

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0316.5
(318 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
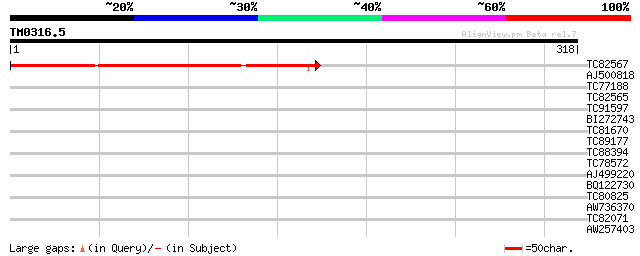
Score E
Sequences producing significant alignments: (bits) Value
TC82567 similar to GP|20161356|dbj|BAB90280. contains EST AU0704... 239 1e-63
AJ500818 weakly similar to GP|10435304|dbj unnamed protein produ... 32 0.25
TC77188 similar to PIR|T04862|T04862 probable serine/threonine-s... 32 0.42
TC82565 similar to GP|6041842|gb|AAF02151.1| unknown protein {Ar... 31 0.55
TC91597 similar to GP|21323350|dbj|BAB97978. Hypothetical protei... 30 1.2
BI272743 homologue to GP|9279730|dbj formin-like protein {Arabid... 30 1.6
TC81670 weakly similar to PIR|E86293|E86293 hypothetical protein... 30 1.6
TC89177 29 2.7
TC88394 similar to PIR|T47576|T47576 FKBP12 interacting protein ... 29 2.7
TC78572 similar to GP|9801280|emb|CAC03581.1 putative 1-deoxy-D-... 28 3.6
AJ499220 similar to SP|Q09863|YAF Hypothetical protein C29E6.10c... 28 3.6
BQ122730 homologue to GP|3204103|emb hypothetical protein {Cicer... 28 3.6
TC80825 similar to PIR|F84614|F84614 probable kinesin heavy chai... 28 3.6
AW736370 27 8.0
TC82071 weakly similar to PIR|T00411|T01608 hypothetical protein... 27 8.0
AW257403 weakly similar to GP|12039277|gb| hypothetical protein ... 27 8.0
>TC82567 similar to GP|20161356|dbj|BAB90280. contains EST
AU070442(S15668)~unknown protein {Oryza sativa (japonica
cultivar-group)}, partial (43%)
Length = 680
Score = 239 bits (610), Expect = 1e-63
Identities = 129/177 (72%), Positives = 146/177 (81%), Gaps = 3/177 (1%)
Frame = +1
Query: 1 MAETVNHLNNHDSKEKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYL 60
M + NH NN D++EK+AS LAALLKEMKEGLD V+ KIQ+LTAKVK Q TADGFSYL
Sbjct: 142 MEKPENHFNNDDNREKQASQLAALLKEMKEGLDNVKSKIQTLTAKVKN-QDSTADGFSYL 318
Query: 61 EAKNLLLLNYCQSLVYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKL 120
EAKNLLLLNYCQSLVYYLLRKAKG SIEEHPVVRSIVEIRL+LEKIRPIDKKQQYQIQKL
Sbjct: 319 EAKNLLLLNYCQSLVYYLLRKAKGCSIEEHPVVRSIVEIRLFLEKIRPIDKKQQYQIQKL 498
Query: 121 MKVGENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQ---EGHDAYRP 174
+K E+A ++ +KEP AS KSEDVSKYRPNPDMLVS+ + + +G + YRP
Sbjct: 499 IKASESATSN--TGEKEPAASKKSEDVSKYRPNPDMLVSKVEPTAEDDGDGDNVYRP 663
>AJ500818 weakly similar to GP|10435304|dbj unnamed protein product {Homo
sapiens}, partial (12%)
Length = 457
Score = 32.3 bits (72), Expect = 0.25
Identities = 23/86 (26%), Positives = 42/86 (48%), Gaps = 2/86 (2%)
Frame = -3
Query: 184 QDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKME 243
++K +++R +R++E +E K R EEK +E + E +E +R K +
Sbjct: 266 EEKRQEEKRQEEKRQEEKRQEEKRQEEKRQEEKRQEEKRQEEKRQEEKRQEEKRQEEKRQ 87
Query: 244 ERAWQEE--ELFNRVPLTKHERKREK 267
E QEE + R+ + E KR++
Sbjct: 86 EEKRQEEKRQEEKRLEEKRQEEKRQE 9
Score = 30.0 bits (66), Expect = 1.2
Identities = 19/68 (27%), Positives = 34/68 (49%)
Frame = -3
Query: 184 QDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKME 243
