

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0314.4
(199 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
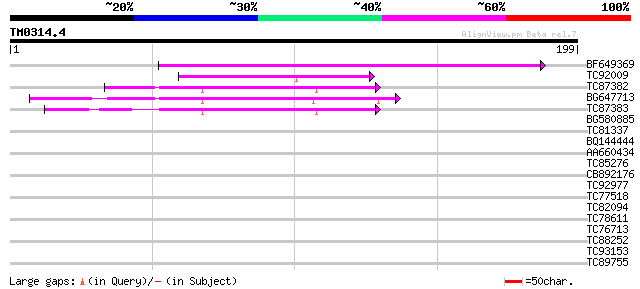
Score E
Sequences producing significant alignments: (bits) Value
BF649369 87 4e-18
TC92009 55 1e-08
TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulch... 50 4e-07
BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent vir... 44 5e-05
TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA PO... 40 8e-04
BG580885 36 0.009
TC81337 similar to GP|15723293|gb|AAL06332.1 U2 auxiliary factor... 33 0.073
BQ144444 GP|18376070|em related to tpa inducible protein {Neuros... 32 0.16
AA660434 similar to GP|15081711|gb At2g38020/T8P21.7 {Arabidopsi... 31 0.28
TC85276 weakly similar to GP|22294973|dbj|BAC08801. ORF_ID:tlr12... 30 0.81
CB892176 PIR|T09286|T09 malate dehydrogenase (EC 1.1.1.37) precu... 28 1.8
TC92977 similar to GP|15528625|dbj|BAB64646. putative splicing f... 27 4.0
TC77518 similar to SP|Q94HF1|IF3B_ORYSA Eukaryotic translation i... 27 5.2
TC82094 similar to GP|20259195|gb|AAM14313.1 unknown protein {Ar... 27 5.2
TC78611 similar to GP|15450599|gb|AAK96571.1 AT5g17440/K3M16_10 ... 27 5.2
TC76713 similar to PIR|S33622|S33622 ADR6 protein - soybean, par... 27 5.2
TC88252 similar to GP|6137207|gb|AAF04377.1| P72 DEAD box protei... 27 6.8
TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Ci... 26 8.9
TC89755 similar to GP|10177798|dbj|BAB11289. emb|CAB87778.1~gene... 26 8.9
>BF649369
Length = 631
Score = 87.0 bits (214), Expect = 4e-18
Identities = 42/136 (30%), Positives = 76/136 (55%)
Frame = +3
Query: 53 VTGRRVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQA 112
+ G++V +P+F+G D WI R E +F + ++ R+++ ++MEG + WF E
Sbjct: 111 LAGKKVKLPLFEGDDPVAWITRAEIYFDVQNTPDDMRVKLSRLSMEGPTIHWFNLLMETE 290
Query: 113 PERAWEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGADREILK 172
+ + E K ALI R+ ++NPF L +++Q GSV + E+FE ++ + E
Sbjct: 291 DDLSREKLKKALIARYDGRRLENPFEELSTLRQIGSVEEFVEAFELLSSQVGRLPEEQYL 470
Query: 173 GVFLNGLQEEIRAEMK 188
G F++GL+ IR ++
Sbjct: 471 GYFMSGLKAHIRRRVR 518
>TC92009
Length = 974
Score = 55.5 bits (132), Expect = 1e-08
Identities = 26/70 (37%), Positives = 40/70 (57%), Gaps = 1/70 (1%)
Frame = +2
Query: 60 IPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEG-KALGWFQWWEEQAPERAWE 118
+ F G+D GWI R E FF + + ++L+ ++ME KAL WF W ++ P+ W+
Sbjct: 689 LSQFTGTDPAGWITRAEMFFADNEIHSCDKLQWAFMSMEDEKALLWFYSWCQENPDADWK 868
Query: 119 PFKHALIRRF 128
F A+IR F
Sbjct: 869 SFSMAMIREF 898
>TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulchellus},
partial (7%)
Length = 2304
Score = 50.4 bits (119), Expect = 4e-07
Identities = 30/107 (28%), Positives = 55/107 (51%), Gaps = 10/107 (9%)
Frame = +2
