTVRFLVEKALHGEFGN-------WSSNDATK- 1413
Query: 331 VVSIVAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMMADQAMVRKLSACETMGSATVICTD 390
++ A AVTI+VVA+PEGLPLAVTL+LA++MK++M D A+VR LSACETMGSA+ ICTD
Sbjct: 1414 LLDFFAIAVTIIVVAVPEGLPLAVTLSLAFAMKKLMNDMALVRHLSACETMGSASCICTD 1593
Query: 391 KTGTLTLNQMRVTKFWL--------GLENVVENFSNAMAPTVLELFHQGVGLNTTGSVYK 442
KTGTLT N M V K W+ G E+ E +N ++ VL + Q + NT+ V K
Sbjct: 1594 KTGTLTTNHMVVNKIWICENTTQLKGDESADELKTN-ISEGVLSILLQAIFQNTSAEVVK 1770
Query: 443 PSAESEPEISGSPTEKAMLLWAVSDLGMDMDELKQK--HKVLHVETFNSEKKRSGVAVRK 500
+ I GSPTE A+L + + LG + D +K+L +E FNS +K+ V V
Sbjct: 1771 DK-NGKNTILGSPTESALLEFGLL-LGSEFDARNHSKAYKILKLEPFNSVRKKMSVLVGL 1944
Query: 501 ETNNTVHVHWKGAAEMVLAMCSNYIDSNGTQKSLDEERSKI-EKIIQGMAASSLRCIAFA 559
N V KGA+E++L MC ID NG L +R+ I +I A+ +LR + A
Sbjct: 1945 P-NGRVQAFCKGASEIILEMCDKMIDCNGEVVDLPADRANIVSDVINSFASEALRTLCLA 2121
Query: 560 YMEISEGGDYIEKGKPRQVLREDGLTLLGIVGLKDPCRPNVKKAVETCKLAGVDIKMITG 619
+I+E +G+ + + G TL+ +VG+KDP RP VK+AV+TC AG+ ++M+TG
Sbjct: 2122 VRDINE-----TQGETN--IPDSGYTLIALVGIKDPVRPGVKEAVQTCIAAGITVRMVTG 2280
Query: 620 DNIFTAKAIATECGILDLNDAGGVVVEGVEFRNYTEEERMEKVDKIRVMARSSPMDKLLM 679
DNI TAKAIA ECGIL + GV +EG FR ++E+ + + +I+VMARS P+DK +
Sbjct: 2281 DNINTAKAIAKECGILTDD---GVAIEGPSFRELSDEQMKDIIPRIQVMARSLPLDKHKL 2451
Query: 680 VQCLKKK-GHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFNSVAT 738
V L+ G VVAVTGDGTNDAPAL EADIGL+MGI GTEVAKE +D++I+DDNF ++
Sbjct: 2452 VTNLRNMFGEVVAVTGDGTNDAPALHEADIGLAMGIAGTEVAKEKADVIIMDDNFATIVN 2631
Query: 739 VLRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTTVQLLWVNLIMDTLGAL 798
V++WGR VY NIQKF+QFQLTVNV AL+INF++A +G PLT VQLLWVNLIMDTLGAL
Sbjct: 2632 VVKWGRAVYINIQKFVQFQLTVNVVALIINFVSACITGSAPLTAVQLLWVNLIMDTLGAL 2811
Query: 799 ALATERPTKELMQKKPIGRTEPLITKIMWRNLLAQALYQIAVLLVFQFYGKSIFNV---- 854
ALATE P L+++ P+GR ITK MWRN++ Q++YQ+ VL + F GK + +
Sbjct: 2812 ALATEPPNDGLLKRPPVGRGASFITKTMWRNIIGQSIYQLIVLAILNFDGKRLLGINGSD 2991
Query: 855 SKEVKNTLIFNTFVLCQVFNEFNSRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMVELL 914
+ EV NTLIFN+FV CQVFNE NSR +EK+N+F G+ + +FL I+ T+ QV++VE L
Sbjct: 2992 ATEVLNTLIFNSFVFCQVFNEINSRDIEKINIFRGMFDSWIFLLIIFSTVAFQVVIVEFL 3171
Query: 915 RKFADTERLNWEQWGICIGIAAVSWPIAWLTKLTPVPSK 953
FA T L+W+ W + + I A+S P+A + K PV K
Sbjct: 3172 GAFASTVPLSWQLWLLSVLIGAISMPLAVIVKCIPVERK 3288
>TC85483 type IIB calcium ATPase MCA5 [Medicago truncatula]
Length = 2541
Score = 634 bits (1634), Expect = 0.0
Identities = 354/721 (49%), Positives = 475/721 (65%), Gaps = 11/721 (1%)
Frame = +2
Query: 241 GYAQMLVTAVGANTAWGQMMSSISGDNSERTPLQARLDKLTSSIGKIGLAVAFLVLLVLL 300
G +MLVT VG T WG++M+++S + TPLQ +L+ + + IGKIGL A + VL+
Sbjct: 2 GSCKMLVTTVGMRTQWGKLMATLSEGGDDETPLQVKLNGVATIIGKIGLFFAIVTFAVLV 181
Query: 301 IRYFTGNTEDENGNKEYKGSKTDINDVCNAVVSIVAAAVTIVVVAIPEGLPLAVTLTLAY 360
+ + EN + G D ++ A AVTIVVVA+PEGLPLAVTL+LA+
Sbjct: 182 QGLVSLKLQQENF-WNWNG------DDALEMLEYFAIAVTIVVVAVPEGLPLAVTLSLAF 340
Query: 361 SMKRMMADQAMVRKLSACETMGSATVICTDKTGTLTLNQMRVTKFWLGLE-----NVVEN 415
+MK+MM D+A+VR L+ACETMGSAT IC+DKTGTLT N M V K + ++ N +
Sbjct: 341 AMKKMMNDKALVRNLAACETMGSATTICSDKTGTLTTNHMTVVKTCICMKSKEVSNKTSS 520
Query: 416 FSNAMAPTVLELFHQGVGLNTTGSVYKPSAESEPEISGSPTEKAMLLWAVSDLGMDMDEL 475
+ + +V++L Q + NT G V + + + EI G+PTE A+L + +S LG D
Sbjct: 521 LCSELPESVVKLLQQSIFNNTGGEVVV-NKQGKHEILGTPTETAILEFGLS-LGGDFQGE 694
Query: 476 KQKHKVLHVETFNSEKKRSGVAVRKETNNTVHVHWKGAAEMVLAMCSNYIDSNGTQKSLD 535
+Q K++ VE FNS KKR G V + + H KGA+E+VLA C ++SNG LD
Sbjct: 695 RQACKLVKVEPFNSTKKRMGAVVELPSGG-LRAHCKGASEIVLAACDKVLNSNGEVVPLD 871
Query: 536 EERSK-IEKIIQGMAASSLRCIAFAYMEISEGGDYIEKGKPRQVLREDGLTLLGIVGLKD 594
EE + + I A +LR + AYME+ G + G T +G+VG+KD
Sbjct: 872 EESTNHLTNTINQFANEALRTLCLAYMELENGFS------AEDTIPVTGYTCIGVVGIKD 1033