++K +++R +R++E +E K R EEK +E + E +E +R K +
Sbjct: 206 EEKRQEEKRQEEKRQEEKRQEEKRQEEKRQEEKRQEEKRQEEKRQEEKRQEEKRLEEKRQ 27
Query: 244 ERAWQEEE 251
E QEE+
Sbjct: 26 EEKRQEEK 3
>TC77188 similar to PIR|T04862|T04862 probable serine/threonine-specific
protein kinase (EC 2.7.1.-) F28A21.110 - Arabidopsis
thaliana, partial (79%)
Length = 1881
Score = 31.6 bits (70), Expect = 0.42
Identities = 24/96 (25%), Positives = 44/96 (45%), Gaps = 11/96 (11%)
Frame = +1
Query: 128 ATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPT------- 180
+T D PS A K+ ++K + NP++L+ R +L GH + V A
Sbjct: 94 STPDYPSMAVVAAPKKNNSMNK-KDNPNLLLGRFELGKLLGHGTFAKVHLAKNLKTGESV 270
Query: 181 ---SMDQDKTSKQER-NALRREKEILKESKHSSFIR 212
+ +DK K + ++RE IL+ +H + ++
Sbjct: 271 AIKIISKDKILKSGLVSHIKREISILRRVRHPNIVQ 378
>TC82565 similar to GP|6041842|gb|AAF02151.1| unknown protein {Arabidopsis
thaliana}, partial (65%)
Length = 1141
Score = 31.2 bits (69), Expect = 0.55
Identities = 25/86 (29%), Positives = 42/86 (48%), Gaps = 7/86 (8%)
Frame = +1
Query: 185 DKTSKQERNA-LRREKEILKESKHSSFIRTLM---NDIEEKPEEIRDYEGSSKEVERHIA 240
+K KQ R +RR + +LKES+H + + M N++ E E + EG S ++ R A
Sbjct: 259 NKRRKQLREQFMRRTRLVLKESEHDPWCKRYMELYNELRENWERLYWDEGFSNKLARDHA 438
Query: 241 KMEERAWQEEELF---NRVPLTKHER 263
E +E+ NR P ++ +
Sbjct: 439 NYESAEDDDEDFSPYRNRRPQMEYNK 516
>TC91597 similar to GP|21323350|dbj|BAB97978. Hypothetical protein
{Corynebacterium glutamicum ATCC 13032}, partial (5%)
Length = 718
Score = 30.0 bits (66), Expect = 1.2
Identities = 31/155 (20%), Positives = 63/155 (40%), Gaps = 3/155 (1%)
Frame = +2
Query: 120 LMKVGENAATSDLPSKKEPVASDKSEDV---SKYRPNPDMLVSREDLAPQEGHDAYRPVK 176
L ++ ++ +P KK KS S+ NP + D +G V
Sbjct: 14 LSRIAHQNQSTAMPRKKSSAPRSKSPSKGQPSRKSSNPQLATQASDSKDTKGRKV-DVVA 190
Query: 177 FAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVE 236
AP +K+ + ++ +++E K ++ + + +EKP + + + VE
Sbjct: 191 EAPELGPPAALTKETKESVEKQEE-----KPATKLEEKPVEKQEKPIKKPAKKQEDRPVE 355
Query: 237 RHIAKMEERAWQEEELFNRVPLTKHERKREKYLKK 271
+H +EE ++E+ + + +R EK KK
Sbjct: 356 KHEKPVEEPIEKQEKPIKKPAKKQEDRPIEKQEKK 460
>BI272743 homologue to GP|9279730|dbj formin-like protein {Arabidopsis
thaliana}, partial (2%)
Length = 370
Score = 29.6 bits (65), Expect = 1.6
Identities = 22/98 (22%), Positives = 46/98 (46%), Gaps = 12/98 (12%)
Frame = +1
Query: 191 ERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKE-----------VERHI 239
E NA EKE+ ++ + FI ++++ EE+ ++ ++ +E +
Sbjct: 67 EHNAAVEEKEVQEKVEKIIFIEEEQEEMKDVEEEVELFDADDEDGDKGSEIDYILIEENN 246
Query: 240 AKMEERAWQEEELFNRVPLTKH-ERKREKYLKKSRNGL 276
+ EE +EEE + V T+ +K E +++K + L
Sbjct: 247 IEEEEHEEEEEEEESSVLSTEELNKKFEDFIRKMKEDL 360
>TC81670 weakly similar to PIR|E86293|E86293 hypothetical protein AAF18488.1
[imported] - Arabidopsis thaliana, partial (41%)
Length = 1797
Score = 29.6 bits (65), Expect = 1.6
Identities = 34/207 (16%), Positives = 85/207 (40%)
Frame = +2