Query: 34 FSDHSRERFDHPHRVHLRAVTGRRVDIPMFDGS----DAYGWINRVERFFQLSRVGEEER 89
+SD S R H R L+ + +VDIP F+G+ D W+ +ER F+ V EE++
Sbjct: 551 YSDSSSSRSSHSQRRQLQ-MNDIKVDIPDFEGNLQLDDFLDWLQTIERVFEYKEVPEEQK 727
Query: 90 LEMVMIAMEGKALGWFQ------WWEEQAPERAWEPFKHALIRRFQP 130
+++V ++ AL W++ E ++ + W+ + L R++ P
Sbjct: 728 VKIVAAKLKKHALIWWENLKRRRKREGKSKIKTWDKMRQKLTRKYLP 868
>BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent virus}, partial
(1%)
Length = 726
Score = 43.5 bits (101), Expect = 5e-05
Identities = 37/141 (26%), Positives = 63/141 (44%), Gaps = 11/141 (7%)
Frame = +2
Query: 8 TPRFRQASRERDREASSHSGSRTGSRFSDHSRERFDHPHRVHLRAVTGRRVDIPMFDGS- 66
T +QA E R + GS SD S R HR R ++ +VDIP F G
Sbjct: 206 TVNAQQALLEAQRRRNDDDGSG-----SDSSSSRSSRSHRRQTR-MSKIKVDIPDF*GKL 367
Query: 67 ---DAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWF------QWWEEQAPERAW 117
+ W+ +ER F+ V EE+++++V ++ A W+ + E ++ + W
Sbjct: 368 QPDEFVDWLQTIERVFKYKEVAEEQKVKIVAAKLKKHASIWWKNLKRKRNCEGKSKIKTW 547
Query: 118 EPFKHALIRRF-QPTLVQNPF 137
+ + L R++ P Q+ F
Sbjct: 548 DKMRQKLTRKYLHPHYYQDNF 610
>TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA POLYMERASE II
{Encephalitozoon cuniculi}, partial (0%)
Length = 1247
Score = 39.7 bits (91), Expect = 8e-04
Identities = 34/128 (26%), Positives = 58/128 (44%), Gaps = 10/128 (7%)
Frame = -2
Query: 13 QASRERDREASSHSGSRTGSRFSDHSRERFDHPHRVHLRAVTGRRVDIPMFDGS----DA 68
Q R +D +SS S S SR S R F + + DIP F+G+ D
Sbjct: 832 QQKRFKDHVSSSDSLS---SRSSRSQRREFQ---------MNDIK*DIPDFEGNLQPDDL 689
Query: 69 YGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQ------WWEEQAPERAWEPFKH 122
W+ +ER F+ V EE+++++V ++ A W++ E ++ + WE +
Sbjct: 688 LDWLQIMERLFKYKEVLEEQKVKIVAAKLKKLASIWWENVKRRRKREGKSKIKTWEKMRQ 509
Query: 123 ALIRRFQP 130
L R++ P
Sbjct: 508 KLTRKYLP 485
>BG580885
Length = 609
Score = 36.2 bits (82), Expect = 0.009
Identities = 33/133 (24%), Positives = 62/133 (45%), Gaps = 10/133 (7%)
Frame = +3
Query: 60 IPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVM----IAMEGKALGWFQW-WEEQAPE 114
+P+F G+ I + RF ++ R +EM + +E ++ W+ E
Sbjct: 45 LPIFRGTPNESPITHLSRFNKVCRANNASSVEMQKKIFPVTLEEESALWYDLNIEPYYIS 224
Query: 115 RAWEPFKHALIRRF-QPTLVQNPFGPLLSIKQ--KGSVMAY--RESFEEAAAPMRGADRE 169
+W+ K + ++ + + V+ L+ I Q K V +Y R + P G + +
Sbjct: 225 LSWDEIKLSFLQAYYEIEPVEELRSELMGIHQGEKERVRSYFLRLQWILKRWPEHGLEDD 404
Query: 170 ILKGVFLNGLQEE 182
++KGVF+NGL+EE
Sbjct: 405 VIKGVFVNGLREE 443
>TC81337 similar to GP|15723293|gb|AAL06332.1 U2 auxiliary factor small
subunit {Arabidopsis thaliana}, partial (81%)
Length = 1190
Score = 33.1 bits (74), Expect = 0.073
Identities = 19/42 (45%), Positives = 23/42 (54%), Gaps = 1/42 (2%)
Frame = +2
Query: 1 RTRGRSNTPRFRQASRERDRE-ASSHSGSRTGSRFSDHSRER 41
R+R RSN+PR R+ SR R+RE G R SD R R
Sbjct: 653 RSRSRSNSPRRRRRSRSRERERPRDRDYDSRGRRSSDRDRVR 778