Query: 595 PCRPNVKKAVETCKLAGVDIKMITGDNIFTAKAIATECGILDLNDAGGVVVEGVEFRNYT 654
P RP VK++V C+ AG+ ++M+TGDNI TAKAIA ECGIL + G+ +EG EFR +
Sbjct: 1034PVRPGVKESVALCRSAGITVRMVTGDNINTAKAIARECGILTDD---GIAIEGPEFREKS 1204
Query: 655 EEERMEKVDKIRVMARSSPMDKLLMVQCLKKK-GHVVAVTGDGTNDAPALKEADIGLSMG 713
EE +E + KI+VMARSSP+DK +V+ L+ G VVAVTGDGTNDAPAL EADIGL+MG
Sbjct: 1205LEELLELIPKIQVMARSSPLDKHTLVRHLRTTFGEVVAVTGDGTNDAPALHEADIGLAMG 1384
Query: 714 IQGTEVAKESSDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAV 773
I GTEVAKES+D++ILDDNF+++ TV +WGR VY NIQKF+QFQLTVN+ AL++NF +A
Sbjct: 1385IAGTEVAKESADVIILDDNFSTIVTVAKWGRSVYINIQKFVQFQLTVNIVALIVNFTSAC 1564
Query: 774 SSGDVPLTTVQLLWVNLIMDTLGALALATERPTKELMQKKPIGRTEPLITKIMWRNLLAQ 833
+G PLT VQLLWVN+IMDTLGALALATE P +LM++ P+GR I+ +MWRN+L Q
Sbjct: 1565LTGTAPLTAVQLLWVNMIMDTLGALALATEPPNDDLMKRAPVGRKGNFISNVMWRNILGQ 1744
Query: 834 ALYQIAVLLVFQFYGKSIFNV----SKEVKNTLIFNTFVLCQVFNEFNSRSMEKLNVFEG 889
+LYQ V+ Q GK+IF++ S V NTLIFN FV CQVFNE NSR MEK+NVF+G
Sbjct: 1745SLYQFMVIWFLQSKGKTIFSLDGPNSDLVLNTLIFNAFVFCQVFNEINSREMEKINVFKG 1924
Query: 890 ILKNHLFLGIVGITIVLQVLMVELLRKFADTERLNWEQWGICIGIAAVSWPIAWLTKLTP 949
IL N++F+G++ TI Q+++VE L FA+T L QW C+ + + PIA K P
Sbjct: 1925ILDNYVFVGVISATIFFQIIIVEYLGTFANTTPLTLVQWFFCLFVGFMGMPIAARLKKIP 2104
Query: 950 V 950
V
Sbjct: 2105V 2107
>TC88944 type IIB calcium ATPase [Medicago truncatula]
Length = 1883
Score = 560 bits (1443), Expect = e-159
Identities = 306/618 (49%), Positives = 423/618 (67%), Gaps = 13/618 (2%)
Frame = +3
Query: 348 EGLPLAVTLTLAYSMKRMMADQAMVRKLSACETMGSATVICTDKTGTLTLNQMRVTKFWL 407
+GLPLAVTL LA++MK+MM D+A+VR L+ACETMGSAT IC+DKTGTLT N M V K +
Sbjct: 3 DGLPLAVTLILAFAMKKMMNDKALVRHLAACETMGSATTICSDKTGTLTTNHMTVVKACI 182
Query: 408 -----GLENVVEN--FSNAMAPTVLELFHQGVGLNTTGSVYKPSAESEPEISGSPTEKAM 460
+ + +++ FS+ + + + + + + NT G V K + + EI GSPTE A+
Sbjct: 183 CGKIKEVNSSIDSSDFSSDLPDSAIAILLESIFNNTGGEVVK-NENGKIEILGSPTETAI 359
Query: 461 LLWAVSDLGMDMDELKQKHKVLHVETFNSEKKRSGVAVRKETNNTVHVHWKGAAEMVLAM 520
L + +S LG D + +Q K++ VE FNS KKR GV ++ H KGA+E++LA
Sbjct: 360 LEFGLS-LGGDFHKERQASKLVKVEPFNSIKKRMGVVLQLPDGG-YRAHCKGASEIILAA 533
Query: 521 CSNYIDSNGTQKSLDEER-SKIEKIIQGMAASSLRCIAFAYMEISEGGDYIEKGKPRQVL 579
C ++DSN LDE+ S + I+ A +LR + AY++I D G P V
Sbjct: 534 CDKFVDSNSKIVPLDEDSISHLNDTIEKFANEALRTLCLAYIDIH---DEFLVGSPIPV- 701
Query: 580 REDGLTLLGIVGLKDPCRPNVKKAVETCKLAGVDIKMITGDNIFTAKAIATECGILDLND 639
+G T +GIVG+KDP RP V+++V C+ AG+ ++M+TGDNI TAKAIA ECGIL
Sbjct: 702 --NGYTCVGIVGIKDPVRPGVRESVAICRSAGITVRMVTGDNINTAKAIARECGIL---- 863
Query: 640 AGGVVVEGVEFRNYTEEERMEKVDKIRVMARSSPMDKLLMVQCLKKK-GHVVAVTGDGTN 698
G+ +EG EFR +E+E ++ + KI+VMARSSPMDK +V+ L+ VVAVTGDGTN
Sbjct: 864 TDGIAIEGPEFREMSEKELLDIIPKIQVMARSSPMDKHTLVKHLRTTFEEVVAVTGDGTN 1043
Query: 699 DAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQL 758
DAPAL EADIGL+MGI GTEVAKES+D++ILDDNF+++ TV +WGR VY NIQKF+QFQL
Sbjct: 1044DAPALHEADIGLAMGIAGTEVAKESADVIILDDNFSTIVTVAKWGRSVYINIQKFVQFQL 1223
Query: 759 TVNVAALVINFIAAVSSGDVPLTTVQLLWVNLIMDTLGALALATERPTKELMQKKPIGRT 818
TVNV AL++NF +A +G+ PLT VQLLWVN+IMDTLGALALATE P ELM++ P+GR
Sbjct: 1224TVNVVALIVNFTSACLTGNAPLTAVQLLWVNMIMDTLGALALATEPPNDELMKRAPVGRK 1403
Query: 819 EPLITKIMWRNLLAQALYQIAVLLVFQFYGKSIFNV----SKEVKNTLIFNTFVLCQVFN 874
I+ +MWRN+ Q++YQ ++ + Q GK++F++ S + NTLIFN+FV CQVFN
Sbjct: 1404GNFISNVMWRNITGQSIYQFVIIWLLQTRGKTVFHLDGPDSDLILNTLIFNSFVFCQVFN 1583
Query: 875 EFNSRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMVELLRKFADTERLNWEQWGICIGI 934
E +SR ME++NVFEGILKN++F ++ T + Q+++VE L +A+T L+ + W I + +
Sbjct: 1584EISSRDMERINVFEGILKNYVFTAVLTCTAIFQIIIVEFLGTYANTSPLSLKLWLISVFL 1763
Query: 935 AAVSWPIAWLTKLTPVPS 952
+ PI K+ PV S
Sbjct: 1764GVLGMPIGAALKMIPVGS 1817
