

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0321.6
(497 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
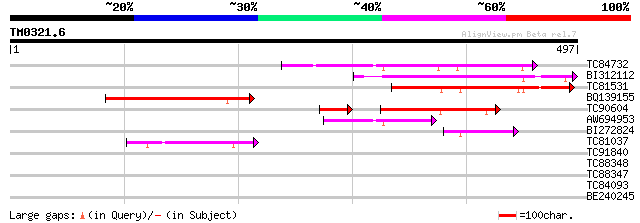
Score E
Sequences producing significant alignments: (bits) Value
TC84732 homologue to GP|14334996|gb|AAK59762.1 At1g05230/YUP8H12... 126 2e-29
BI312112 similar to GP|22475195|gb homeodomain protein GhHOX1 {G... 117 8e-27
TC81531 homologue to GP|3925363|gb|AAC79430.1| homeodomain prote... 117 1e-26
BQ139155 similar to GP|3925363|gb| homeodomain protein {Malus x ... 109 2e-24
TC90604 weakly similar to PIR|B96760|B96760 probable homeobox pr... 77 5e-17
AW694953 similar to GP|14334996|gb At1g05230/YUP8H12_16 {Arabido... 63 2e-10
BI272824 similar to GP|1814424|gb|A homeodomain protein AHDP {Ar... 46 4e-05
TC81037 homologue to GP|18076740|emb|CAC84277. HD-Zip protein {Z... 42 4e-04
TC91840 similar to PIR|T10695|T10695 transcription factor HD-zip... 40 0.002
TC88348 homologue to GP|13774427|gb|AAK38870.1 translation initi... 29 3.6
TC88347 homologue to GP|13774427|gb|AAK38870.1 translation initi... 29 3.6
TC84093 similar to GP|14324717|dbj|BAB59644. unknown product {Th... 28 6.2
BE240245 similar to GP|9759334|dbj gene_id:MUP24.12~unknown prot... 28 8.1
>TC84732 homologue to GP|14334996|gb|AAK59762.1 At1g05230/YUP8H12_16
{Arabidopsis thaliana}, partial (35%)
Length = 808
Score = 126 bits (317), Expect = 2e-29
Identities = 85/244 (34%), Positives = 130/244 (52%), Gaps = 20/244 (8%)
Frame = +3
Query: 239 GAKRWLLELQRTFEKKACSEFSYTASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNT 298
GAKRW+ L R E+ A + + + + G + DGRKS++ L+ RM SF ++
Sbjct: 3 GAKRWIATLDRQCERLASAMATNIPTVD--VGVITNQDGRKSMLKLAERMCISFCAGVSA 176
Query: 299 QGIQNLLDPSNQLTTSIKLIAHRNTILD-----GIVVIAVSSIWLPVTPKNLFLFLKDEF 353
S ++++ R ++ D GIV+ A +S WLPV PK +F FL+DE
Sbjct: 177 STAHTWTTLSGTGADDVRVMT-RKSVDDPGRPAGIVLSAATSFWLPVPPKRVFEFLRDEN 353
Query: 354 RKAEWDFLAGGRPMRQKAHIS--INESNHVSILQTSSIIG--EHMVIFQESSIDDLGGYV 409
++EWD L+ G +++ AHI+ + N VS+L+ +S +M+I QES D G +V
Sbjct: 354 SRSEWDILSNGGVVQEMAHIANGRDTGNCVSLLRVNSANSSQSNMLILQESCTDTTGSFV 533
Query: 410 VYAPIDKLDAYMIVNGCDSSKKLILPSGFTISRDGREGN-----------EGSLLTMCFQ 458
+YAP+D + +++NG D +LPSGF I DG N GSLLT+ Q
Sbjct: 534 IYAPVDIVAMNVVLNGGDPDYVALLPSGFAILPDGTTTNGGKVLVKLDMGGGSLLTVALQ 713
Query: 459 VLVD 462
+LVD
Sbjct: 714 ILVD 725
>BI312112 similar to GP|22475195|gb homeodomain protein GhHOX1 {Gossypium
hirsutum}, partial (27%)
Length = 727
Score = 117 bits (294), Expect = 8e-27
Identities = 77/214 (35%), Positives = 112/214 (51%), Gaps = 18/214 (8%)
Frame = +2
Query: 302 QNLLDPSNQLTTSIKLIAHRNTILDGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFL 361