Score = 26.2 bits (56), Expect = 8.9
Identities = 13/32 (40%), Positives = 17/32 (52%)
Frame = +2
Query: 2 TRGRSNTPRFRQASRERDREASSHSGSRTGSR 33
+RGR ++ R R +RDRE S R G R
Sbjct: 740 SRGRRSSDRDRVRDGDRDRERKRDSRDRDGGR 835
>BQ144444 GP|18376070|em related to tpa inducible protein {Neurospora
crassa}, partial (2%)
Length = 1256
Score = 32.0 bits (71), Expect = 0.16
Identities = 17/35 (48%), Positives = 22/35 (62%), Gaps = 2/35 (5%)
Frame = -3
Query: 22 ASSHSGSRTGSRFSDHSRERFDHPHRV--HLRAVT 54
AS+HS SRT R + H+R R DHP H+R V+
Sbjct: 867 ASAHSSSRTARR-APHNRNRLDHPSTTSQHIRRVS 766
>AA660434 similar to GP|15081711|gb At2g38020/T8P21.7 {Arabidopsis thaliana},
partial (16%)
Length = 579
Score = 31.2 bits (69), Expect = 0.28
Identities = 23/68 (33%), Positives = 30/68 (43%)
Frame = -2
Query: 3 RGRSNTPRFRQASRERDREASSHSGSRTGSRFSDHSRERFDHPHRVHLRAVTGRRVDIPM 62
RG + +AS ER R + +R GSR DH +E R + GRRV I +
Sbjct: 233 RGDDRRRKREEASEERTRRGAGRFWNRRGSRRGDHRKE------RRRFCSERGRRVSIAL 72
Query: 63 FDGSDAYG 70
D YG
Sbjct: 71 -DRVRVYG 51
>TC85276 weakly similar to GP|22294973|dbj|BAC08801.
ORF_ID:tlr1249~hypothetical protein {Thermosynechococcus
elongatus BP-1}, partial (33%)
Length = 1315
Score = 29.6 bits (65), Expect = 0.81
Identities = 15/37 (40%), Positives = 19/37 (50%)
Frame = +1
Query: 10 RFRQASRERDREASSHSGSRTGSRFSDHSRERFDHPH 46
R R+ RER+RE + G+R S S SR PH
Sbjct: 583 REREREREREREPRAEFGTRGNSCRSSPSRPNLQRPH 693
>CB892176 PIR|T09286|T09 malate dehydrogenase (EC 1.1.1.37) precursor -
alfalfa, partial (41%)
Length = 615
Score = 28.5 bits (62), Expect = 1.8
Identities = 15/40 (37%), Positives = 21/40 (52%)
Frame = +1
Query: 1 RTRGRSNTPRFRQASRERDREASSHSGSRTGSRFSDHSRE 40
+TRG R R+ RER+RE + G R+SD S +
Sbjct: 109 KTRGERERERERERERERERERAFSKHP**GRRYSDPSSQ 228
>TC92977 similar to GP|15528625|dbj|BAB64646. putative splicing
factor-like protein {Oryza sativa (japonica
cultivar-group)}, partial (10%)
Length = 733
Score = 27.3 bits (59), Expect = 4.0
Identities = 14/36 (38%), Positives = 17/36 (46%)
Frame = +1
Query: 1 RTRGRSNTPRFRQASRERDREASSHSGSRTGSRFSD 36
R+ RS +PR R R R S R S+FSD
Sbjct: 58 RSYSRSRSPRSRSRGRSYSRSRSPKKTRRVKSKFSD 165
>TC77518 similar to SP|Q94HF1|IF3B_ORYSA Eukaryotic translation initiation
factor 3 subunit 11 (eIF-3 p25) (eIF3k). [Rice] {Oryza
sativa}, partial (94%)
Length = 1020
Score = 26.9 bits (58), Expect = 5.2
Identities = 16/36 (44%), Positives = 20/36 (55%)
Frame = +3
Query: 6 SNTPRFRQASRERDREASSHSGSRTGSRFSDHSRER 41
++T R R+ RER+RE R G R S SRER
Sbjct: 51 ASTEREREREREREREF------RNGKRSSSSSRER 140
>TC82094 similar to GP|20259195|gb|AAM14313.1 unknown protein {Arabidopsis
thaliana}, partial (83%)
Length = 1759
Score = 26.9 bits (58), Expect = 5.2
Identities = 14/38 (36%), Positives = 19/38 (49%)