>TC88464 similar to SP|Q9LU41|ACA9_ARATH Potential calcium-transporting
ATPase 9 plasma membrane-type (EC 3.6.3.8) (Ca2+-ATPase
isoform 9)., partial (43%)
Length = 1608
Score = 541 bits (1395), Expect = e-154
Identities = 281/457 (61%), Positives = 339/457 (73%), Gaps = 11/457 (2%)
Frame = +3
Query: 506 VHVHWKGAAEMVLAMCSNYIDSNGTQKSLDEERSKIEKIIQGMAASSLRCIAFAYMEISE 565
VH+HWKGAAE+VL C+ Y+DSN +S+++E+ +++ I MAA SLRC+A AY
Sbjct: 15 VHIHWKGAAEIVLGACTQYLDSNSHLQSIEQEKDFLKEAIDDMAARSLRCVAIAYRSYEL 194
Query: 566 GGDYI---EKGKPRQVLREDGLTLLGIVGLKDPCRPNVKKAVETCKLAGVDIKMITGDNI 622
D I E+ + L ED L LL IVG+KDPCRP VK AV C AGV ++M+TGDN+
Sbjct: 195 --DKIPSNEEDLAQWSLPEDELVLLAIVGIKDPCRPGVKDAVRICTDAGVKVRMVTGDNL 368
Query: 623 FTAKAIATECGILDLNDAG--GVVVEGVEFRNYTEEERMEKVDKIRVMARSSPMDKLLMV 680
TAKAIA ECGIL N+ ++EG FR +E+ER + KI VM RSSP DKLL+V
Sbjct: 369 QTAKAIALECGILASNEEAVDPCIIEGKVFRELSEKEREQVAKKITVMGRSSPNDKLLLV 548
Query: 681 QCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFNSVATVL 740
Q L+K G VVAVTGDGTNDAPAL EADIGLSMGIQGTEVAKESSDI+ILDDNF SV V+
Sbjct: 549 QALRKGGDVVAVTGDGTNDAPALHEADIGLSMGIQGTEVAKESSDIIILDDNFASVVKVV 728
Query: 741 RWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTTVQLLWVNLIMDTLGALAL 800
RWGR VY NIQKFIQFQLTVNVAALVIN +AA+SSGDVPL VQLLWVNLIMDTLGALAL
Sbjct: 729 RWGRSVYANIQKFIQFQLTVNVAALVINVVAAISSGDVPLNAVQLLWVNLIMDTLGALAL 908
Query: 801 ATERPTKELMQKKPIGRTEPLITKIMWRNLLAQALYQIAVLLVFQFYGKSIFNVSK---- 856
ATE PT LM + P+GR EPLIT IMWRNL+ QALYQ++VLL F G+SI
Sbjct: 909 ATEPPTDHLMHRSPVGRREPLITNIMWRNLIVQALYQVSVLLFLNFCGESILPKQDTKAH 1088
Query: 857 --EVKNTLIFNTFVLCQVFNEFNSRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMVELL 914
+VKNTLIFN FV+CQ+FNEFN+R +++NVF G+ KN LF+GI+GIT +LQ++++E L
Sbjct: 1089DYQVKNTLIFNAFVMCQIFNEFNARKPDEMNVFPGVTKNRLFMGIIGITFILQIVIIEFL 1268
Query: 915 RKFADTERLNWEQWGICIGIAAVSWPIAWLTKLTPVP 951
KFA T RL+W+ W + + I VSWP+A K PVP
Sbjct: 1269GKFASTVRLDWKLWLVSLIIGLVSWPLAIAGKFIPVP 1379
>TC87649 similar to SP|Q9SZR1|ACAA_ARATH Potential calcium-transporting
ATPase 10 plasma membrane-type (EC 3.6.3.8), partial
(43%)
Length = 2159
Score = 400 bits (1028), Expect(3) = e-152
Identities = 216/342 (63%), Positives = 257/342 (74%), Gaps = 6/342 (1%)
Frame = +3
Query: 491 KKRSGVAVRKETNNTVHVHWKGAAEMVLAMCSNYIDSNGTQKSLDEER-SKIEKIIQGMA 549
K+ +GVA++ ++ VH+HWKGAAE+VLA C+ YID+N +DEE+ + +K I+ MA
Sbjct: 63 KREAGVAIQT-ADSDVHIHWKGAAEIVLACCTGYIDANDQLVEIDEEKMTFFKKAIEDMA 239
Query: 550 ASSLRCIAFAYMEI-SEGGDYIEKGKPRQVLREDGLTLLGIVGLKDPCRPNVKKAVETCK 608
+ SLRC+A AY E E+ L E+ L LL IVG+KDPCRP VK +V+ C+
Sbjct: 240 SDSLRCVAIAYRPYEKEKVPDNEEQLADWSLPEEELVLLAIVGIKDPCRPGVKNSVQLCQ 419
Query: 609 LAGVDIKMITGDNIFTAKAIATECGIL----DLNDAGGVVVEGVEFRNYTEEERMEKVDK 664
AGV +KM+TGDN+ TAKAIA ECGIL D+ + V+EG FR ++ ER E +
Sbjct: 420 KAGVKVKMVTGDNVKTAKAIALECGILSSLADVTERS--VIEGKTFRALSDSEREEIAES 593
Query: 665 IRVMARSSPMDKLLMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESS 724
I VM RSSP DKLL+VQ L++KGHVVAVTGDGTNDAPAL EADIGL+MGI GTEVAKESS
Sbjct: 594 ISVMGRSSPNDKLLLVQALRRKGHVVAVTGDGTNDAPALHEADIGLAMGIAGTEVAKESS 773
Query: 725 DIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTTVQ 784
DI+ILDDNF SV V+RWGR VY NIQKFIQFQLTVNVAALVIN +AAVSSGDVPL VQ
Sbjct: 774 DIIILDDNFASVVKVVRWGRSVYANIQKFIQFQLTVNVAALVINVVAAVSSGDVPLNAVQ 953
Query: 785 LLWVNLIMDTLGALALATERPTKELMQKKPIGRTEPLITKIM 826
LLWVNLIMDTLGALALATE PT LM + P+GR EPLIT IM
Sbjct: 954 LLWVNLIMDTLGALALATEPPTDHLMDRSPVGRREPLITNIM 1079
Score = 155 bits (392), Expect(3) = e-152
Identities = 72/135 (53%), Positives = 98/135 (72%), Gaps = 7/135 (5%)
Frame = +1
Query: 825 IMWRNLLAQALYQIAVLLVFQFYGKSIFNVSKE-------VKNTLIFNTFVLCQVFNEFN 877
+ WRNLL QA+YQ++VLLV F G SI + + VKNTLIFN FV+CQ+FNEFN
Sbjct: 1075 LWWRNLLIQAMYQVSVLLVLNFRGISILGLEHQPTEHAIKVKNTLIFNAFVICQIFNEFN 1254