+NL DPS L G++V AVSSIWLP++P LF FL+DE R+ EWD +
Sbjct: 44 KNLNDPSEPL---------------GLIVCAVSSIWLPISPNVLFDFLRDETRRTEWDIM 178
Query: 362 AGGRPMRQKAHISINESNHVSI-LQTSSIIGEHMVIFQESSIDDLGGYVVYAPIDKLDAY 420
+ G ++ A+++ + ++ +QT +M I Q+S + VVYAP D
Sbjct: 179 SNGGTVQSIANLAKGQDRGNAVTIQTIKSKENNMWILQDSCTNSYESMVVYAPADITGIQ 358
Query: 421 MIVNGCDSSKKLILPSGFTISRDGREGNE--------------GSLLTMCFQVLVDEEGS 466
++ GCDSS ILPSGF+I DG E + GSL T+ FQ+L +
Sbjct: 359 SVMTGCDSSNLAILPSGFSIVSDGLESRQMVITSRREEKNTEGGSLFTIAFQILT----N 526
Query: 467 SSQVRKKTLESIDSSCMLM---LQRIKDALHCSD 497
+S K T+ES+DS L+ L+ IK +L+C D
Sbjct: 527 ASPTAKLTMESVDSMNSLVSCTLRHIKTSLNCED 628
>TC81531 homologue to GP|3925363|gb|AAC79430.1| homeodomain protein {Malus x
domestica}, partial (25%)
Length = 888
Score = 117 bits (292), Expect = 1e-26
Identities = 70/182 (38%), Positives = 111/182 (60%), Gaps = 21/182 (11%)
Frame = +3
Query: 335 SIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHISINE--SNHVSILQTSSIIGE 392
S+WLPV+P+ LF FL+DE ++EWD L+ G PM++ AHI+ + N VS+L+ S++
Sbjct: 3 SVWLPVSPQRLFDFLRDERLRSEWDILSNGGPMQEMAHIAKGQDHGNCVSLLRASAMNSN 182
Query: 393 H--MVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLILPSGFTISRDG------ 444
M+I QE+ ID+ G VVYAP+D ++++NG DS+ +LPSGF I DG
Sbjct: 183 QSSMLILQETCIDEAGSLVVYAPVDIPAMHVVMNGGDSAYVALLPSGFAIVPDGPGSRGL 362
Query: 445 -------REGNE----GSLLTMCFQVLVDEEGSSSQVRKKTLESIDSSCMLMLQRIKDAL 493
G E GSLLT+ FQ+LV+ ++++ +++E++++ +Q+IK AL
Sbjct: 363 ENGDATANGGGEARVSGSLLTVAFQILVNSL-PTAKLTVESVETVNNLISCTVQKIKAAL 539
Query: 494 HC 495
C
Sbjct: 540 QC 545
>BQ139155 similar to GP|3925363|gb| homeodomain protein {Malus x domestica},
partial (24%)
Length = 505
Score = 109 bits (273), Expect = 2e-24
Identities = 51/134 (38%), Positives = 88/134 (65%), Gaps = 4/134 (2%)
Frame = +1
Query: 85 ESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALRLMWN 144
E+S++S +V+ + + LV+ +DS++W+++ P +++++ T +V G +GAL+LM
Sbjct: 85 EASRESGVVIINSLALVETLMDSNRWSEMFPCVIARSSTTEVISSGINGTRNGALQLMQA 264
Query: 145 EVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRS----VAKKFPSGCRI 200
E+ VLSPLVP RE FLR+C+Q G W +VDVS+D+ ++ + ++ PSG +
Sbjct: 265 ELQVLSPLVPVREVSFLRFCKQHAEGVWAVVDVSIDTIXETSAGAPTFLTCRRLPSGXVV 444
Query: 201 RELPNGCCMVTWVE 214
+++PNG VTWVE
Sbjct: 445 QDMPNGYSKVTWVE 486
>TC90604 weakly similar to PIR|B96760|B96760 probable homeobox protein
T9L24.43 [imported] - Arabidopsis thaliana, partial (8%)
Length = 1097
Score = 77.4 bits (189), Expect(2) = 5e-17
Identities = 41/109 (37%), Positives = 67/109 (60%), Gaps = 4/109 (3%)
Frame = +1
Query: 326 DGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHISIN--ESNHVSI 383