Query: 102 YLEKIRPIDKKQQYQIQKLMKVGENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSRE 161
++E+ R + +K + +++++++ ++E + ++ E + K L RE
Sbjct: 947 FVEESRSMQRKAREEVRRILE------------EQEKLRNELDEKMRKLDTWSRDLNKRE 1090
Query: 162 DLAPQEGHDAYRPVKFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEK 221
L QE +++DK K RN E +L + + +EE+
Sbjct: 1091 VLTDQERQ-----------KLEEDKKKKDSRN----ESLMLASKEQKIADENVFRLVEEQ 1225
Query: 222 PEEIRDYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDGLTE 281
E + ++E+ + ++ + EEL ++ + KH ++ K + ++ +
Sbjct: 1226 KREKEEALNKILQLEKQLDAKQKLEMEIEELRGKLQVMKHLGDQDDTAIKKK--MEEMNS 1399
Query: 282 SFFDEIKTLPFEDETGEQVMGPSKGSN 308
D+I++L + ++ + SN
Sbjct: 1400 ELEDKIESLEDMESMNSTLIVKERQSN 1480
>TC89177
Length = 1201
Score = 28.9 bits (63), Expect = 2.7
Identities = 19/59 (32%), Positives = 29/59 (48%), Gaps = 3/59 (5%)
Frame = +2
Query: 158 VSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERNALRREK---EILKESKHSSFIRT 213
+S+ED P +G + F P D +K+ R LRR + +ILK ++ S RT
Sbjct: 116 LSKEDFVPIKGKIERAFIMFNPDGSHADDKTKRLRRLLRRREARNDILKHNRAVSASRT 292
>TC88394 similar to PIR|T47576|T47576 FKBP12 interacting protein (FIP37) -
Arabidopsis thaliana, partial (56%)
Length = 1285
Score = 28.9 bits (63), Expect = 2.7
Identities = 25/91 (27%), Positives = 43/91 (46%), Gaps = 10/91 (10%)
Frame = +1
Query: 194 ALRREKEILKESKHSSFIRTLMNDIEEKPEEIRD---------YEGSSKEVERHIAKMEE 244
A+ KE++ S S + T + +EEK ++I+D + SK + +AK
Sbjct: 682 AINAGKEVVTRSGCS*RVHTFEDLVEEKDKKIKDLQDNITAITFTSQSKMGKMLMAKCRT 861
Query: 245 RAWQEEELFNRVPLTK-HERKREKYLKKSRN 274
+ EE+ N+ K HE + L+KS+N
Sbjct: 862 LQEENEEIGNQASEGKIHELTMKLALQKSQN 954
>TC78572 similar to GP|9801280|emb|CAC03581.1 putative 1-deoxy-D-xylulose
5-phosphate reductoisomerase {Zea mays}, partial (88%)
Length = 1654
Score = 28.5 bits (62), Expect = 3.6
Identities = 25/77 (32%), Positives = 36/77 (46%), Gaps = 5/77 (6%)
Frame = +2
Query: 167 EGHDAYRPVKFAPTS-----MDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEK 221
E D +R V A S +DQ K K + A+R E + + ++ + D+EEK
Sbjct: 353 ENPDKFRVVALAAGSNITLLVDQVKRFKPQLVAVRNESLLAE-------LQEALKDVEEK 511
Query: 222 PEEIRDYEGSSKEVERH 238
PE I +G EV RH
Sbjct: 512 PEIIPGEQGVI-EVARH 559
>AJ499220 similar to SP|Q09863|YAF Hypothetical protein C29E6.10c in
chromosome I. [Fission yeast] {Schizosaccharomyces
pombe}, partial (12%)
Length = 518
Score = 28.5 bits (62), Expect = 3.6
Identities = 33/147 (22%), Positives = 57/147 (38%), Gaps = 22/147 (14%)
Frame = -3
Query: 142 DKSEDVSKYRPNPDMLVSREDLAPQEGH----------------DAYRPVKFAPTSMDQD 185
D+ ED +Y + + E+ +EG +AYR K A +
Sbjct: 438 DEDEDEDEYEEDGEEDNQTEEQRMEEGRRMFQIFAARMFEQRVLNAYRE-KVAQERQQKL 262
Query: 186 KTSKQERNALRREKEI--LKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKE----VERHI 239
+E N L+ E+E+ LKE + + +E+ R+ E ++E ER
Sbjct: 261 LEELEEENRLKEERELKKLKEKERKKAKNRALKQQKEEERARREAERLAEEKAQRAEREK 82
Query: 240 AKMEERAWQEEELFNRVPLTKHERKRE 266
EER +EEE + + +K +
Sbjct: 81 KLEEERKRREEERLKKEAERRQRKKND 1