Query: 878 SRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMVELLRKFADTERLNWEQWGICIGIAAV 937
+R ++ N+F+G+ +N+LF+GIVG T+VLQV++VE L KF T RLNW+QW I + I +
Sbjct: 1255 ARKPDEYNIFKGVTRNYLFMGIVGFTVVLQVIIVEFLGKFTTTTRLNWKQWLISVAIGFI 1434
Query: 938 SWPIAWLTKLTPVPS 952
WP+A + KL PVP+
Sbjct: 1435 GWPLAVVGKLIPVPA 1479
Score = 25.0 bits (53), Expect(3) = e-152
Identities = 10/18 (55%), Positives = 13/18 (71%)
Frame = +2
Query: 476 KQKHKVLHVETFNSEKKR 493
+ + +LHV FNSEKKR
Sbjct: 17 RSESSILHVFPFNSEKKR 70
>TC92182 type IIA calcium ATPase [Medicago truncatula]
Length = 3148
Score = 320 bits (821), Expect = 1e-87
Identities = 265/880 (30%), Positives = 440/880 (49%), Gaps = 80/880 (9%)
Frame = +1
Query: 33 VKDKNLEAYSEFGGVEGVADVLGTIPAKGILGSDDDTAARRELFGTNTYVRPPPKIFLHF 92
+++K A+S V+ + G KG+ S ++ RRE +G N + K
Sbjct: 1 MEEKPFPAWS--WSVDECLEEYGVKLEKGL--SSNEVQKRREKYGWNELAKEKGKPLWKL 168
Query: 93 VLEALNDTTILILLGCAGLSLGFGIKEHGPGEGWYEGGSIFLAVFLVVVVSALSNFRQDR 152
VLE +D + ILL A +S F + E +E L + L++V++A+ Q+
Sbjct: 169 VLEQFDDMLVKILLAAAFIS--FLLAYFEGSESGFEAYVEPLVIILILVLNAIVGVWQEN 342
Query: 153 QFDKLSKISNDIKVE---VVRNGR-PQQISIFDVLVGDVIYLKIGDQIPADGLF--LGGH 206
+K + +++ E V+R+G + +++ GD++ L++GD++PAD L
Sbjct: 343 NAEKALEALKELQCESIKVLRDGYFVPDLPARELVPGDIVELRVGDKVPADMRVAALKTS 522
Query: 207 SLQVDESSMTGES------------DHVEIEPLKAPFLLSGAKVVDGYAQMLVTAVGANT 254
+L++++SS+TGE+ D E++ K + +G VV+G +V NT
Sbjct: 523 TLRLEQSSLTGEAMPVLKGTNPIFMDDCELQA-KENMVFAGTTVVNGSCICIVITTAMNT 699
Query: 255 AWGQMMSSISGDNSER--TPLQARLDKLTSSIGKIGLAVAFLVLLVLLIRYFTGNTEDEN 312
G++ I + E TPL+ +LD+ G++ ++ + L+V +I Y + D
Sbjct: 700 EIGKIQKQIHEASLEESDTPLKKKLDEFG---GRLTTSIGIVCLVVWIINYKNFISWDV- 867
Query: 313 GNKEYKGSKTDINDVCNAVVSIVAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMMADQAMV 372
G T+I AV + V AIPEGLP +T LA ++M A+V
Sbjct: 868 ----VDGWPTNIQFSFQKCTYYFKIAVALAVAAIPEGLPAVITTCLALGTRKMAQKNAIV 1035
Query: 373 RKLSACETMGSATVICTDKTGTLTLNQMRVTKFW-----------LGLEN---------V 412
RKL + ET+G TVIC+DKTGTLT NQM T+F+ + +E +
Sbjct: 1036RKLPSVETLGCTTVICSDKTGTLTTNQMSATEFFTLGGKTTACRVISVEGTTYDPKDGGI 1215
Query: 413 VENFSNAMAPTVLEL------------FHQGVGLNTTGSVYKPSAESEPEISGSPTEKAM 460
V+ M +L + + G TG + + + E G P K+
Sbjct: 1216VDWTCYNMDANLLAMAEICAVCNDAGVYFDGRLFRATGLPTEAALKVLVEKMGFPDTKSR 1395
Query: 461 -----LLWAVSDLGMDMDELK--------QKHKVLHVETFNSEKKRSGVAVRKETNNTVH 507
L A +++ +D + LK ++ K + F+ +K V VR E +
Sbjct: 1396NKTHDALVATNNM-VDCNTLKLGCCEWWNRRSKRVATLEFDRVRKSMSVIVR-EPDGQNR 1569
Query: 508 VHWKGAAEMVLAMCSNYIDSNGTQKSLDEE-RSKIEKIIQGMAASSLRCIAFAYM-EISE 565
+ KGA E +L S ++G+ +D++ R + + + M++ LRC+ A E+ E
Sbjct: 1570LLVKGAVESLLERSSYVQLADGSLVPIDDQCRELLLQRLHEMSSKGLRCLGLACKDELGE 1749
Query: 566 GGDYIEKGKP--RQVLR-------EDGLTLLGIVGLKDPCRPNVKKAVETCKLAGVDIKM 616
DY P +++L E L +G+VGL+DP R V KA+E CK AG+ + +
Sbjct: 1750FSDYYADTHPAHKKLLDPTYYSSIESDLIFVGVVGLRDPPREEVHKAIEDCKQAGIRVMV 1929
Query: 617 ITGDNIFTAKAIATECGILDLN-DAGGVVVEGVEFRNYTEEERMEKVDKI--RVMARSSP 673
ITGDN TA+AI E + + D G + G EF + + E+++ + + +V +R+ P
Sbjct: 1930ITGDNKSTAEAICKEIKLFSTDEDLTGQSLTGKEFMSLSHSEQVKLLLRNGGKVFSRAEP 2109
Query: 674 MDKLLMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNF 733
K +V+ LK+ G +VA+TGDG NDAPALK ADIG++MGI GTEVAKE+SD+V+ DDNF
Sbjct: 2110RHKQEIVRLLKEMGEIVAMTGDGVNDAPALKLADIGIAMGITGTEVAKEASDMVLADDNF 2289
Query: 734 NSVATVLRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTTVQLLWVNLIMD 793
+++ + + GR +YNN++ FI++ ++ NV ++ F+ A + VQLLWVNL+ D
Sbjct: 2290STIVSAIAEGRAIYNNMKAFIRYMISSNVGEVISIFLTAALGIPECMIPVQLLWVNLVTD 2469
Query: 794 TLGALALATERPTKELMQKKPIGRTEPLITK-IMWRNLLA 832
A AL ++MQK P + LI+ +++R L++
Sbjct: 2470GPPATALGFNPADVDIMQKPPRKSDDALISAWVLFRYLVS 2589
>CB894102 similar to GP|8843813|db Ca2+-transporting ATPase-like protein