+G VV A +++WLP+ + +F LKD ++++WD L+ G PM + AHIS N +SI
Sbjct: 628 NGTVVTAATTLWLPLPAQKVFELLKDPTKRSQWDGLSCGNPMHEIAHISNGPYHGNCISI 807
Query: 384 LQTSSIIGEHMVIFQESSIDDLGGYVVYAPIDK--LDAYMIVNGCDSSK 430
+++ MVI QES +G Y++YAPID+ +D + VN +S++
Sbjct: 808 IKSFIPTQRQMVILQESFTSPVGSYIIYAPIDRKTVDVALSVNKKESNQ 954
Score = 28.1 bits (61), Expect(2) = 5e-17
Identities = 13/29 (44%), Positives = 19/29 (64%)
Frame = +3
Query: 272 VKLLDGRKSVINLSNRMVGSFFKVLNTQG 300
++ L+GR SVI L++RMV F + L G
Sbjct: 450 IQTLEGRNSVIKLADRMVKMFCECLTMPG 536
>AW694953 similar to GP|14334996|gb At1g05230/YUP8H12_16 {Arabidopsis
thaliana}, partial (16%)
Length = 404
Score = 63.2 bits (152), Expect = 2e-10
Identities = 36/104 (34%), Positives = 59/104 (56%), Gaps = 5/104 (4%)
Frame = +2
Query: 276 DGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIKLIAHRNTILD-----GIVV 330
DG+KS++ L+ RM +F ++ S ++++ R ++ D GIV+
Sbjct: 68 DGKKSMLKLAERMCITFCAGVSASTAHTWTTLSGTGADDVRVMT-RKSVNDPERPAGIVL 244
Query: 331 IAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS 374
A +S WLPV PK +F FLK+E K+EWD L+ G +++ AHI+
Sbjct: 245 SAATSFWLPVPPKRVFEFLKNENSKSEWDILSNGGVVQKMAHIA 376
>BI272824 similar to GP|1814424|gb|A homeodomain protein AHDP {Arabidopsis
thaliana}, partial (8%)
Length = 268
Score = 45.8 bits (107), Expect = 4e-05
Identities = 25/68 (36%), Positives = 41/68 (59%), Gaps = 2/68 (2%)
Frame = +1
Query: 381 VSILQTSSIIGEH--MVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLILPSGF 438
VS+L+ +++ M+I QE+ +D VVYAP+D ++++G DS+ +LPSGF
Sbjct: 1 VSLLRANAVNANDSSMLILQETWMDTSCSVVVYAPVDGQSLNVVMSGGDSAYVALLPSGF 180
Query: 439 TISRDGRE 446
I DG +
Sbjct: 181 AIVPDGND 204
>TC81037 homologue to GP|18076740|emb|CAC84277. HD-Zip protein {Zinnia
elegans}, partial (25%)
Length = 650
Score = 42.4 bits (98), Expect = 4e-04
Identities = 31/125 (24%), Positives = 62/125 (48%), Gaps = 9/125 (7%)
Frame = +1
Query: 103 VFLDSDKWADILPTIVS---KAKTIQVFDVGSIENHSGALRLMWNEVHVLSPLVPSREYF 159
V L+ K A+IL +S + + ++VF + N G + L++ + + + L P+R+++
Sbjct: 61 VSLEPTKIAEILKDRLSWFRECRNLEVFTMFPAGN-GGTIELIYTQTYAPTTLAPARDFW 237
Query: 160 FLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAKKF------PSGCRIRELPNGCCMVTWV 213
LRY ++ G ++ + SL G + + A +F PSG IR G ++ V
Sbjct: 238 TLRYTTTLDNGSLVVCERSLSGSGTGPNPTAAAQFVRAEMLPSGYLIRPCDGGGSIIHIV 417
Query: 214 EQVEV 218
+ + +
Sbjct: 418 DHLNL 432
>TC91840 similar to PIR|T10695|T10695 transcription factor HD-zip -
Arabidopsis thaliana, partial (40%)
Length = 1043
Score = 40.0 bits (92), Expect = 0.002
Identities = 32/125 (25%), Positives = 59/125 (46%), Gaps = 9/125 (7%)
Frame = +3
Query: 103 VFLDSDKWADIL---PTIVSKAKTIQVFDVGSIENHSGALRLMWNEVHVLSPLVPSREYF 159