Frame = +2
Query: 102 LGWFQWWEEQAPERAWEPFKHALIRRFQPTLVQNPFGP 139
+G ++ E PE PF LIRRF + + NP P
Sbjct: 1295 MGGIMYYLEYEPE-TMAPFLRGLIRRFLASRISNPSPP 1405
>TC78611 similar to GP|15450599|gb|AAK96571.1 AT5g17440/K3M16_10 {Arabidopsis
thaliana}, partial (91%)
Length = 1636
Score = 26.9 bits (58), Expect = 5.2
Identities = 21/72 (29%), Positives = 26/72 (35%)
Frame = +1
Query: 1 RTRGRSNTPRFRQASRERDREASSHSGSRTGSRFSDHSRERFDHPHRVHLRAVTGRRVDI 60
R RG S+ R RDR+ H R R D R+D R R+ R D
Sbjct: 1084 RERGESHERGRDNDRRSRDRDNRHHDRDRGYDRDRDRDSSRYDSRSRRRSRS-RERSRDY 1260
Query: 61 PMFDGSDAYGWI 72
D Y W+
Sbjct: 1261 DRHRRHDRY*WL 1296
>TC76713 similar to PIR|S33622|S33622 ADR6 protein - soybean, partial (67%)
Length = 1790
Score = 26.9 bits (58), Expect = 5.2
Identities = 9/27 (33%), Positives = 15/27 (55%)
Frame = +3
Query: 88 ERLEMVMIAMEGKALGWFQWWEEQAPE 114
+RL +++I L WF+WW P+
Sbjct: 1260 KRLYIIVIEFVQPQLSWFRWWPVMEPK 1340
>TC88252 similar to GP|6137207|gb|AAF04377.1| P72 DEAD box protein {Pisum
sativum}, partial (20%)
Length = 990
Score = 26.6 bits (57), Expect = 6.8
Identities = 15/42 (35%), Positives = 18/42 (42%)
Frame = +3
Query: 4 GRSNTPRFRQASRERDREASSHSGSRTGSRFSDHSRERFDHP 45
GR F S+ DR+ S GS RF +RER P
Sbjct: 171 GRGGGRGFDNDSQRNDRDQSPDRGSSWSDRFKTVNRERSRSP 296
>TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Cicer
arietinum}, partial (8%)
Length = 516
Score = 26.2 bits (56), Expect = 8.9
Identities = 25/101 (24%), Positives = 41/101 (39%), Gaps = 11/101 (10%)
Frame = +2
Query: 71 WINRVERFFQLSRVGEEERLEM--VMIAMEGKALGWFQWW-------EEQAPERAWEPFK 121
W+ +ER F++ + E ++++ M+A E WW E+ W F+
Sbjct: 176 WLKEIERIFRVMQCFETQKVQFGTHMLAEEAD-----DWWISLLPVLEQDDAVVTWAMFR 340
Query: 122 HALIRRFQPTLVQNPFG-PLLSIKQKG-SVMAYRESFEEAA 160
+ R+ P V+ L +KQ SV Y F E A
Sbjct: 341 KEFLGRYFPEDVRGKKEIEFLELKQGDMSVTEYAAKFVELA 463
>TC89755 similar to GP|10177798|dbj|BAB11289.
emb|CAB87778.1~gene_id:MXA21.9~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (59%)
Length = 864
Score = 26.2 bits (56), Expect = 8.9
Identities = 7/12 (58%), Positives = 10/12 (83%)
Frame = -2
Query: 99 GKALGWFQWWEE 110
G+ +GW+ WWEE
Sbjct: 275 GECVGWWLWWEE 240
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.136 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,390,880
Number of Sequences: 36976
Number of extensions: 70142
Number of successful extensions: 409
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 403
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 408
length of query: 199
length of database: 9,014,727
effective HSP length: 91
effective length of query: 108
effective length of database: 5,649,911
effective search space: 610190388
effective search space used: 610190388
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0314.4