{Arabidopsis thaliana}, partial (15%)
Length = 802
Score = 189 bits (479), Expect = 6e-48
Identities = 105/240 (43%), Positives = 147/240 (60%), Gaps = 31/240 (12%)
Frame = +2
Query: 135 AVFLVVVVSALSNFRQDRQFDKLSKISNDIKVEVVRNGRPQQISIFDVLVGDVIYLKIGD 194
AV LV+VV+A+S+++Q QF L++ +I +EV+R GR +ISI+D++VGDVI L IG+
Sbjct: 5 AVILVIVVTAVSDYKQSLQFRDLNEEKRNIHLEVIRGGRRVEISIYDLVVGDVIPLNIGN 184
Query: 195 QIPADGLFLGGHSLQVDESSMTGESDHVEIEPLKAPFLLSGAKVVDGYAQMLVTAVGANT 254
Q+PADG+ + GHSL +DESSMTGES V + K PF++SG KV DG MLVT VG NT
Sbjct: 185 QVPADGVVITGHSLSIDESSMTGESKIVH-KDSKDPFMMSGCKVADGSGTMLVTGVGINT 361
Query: 255 AWGQMMSSISGDNSERTPLQARLDKLTSSIGKIGLAVAFLVLLVLL-------------- 300
WG +M+SIS D E TPLQ RL+ + + IG +GL+VA LVL+VLL
Sbjct: 362 EWGLLMASISEDTGEETPLQVRLNGVATFIGIVGLSVAVLVLIVLLAR*LVDGPCLPYFS 541
Query: 301 -----------------IRYFTGNTEDENGNKEYKGSKTDINDVCNAVVSIVAAAVTIVV 343
IRYF+G+T + +G K++ KT + + I+ AV ++
Sbjct: 542 *RESSMLFCMLLKWELVIRYFSGHTRNSDGTKQFIAGKTKAGHAIDGAIKIITVAVGTLI 721
>TC78163 H+-ATPase
Length = 3295
Score = 170 bits (431), Expect = 2e-42
Identities = 165/738 (22%), Positives = 332/738 (44%), Gaps = 11/738 (1%)
Frame = +1
Query: 67 DDTAARRELFGTNTYV-RPPPKI--FLHFVLEALNDTTILILLGCAGLSLGFGIKEHGPG 123
++ R ELFG N + KI FL F+ L+ + ++ G +
Sbjct: 244 EEVQERLELFGYNKLEEKKESKILKFLGFMWNPLSWVMEAAAIMAIAMAHGGRNMDGTKK 423
Query: 124 EGWYEGGSIFLAVFLVVVVSALSNFRQDRQFDK-----LSKISNDIKVEVVRNGRPQQIS 178
+G Y+ F+ + +++++++ +F ++ +++++ K +V+R+G+ +
Sbjct: 424 QGDYQD---FVGIIILLIINSTISFIEENNAGNAAAALMARLAP--KAKVLRDGKWSEED 588
Query: 179 IFDVLVGDVIYLKIGDQIPADGLFLGGHSLQVDESSMTGESDHVEIEPLKAPFLLSGAKV 238
++ GD++ +K+GD IPAD L G L++D+S++TGES V P + + SG+
Sbjct: 589 ASVLVPGDIVSIKLGDIIPADARLLEGDPLKIDQSALTGESLPVTKHPGEG--IYSGSTC 762
Query: 239 VDGYAQMLVTAVGANTAWGQMMSSISGDNSERTPLQARLDKLTSSIGKIGL-AVAFLVLL 297
G + +V A G +T +G+ + E T K+ +SIG + ++A +++
Sbjct: 763 KQGEIEAIVIATGVHTFFGKAAHLV-----ENTTHVGHFQKVLTSIGNFCICSIAIGMVI 927
Query: 298 VLLIRYFTGNTEDENGNKEYKGSKTDINDVCNAVVSIVAAAVTIVVVAIPEGLPLAVTLT 357
+++ Y KG + I+++ + +++ IP +P +++T
Sbjct: 928 EIIVIYGVHG----------KGYRNGIDNL-----------LVLLIGGIPIAMPTVLSVT 1044
Query: 358 LAYSMKRMMADQAMVRKLSACETMGSATVICTDKTGTLTLNQMRVTKFWLGLENVVENFS 417
+A ++ A+ ++++A E M V+C+DKTGTLTLN++ V K
Sbjct: 1045MAIGSHKLSQQGAITKRMTAIEEMAGMDVLCSDKTGTLTLNKLTVDK------------- 1185
Query: 418 NAMAPTVLELFHQGVGLNTTGSVYKPSAESEPEISGSPTEKAMLLWAVSDLGMDMDELKQ 477
++E+F +GV + + ++ E + + A+ + D E +
Sbjct: 1186-----DMIEVFAKGVDKDLVVLMAARASRLE--------NQDAIDCAIVSMLADPKEART 1326
Query: 478 KHKVLHVETFNSEKKRSGVAVRKETNNTVHVHWKGAAEMVLAMCSNYIDSNGTQKSLDEE 537
K +H FN KR+ + N +H KGA E +L + N E
Sbjct: 1327GIKEVHFLPFNPTDKRTALTYIDAAGN-MHRVSKGAPEQILNLARNKA----------EI 1473
Query: 538 RSKIEKIIQGMAASSLRCIAFAYMEISEGGDYIEKGKPRQVLREDGLTLLGIVGLKDPCR 597
K+ +I A LR + A E+ EG G P + + ++ L DP R
Sbjct: 1474AQKVHSMIDKFAERGLRSLGVARQEVPEGSK-DSPGGPWE--------FVALLPLFDPPR 1626
Query: 598 PNVKKAVETCKLAGVDIKMITGDNIFTAKAIATECGI-LDLNDAGGVVVEGV-EFRNYTE 655
+ + + GV +KMITGD + K G+ ++ + ++ + + +
Sbjct: 1627HDSAETIRRALDLGVSVKMITGDQLAIGKETGRRLGMGTNMYPSSSLLGDNKDQLGAVSI 1806
Query: 656 EERMEKVDKIRVMARSSPMDKLLMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQ 715
++ +E D A P K +V+ L+ + H+ +TGDG NDAPALK ADIG+++
Sbjct: 1807DDLIENADG---FAGVFPEHKYEIVKRLQARKHICGMTGDGVNDAPALKIADIGIAVA-D 1974
Query: 716 GTEVAKESSDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSS 775
T+ A+ +SDIV+ + + + + + R ++ ++ + + +++ + +V+ F+ S
Sbjct: 1975STDAARSASDIVLTEPGLSVIISAVLTSRAIFQRMKNYTIYAVSITI-RIVLGFMLLNSF 2151
Query: 776 GDVPLTTVQLLWVNLIMD 793
+L + ++ D
Sbjct: 2152WSFDFPPFMVLIIAILND 2205
>TC77796 H+-ATPase
Length = 3416
Score = 110 bits (275), Expect(2) = 2e-41
Identities = 97/380 (25%), Positives = 170/380 (44%), Gaps = 3/380 (0%)
Frame = +1