V L+ + A+IL P + + + +V N G + L++ +++ + L P+R+++
Sbjct: 531 VGLEPTRVAEILKDRPLWFRDCRAVDIVNVLPTAN-GGTIELLYMQLYAPTTLAPARDFW 707
Query: 160 FLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAKKF------PSGCRIRELPNGCCMVTWV 213
LRY VE G +I + SL + N S F PSG IR G ++ V
Sbjct: 708 LLRYTSVVEDGSLVICERSLKNTQNGPSMPPVPHFVRADMLPSGYLIRPCEGGGSIIHIV 887
Query: 214 EQVEV 218
+ +++
Sbjct: 888 DHMDL 902
>TC88348 homologue to GP|13774427|gb|AAK38870.1 translation initiation
factor IF1 {Glycine max}, partial (53%)
Length = 688
Score = 29.3 bits (64), Expect = 3.6
Identities = 13/34 (38%), Positives = 20/34 (58%)
Frame = -1
Query: 425 GCDSSKKLILPSGFTISRDGREGNEGSLLTMCFQ 458
GCD S+ L + + + + DGR G SL+ M F+
Sbjct: 133 GCDGSRGLAMVNDWGVEWDGR*GKHVSLMMMAFR 32
>TC88347 homologue to GP|13774427|gb|AAK38870.1 translation initiation
factor IF1 {Glycine max}, partial (42%)
Length = 765
Score = 29.3 bits (64), Expect = 3.6
Identities = 13/34 (38%), Positives = 20/34 (58%)
Frame = -3
Query: 425 GCDSSKKLILPSGFTISRDGREGNEGSLLTMCFQ 458
GCD S+ L + + + + DGR G SL+ M F+
Sbjct: 214 GCDGSRGLAMVNDWGVEWDGR*GKHVSLMMMAFR 113
>TC84093 similar to GP|14324717|dbj|BAB59644. unknown product {Thermoplasma
volcanium}, partial (6%)
Length = 731
Score = 28.5 bits (62), Expect = 6.2
Identities = 19/77 (24%), Positives = 34/77 (43%)
Frame = +1
Query: 365 RPMRQKAHISINESNHVSILQTSSIIGEHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVN 424
R R + H + +N ++ + +M++ + I Y+PI+ L I N
Sbjct: 208 RNRRSRPHRKWSLTNRWITIRVRPLNFRYMILHRSIIIRCQNRIPRYSPIENLRRVRISN 387
Query: 425 GCDSSKKLILPSGFTIS 441
C +K+ P GFTI+
Sbjct: 388 SC*PIRKMEDPVGFTIT 438
>BE240245 similar to GP|9759334|dbj gene_id:MUP24.12~unknown protein
{Arabidopsis thaliana}, partial (17%)
Length = 487
Score = 28.1 bits (61), Expect = 8.1
Identities = 13/41 (31%), Positives = 22/41 (52%)
Frame = +1
Query: 321 RNTILDGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFL 361
R I+ G ++ AVS W P + N L L+D+ + + +L
Sbjct: 124 RRRIIQGGLIRAVSESWYPSSDDNFGLLLEDDIEVSPYYYL 246
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.134 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,991,893
Number of Sequences: 36976
Number of extensions: 206171
Number of successful extensions: 881
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 863
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 872
length of query: 497
length of database: 9,014,727
effective HSP length: 100
effective length of query: 397
effective length of database: 5,317,127
effective search space: 2110899419
effective search space used: 2110899419
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0321.6