>BQ122730 homologue to GP|3204103|emb hypothetical protein {Cicer arietinum},
partial (98%)
Length = 492
Score = 28.5 bits (62), Expect = 3.6
Identities = 18/59 (30%), Positives = 29/59 (48%)
Frame = -2
Query: 16 KEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQSL 74
+EAS LLK K ++ Q+L K++ +D ++ LE NL L N ++L
Sbjct: 461 QEASKRLELLKAEKAKIESEATMYQNLAGKMESDLQSLSDAYNSLEQSNLQLENEVKAL 285
>TC80825 similar to PIR|F84614|F84614 probable kinesin heavy chain
[imported] - Arabidopsis thaliana, partial (16%)
Length = 1771
Score = 28.5 bits (62), Expect = 3.6
Identities = 13/37 (35%), Positives = 21/37 (56%)
Frame = +2
Query: 12 DSKEKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKE 48
+ EK+ S L LK +E ++HK++ L K+KE
Sbjct: 863 NQSEKQVSQLCEKLKGKEETCCTLQHKVKELERKIKE 973
>AW736370
Length = 488
Score = 27.3 bits (59), Expect = 8.0
Identities = 23/90 (25%), Positives = 43/90 (47%)
Frame = -2
Query: 174 PVKFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSK 233
P+ F Q++ ++ ER R ++E K+ + T+M +E++ Y+ K
Sbjct: 463 PMTFDTEKEYQERLTEAEREIHRWKREYQKKDQD---YETVMGLLEQEA-----YDSR*K 308
Query: 234 EVERHIAKMEERAWQEEELFNRVPLTKHER 263
+V IAK+ ER + + +R+P K R
Sbjct: 307 DVI--IAKLNERIKENDAALDRIPGRKKRR 224
>TC82071 weakly similar to PIR|T00411|T01608 hypothetical protein At2g44820
[imported] - Arabidopsis thaliana, partial (36%)
Length = 764
Score = 27.3 bits (59), Expect = 8.0
Identities = 28/111 (25%), Positives = 48/111 (43%), Gaps = 10/111 (9%)
Frame = +1
Query: 100 RLYLEKIRPIDKKQQYQIQKLMK----VGENAATSDLPSKKEPVASDKSED----VSKYR 151
RL L RP+ KKQ+ + +K+++ +G S K+P K ED +S+ R
Sbjct: 319 RLPLSVARPMMKKQKQREEKMLQERMILGRFGGKDGGSSSKKPAGKHKPEDRGLKLSEGR 498
Query: 152 PNPDMLVSREDL--APQEGHDAYRPVKFAPTSMDQDKTSKQERNALRREKE 200
+L + L P GHD T + T K++ + + +K+
Sbjct: 499 FRNGILDVKHLLKSTPTRGHD---------TGKNMSNTGKRKGSIWKHDKK 624
>AW257403 weakly similar to GP|12039277|gb| hypothetical protein {Oryza
sativa}, partial (29%)
Length = 555
Score = 27.3 bits (59), Expect = 8.0
Identities = 21/67 (31%), Positives = 32/67 (47%), Gaps = 4/67 (5%)
Frame = -1
Query: 254 NRVPLTKHERK----REKYLKKSRNGLDGLTESFFDEIKTLPFEDETGEQVMGPSKGSNR 309
N+V H+ K +++ LK R +T D+ + L FE G +V+GPSKG
Sbjct: 384 NKVNTHTHKTKF***QDE*LKNPRTQRVNITNRKRDQERRLDFE---GREVVGPSKGR*E 214
Query: 310 NGRLKKR 316
G + R
Sbjct: 213 TGVVTMR 193
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.311 0.130 0.352
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,497,957
Number of Sequences: 36976
Number of extensions: 86186
Number of successful extensions: 456
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 440
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 444
length of query: 318
length of database: 9,014,727
effective HSP length: 96
effective length of query: 222
effective length of database: 5,465,031
effective search space: 1213236882
effective search space used: 1213236882
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0316.5