Query: 386 VICTDKTGTLTLNQMRVTKFWLGLENVVENFSNAMAPTVLELFHQGVGLNTTGSVYKPSA 445
V+C+DKTGTLTLN++ V K N++E F+ + + L +
Sbjct: 1072 VLCSDKTGTLTLNKLSVEK------NLIEVFAKGVEKDYVILL----------AARASRT 1203
Query: 446 ESEPEISGSPTEKAMLLWAVSDLGMDMDELKQKHKVLHVETFNSEKKRSGVAVRKETNNT 505
E++ I A+ + D E + + +H FN KR+ + N
Sbjct: 1204 ENQDAIDA----------AIVGMLADPKEARAGVREVHFFPFNPVDKRTALTYIDADGNW 1353
Query: 506 VHVHWKGAAEMVLAMCSNYIDSNGTQKSLDEERSKIEKIIQGMAASSLRCIAFAYMEISE 565
H KGA E +L +C+ ++ R K I A LR + A EI E
Sbjct: 1354 -HRSSKGAPEQILNLCN----------CKEDVRKKAHSTIDKFAERGLRSLGVARQEIPE 1500
Query: 566 GGDYIEKGKPRQVLREDGLTLLGIVGLKDPCRPNVKKAVETCKLAGVDIKMITGDNIFTA 625
D G P Q +G++ L DP R + + + GV++KMITGD + A
Sbjct: 1501 K-DKDSPGAPWQ--------FVGLLPLFDPPRHDSAETITRALNLGVNVKMITGDQLAIA 1653
Query: 626 KAIATECGI-LDLNDAGGVVVEGVE--FRNYTEEERMEKVDKIRVMARSSPMDKLLMVQC 682
K G+ ++ + ++ + + +E +EK D A P K +V+
Sbjct: 1654 KETGRRLGMGTNMYPSSSLLGQSKDAAVSALPVDELIEKADGF---AGVFPEHKYEIVKK 1824
Query: 683 LKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFNSVATVLRW 742
L+++ H+ +TGDG NDAPALK ADIG+++ T+ A+ +SDIV+ + + + + +
Sbjct: 1825 LQERKHICGMTGDGVNDAPALKRADIGIAVA-DATDAARGASDIVLTEPGLSVIISAVLT 2001
Query: 743 GRCVYNNIQKFIQFQLTVNV 762
R ++ ++ + + +++ +
Sbjct: 2002 SRAIFQRMKNYTIYAVSITI 2061
Score = 78.6 bits (192), Expect(2) = 2e-41
Identities = 74/321 (23%), Positives = 143/321 (44%), Gaps = 4/321 (1%)
Frame = +3
Query: 65 SDDDTAARRELFGTNTYVRPPPKIFLHFVLEALNDTT-ILILLGCAGLSLGFGIKEHGPG 123
+ D+ A R ++FG N FL F+ N + ++ ++L G G
Sbjct: 198 TSDEGANRLQVFGPNKLEEKRESKFLKFLGFMWNPLSWVMEAAAIMAIALANG---SGRP 368
Query: 124 EGWYEGGSIFLAVFLVVVVSALSNFRQDRQFDKLSKISNDI--KVEVVRNGRPQQISIFD 181
W + I + L+V+ S +S ++ + + + + K V+R+GR +
Sbjct: 369 PDWQDFVGI---ISLLVINSTISFIEENNAGNAAAALMAGLAPKTRVLRDGRWSEEDAAI 539
Query: 182 VLVGDVIYLKIGDQIPADGLFLGGHSLQVDESSMTGESDHVEIEPLKAPFLLSGAKVVDG 241
++ GD+I +K+GD IPAD L G +L VD+S++TGES P F SG+ V G
Sbjct: 540 LVPGDIISIKLGDIIPADARLLEGDALSVDQSALTGESLPATKNPSDEVF--SGSTVKKG 713
Query: 242 YAQMLVTAVGANTAWGQMMSSISGDNSERTPLQARLDKLTSSIGKIGL-AVAFLVLLVLL 300
+ +V A G +T +G+ + N K+ ++IG + ++A +L+ L+
Sbjct: 714 EIEAVVIATGVHTFFGKAAHLVDSTNQ-----VGHFQKVLTAIGNFCICSIAVGILIELV 878
Query: 301 IRYFTGNTEDENGNKEYKGSKTDINDVCNAVVSIVAAAVTIVVVAIPEGLPLAVTLTLAY 360
+ Y + + +G + + +++ IP +P +++T+A
Sbjct: 879 VMYPIQHRKYRDG---------------------IDNLLVLLIGGIPIAMPTVLSVTMAI 995
Query: 361 SMKRMMADQAMVRKLSACETM 381
R+ A+ ++++A E M
Sbjct: 996 GSHRLSQQGAITKRMTAIEEM 1058
>TC79651 similar to PIR|T52581|T52581 Ca2+-transporting ATPase (EC 3.6.1.38)
ECA3 [imported] - Arabidopsis thaliana, partial (38%)
Length = 1546
Score = 147 bits (371), Expect = 2e-35
Identities = 104/342 (30%), Positives = 169/342 (49%), Gaps = 32/342 (9%)
Frame = +3
Query: 619 GDNIFTAKAIATECGILD-LNDAGGVVVEGVEFRNYTEEERMEKVDKIRVMARSSPMDKL 677
G+N TA+++ + G D L D EF ++ + ++ + R P K
Sbjct: 18 GNNKSTAESLCRKIGAFDHLIDFTEHSYTASEFEELPALQQTIALQRMALFTRVEPSHKR 197
Query: 678 LMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFNSVA 737
++V+ L+ + VVA+TGDG NDAPALK+ADIG++MG GT VAK +SD+V+ DDNF S+
Sbjct: 198 MLVEALQHQNEVVAMTGDGVNDAPALKKADIGIAMG-SGTAVAKSASDMVLADDNFASIV 374
Query: 738 TVLRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTTVQLLWVNLIMDTLGA 797
+ GR +YNN ++FI++ ++ N+ +V F+AAV L VQLLWVNL+ D L A
Sbjct: 375 AAVAEGRAIYNNTKQFIRYMISSNIGEVVCIFVAAVLGIPDTLAPVQLLWVNLVTDGLPA 554
Query: 798 LALATERPTKELMQKKPIGRTEPLITK-IMWRNLLAQALYQIAVLLVFQF---------- 846
A+ + ++M+ KP E ++T + +R L+ A +A + F +
Sbjct: 555 TAIGFNKQDSDVMKVKPRKVNEAVVTGWLFFRYLVIGAYVGLATVAGFIWWFVYADSGPK 734
Query: 847 ------------------YGKSIFNVSKEVKNTLIFNTFVLCQVFNEFNSRSMEKLNVFE 888
Y SIF +T+ V+ ++FN N+ S + +
Sbjct: 735 LPYTELMNFDTCPTRETTYPCSIF--EDRHPSTVAMTVLVVVEMFNALNNLSENQSLLVI 908
Query: 889 GILKNHLFLGIVGITIVLQVLM--VELLRKFADTERLNWEQW 928
N +G + +T++L +L+ V L L+W W
Sbjct: 909 PPWSNLWLVGSIVLTMLLHILILYVRPLSVLFSVTPLSWADW 1034
>CB893893 similar to GP|8843813|db Ca2+-transporting ATPase-like protein
{Arabidopsis thaliana}, partial (11%)
Length = 645
Score = 124 bits (312), Expect(2) = 7e-33
Identities = 63/106 (59%), Positives = 78/106 (73%)
Frame = +1
Query: 169 VRNGRPQQISIFDVLVGDVIYLKIGDQIPADGLFLGGHSLQVDESSMTGESDHVEIEPLK 228
+R GR +ISI+D++VGDVI L IG+Q+PADG+ + GHSL +DESSMTGES V + K
Sbjct: 1 IRGGRRVEISIYDLVVGDVIPLNIGNQVPADGVVITGHSLSIDESSMTGESKIVH-KDSK 177
Query: 229 APFLLSGAKVVDGYAQMLVTAVGANTAWGQMMSSISGDNSERTPLQ 274
PF++SG KV DG MLVT VG NT WG +M+SIS D E TPLQ
Sbjct: 178 DPFMMSGCKVADGSGTMLVTGVGINTEWGLLMASISEDTGEETPLQ 315
Score = 35.4 bits (80), Expect(2) = 7e-33
Identities = 24/78 (30%), Positives = 36/78 (45%), Gaps = 31/78 (39%)
Frame = +2
Query: 276 RLDKLTSSIGKIGLAVAFLVLLVLL-------------------------------IRYF 304
RL+ + + IG +GL+VA LVL+VLL IRYF
Sbjct: 398 RLNGVATFIGIVGLSVAVLVLIVLLAR*LVDGPCLPYFS*RESSMLFCMLLKWELVIRYF 577
Query: 305 TGNTEDENGNKEYKGSKT 322
+G+T + +G K++ KT
Sbjct: 578 SGHTRNSDGTKQFIAGKT 631
>TC86961 similar to GP|1742951|emb|CAA70946.1 Ca2+-ATPase {Arabidopsis
thaliana}, partial (77%)
Length = 1488
Score = 130 bits (326), Expect = 3e-30
Identities = 72/163 (44%), Positives = 102/163 (62%), Gaps = 1/163 (0%)
Frame = +1
Query: 683 LKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFNSVATVLRW 742
LK+ G +TGDG NDAPALK ADIG++MGI GTEVAKE+SD+V+ DDNF+S+ +
Sbjct: 28 LKRTGRW*PMTGDGVNDAPALKLADIGIAMGIAGTEVAKEASDMVLADDNFSSIVAAVGE 207
Query: 743 GRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTTVQLLWVNLIMDTLGALALAT 802
GR +YNN++ FI++ ++ N+ + F+ A L VQLLWVNL+ D A AL
Sbjct: 208 GRSIYNNMKAFIRYMISSNIGEVASIFLTAALGIPEGLIPVQLLWVNLVTDGPPATALGF 387
Query: 803 ERPTKELMQKKPIGRTEPLITK-IMWRNLLAQALYQIAVLLVF 844
P K++M+K P + LI I++R L+ +A + VF
Sbjct: 388 NPPDKDIMKKPPRRSDDSLINLWILFRYLVIGIYVGLATVGVF 516
>BQ138304 homologue to GP|6688833|emb putative calcium P-type ATPase
{Neurospora crassa}, partial (13%)
Length = 412
Score = 125 bits (315), Expect = 6e-29
Identities = 63/137 (45%), Positives = 93/137 (66%)
Frame = +3
Query: 655 EEERMEKVDKIRVMARSSPMDKLLMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGI 714
E E+ME + +R+ P K +V L+K G VVA+TGDG NDAPALK+ADIG++MG
Sbjct: 3 EAEQMEAAKNASLFSRTEPTHKSKLVDLLQKAGEVVAMTGDGVNDAPALKKADIGVAMG- 179
Query: 715 QGTEVAKESSDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVS 774
GT+VAK ++D+V++DDNF ++ + GR +YNN Q+FI++ ++ N+ +V F+ A +
Sbjct: 180 SGTDVAKLAADMVLVDDNFATIEGAVEEGRSIYNNTQQFIRYLISSNIGEVVSIFLTAAA 359
Query: 775 SGDVPLTTVQLLWVNLI 791
L VQLLWVNL+
Sbjct: 360 GMPEALIPVQLLWVNLV 410
>AI974405 similar to SP|Q9LF79|ACA Calcium-transporting ATPase 8 plasma
membrane-type (EC 3.6.3.8) (Ca2+-ATPase isoform 8).,
partial (10%)
Length = 332
Score = 124 bits (312), Expect = 1e-28
Identities = 62/110 (56%), Positives = 82/110 (74%)
Frame = +2
Query: 263 ISGDNSERTPLQARLDKLTSSIGKIGLAVAFLVLLVLLIRYFTGNTEDENGNKEYKGSKT 322
IS D E TPLQ RL+ + + IG +GL+VA LVL+VLL RYF+G+T + +G K++ KT
Sbjct: 2 ISEDTGEETPLQVRLNGVATFIGIVGLSVAVLVLIVLLARYFSGHTRNSDGTKQFIAGKT 181
Query: 323 DINDVCNAVVSIVAAAVTIVVVAIPEGLPLAVTLTLAYSMKRMMADQAMV 372
+ + I+ AVTIVVVA+PEGLPLAVTLTLAYSM++MMAD+A+V
Sbjct: 182 KAGHAIDGAIKIITVAVTIVVVAVPEGLPLAVTLTLAYSMRKMMADKALV 331
>BF004483 GP|21314227| type IIB calcium ATPase MCA5 {Medicago truncatula},
partial (11%)
Length = 551
Score = 124 bits (312), Expect = 1e-28
Identities = 67/128 (52%), Positives = 84/128 (65%), Gaps = 10/128 (7%)
Frame = +1
Query: 768 NFIAAVSSGDVPLTTVQLLWVNLIMDTLGALALATERPTKELMQKKPIGRTEPLITKIMW 827
N I +SSG PLT VQLLWVN+IMDTLGALALATE P +LM++ P+GR I+ +MW
Sbjct: 46 NIICILSSGTAPLTAVQLLWVNMIMDTLGALALATEPPNDDLMKRAPVGRKGNFISNVMW 225
Query: 828 RNLLAQALYQIAVLLVFQFYGKSIFNV----SKEVKNTLIFNTFVLCQVF------NEFN 877
RN+L Q+LYQ V+ Q GK+IF++ S V NTLIFN FV CQV N F
Sbjct: 226 RNILGQSLYQFMVIWFLQSKGKTIFSLDGPNSDLVLNTLIFNAFVFCQVLFLKRLNNLFT 405
Query: 878 SRSMEKLN 885
+ S E ++
Sbjct: 406 NNSYESMS 429
Score = 29.6 bits (65), Expect = 5.9
Identities = 12/18 (66%), Positives = 15/18 (82%)
Frame = +2
Query: 868 VLCQVFNEFNSRSMEKLN 885
++ QVFNE NSR MEK+N
Sbjct: 497 LVLQVFNEINSREMEKIN 550
>CB892538 similar to SP|Q9XES1|ECA Calcium-transporting ATPase 4 endoplasmic
reticulum-type (EC 3.6.3.8). [Mouse-ear cress], partial
(24%)
Length = 805
Score = 82.8 bits (203), Expect(2) = 1e-26
Identities = 63/190 (33%), Positives = 96/190 (50%), Gaps = 12/190 (6%)
Frame = +2
Query: 487 FNSEKKRSGVAVRKETNNTVHVHWKGAAEMVLAMCSNYIDSNGTQKSLDEE-RSKIEKII 545
F+ ++K GV V + KGA E VL S +G+ LD ++ I + +
Sbjct: 59 FDRDRKSMGVIVDSGVGKKKSLLVKGAVENVLDRSSKVQLRDGSVVKLDNNAKNLILQAL 238
Query: 546 QGMAASSLRCIAFAYM-EISEGGDYI--EKGKPRQVLR--------EDGLTLLGIVGLKD 594
M+ S+LRC+ FAY E++ +Y E Q+L ED L +G+VGL+D
Sbjct: 239 HEMSTSALRCLGFAYKDELTNFENYNGNEDHPAHQLLLDPNNYSSIEDELIFVGLVGLRD 418
Query: 595 PCRPNVKKAVETCKLAGVDIKMITGDNIFTAKAIATECGILDLNDAGGVVVEGVEFRNYT 654
P R V +A+E C+ AG+ + +ITGDN TA+AI E G+ N E + ++ T
Sbjct: 419 PPREEVYQAIEDCRAAGIRVMVITGDNKNTAEAICREIGVFAPN-------ENISSKSLT 577
Query: 655 EEERMEKVDK 664
++ ME DK
Sbjct: 578 GKDFMELRDK 607
Score = 56.2 bits (134), Expect(2) = 1e-26
Identities = 28/39 (71%), Positives = 32/39 (81%)
Frame = +3
Query: 683 LKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAK 721
LK+ G VVA+TGDG NDA ALK ADIG++MGI TEVAK
Sbjct: 687 LKRTGEVVAMTGDGCNDATALKLADIGIAMGIARTEVAK 803
>BG647796 GP|21314227|g type IIB calcium ATPase MCA5 {Medicago truncatula},
partial (9%)
Length = 780
Score = 101 bits (252), Expect = 1e-21
Identities = 49/93 (52%), Positives = 63/93 (67%)
Frame = +1
Query: 858 VKNTLIFNTFVLCQVFNEFNSRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMVELLRKF 917
V NTLIFN FV CQVFNE NSR MEK+NVF+GIL N++F+G++ TI Q+++VE L F
Sbjct: 13 VLNTLIFNAFVFCQVFNEINSREMEKINVFKGILDNYVFVGVISATIFFQIIIVEYLGTF 192
Query: 918 ADTERLNWEQWGICIGIAAVSWPIAWLTKLTPV 950
A+T L QW C+ + + PIA K PV
Sbjct: 193 ANTTPLTLVQWFFCLFVGFMGMPIAARLKKIPV 291
>CB893927 homologue to GP|17342714|g type IIA calcium ATPase {Medicago
truncatula}, partial (13%)
Length = 413
Score = 84.0 bits (206), Expect(2) = 8e-19
Identities = 48/115 (41%), Positives = 74/115 (63%), Gaps = 3/115 (2%)
Frame = +2
Query: 608 KLAGVDIKMITGDNIFTAKAIATECGILDLN-DAGGVVVEGVEFRNYTEEERMEKVDKI- 665
K AG+ + +ITG+N TA+AI E + + D G + G EF + + E+++ + +
Sbjct: 2 KQAGIRVMVITGENKSTAEAICKEIKLFSTDEDLTGQSLTGKEFMSLSHSEQVKLLLRNG 181
Query: 666 -RVMARSSPMDKLLMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEV 719
+ +R+ P K +V+ LK+ G +VA+TGDG NDAPALK ADIG++MGI GTE+
Sbjct: 182 GKGFSRAEPRHKQEIVRLLKEMGEIVAMTGDGVNDAPALKLADIGMAMGITGTEM 346
Score = 28.9 bits (63), Expect(2) = 8e-19
Identities = 9/22 (40%), Positives = 17/22 (76%)
Frame = +1
Query: 730 DDNFNSVATVLRWGRCVYNNIQ 751
DDNF+++ + + GR +YNN++
Sbjct: 343 DDNFSTIVSAIAEGRAIYNNMK 408
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.136 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,894,954
Number of Sequences: 36976
Number of extensions: 319980
Number of successful extensions: 1611
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 1525
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1553
length of query: 967
length of database: 9,014,727
effective HSP length: 105
effective length of query: 862
effective length of database: 5,132,247
effective search space: 4423996914
effective search space used: 4423996914
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0322.1


