

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0310b.12
(1485 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
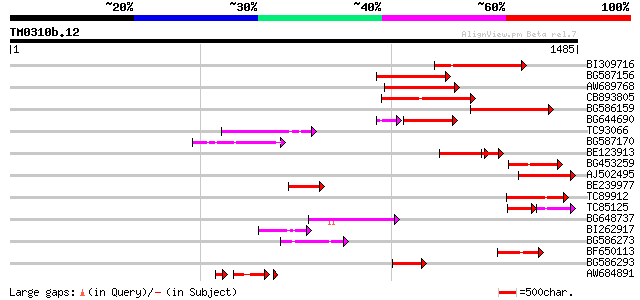
Sequences producing significant alignments: (bits) Value
BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F2... 230 4e-60
BG587156 similar to PIR|G85055|G8 probable polyprotein [imported... 204 2e-52
AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsi... 202 1e-51
CB893805 similar to GP|10177935|d copia-type polyprotein {Arabid... 192 1e-48
BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2... 174 2e-43
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 137 3e-41
TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative ret... 163 5e-40
BG587170 similar to PIR|F86470|F8 probable retroelement polyprot... 161 2e-39
BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sati... 127 4e-29
BG453259 homologue to GP|21434|emb|CA ORF4 {Solanum tuberosum}, ... 125 2e-28
AJ502495 weakly similar to GP|18071369|g putative gag-pol polypr... 117 3e-26
BE239977 weakly similar to GP|23237899|db polyprotein-like {Oryz... 113 6e-25
TC89912 weakly similar to PIR|B84512|B84512 probable retroelemen... 112 1e-24
TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-relate... 67 4e-24
BG648737 weakly similar to GP|21433|emb|CA ORF3 {Solanum tuberos... 105 2e-22
BI262917 weakly similar to GP|19920130|g Putative retroelement {... 103 5e-22
BG586273 weakly similar to PIR|F86470|F8 probable retroelement p... 99 9e-21
BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vu... 96 1e-19
BG586293 weakly similar to PIR|E84473|E84 probable retroelement ... 95 2e-19
AW684891 74 1e-18
>BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (10%)
Length = 744
Score = 230 bits (586), Expect = 4e-60
Identities = 112/242 (46%), Positives = 169/242 (69%)
Frame = +2
Query: 1113 KVCKLKKALYGLKQSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVND 1172
KVC+L+K++YGLKQ+ R W+ + + +++S GY QS D +LF K ++ T LLVYV+D
Sbjct: 17 KVCELQKSIYGLKQASRQWYSKLSESLISFGYLQSSSDFSLFTKF-KDSSFTTLLVYVDD 193
Query: 1173 MIIAGNDELEKQNLRKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKET 1232
+++AGND E Q+++ L +F++KDLG L+YFLG+EVA SKQGI ++QRKY L+LL+++
Sbjct: 194 IVLAGNDISEIQHVKCFLIDRFKIKDLGSLRYFLGLEVARSKQGILLNQRKYTLELLEDS 373
Query: 1233 GKLGCKPMGVPIEQNHKIGSSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFM 1292
G L K P + + K+ +S+ L + QY+RL+GKLIYL+ TRPDI++ V +SQF+
Sbjct: 374 GNLAVKSTLTPYDISLKLHNSDSPLYNDETQYRRLIGKLIYLTTTRPDISFAVQQLSQFV 553
Query: 1293 HDPQERHLQAVDRILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMF 1352
PQ+ H QA R+LQYLK +P +GL + + L + + D+D+A R+S TGY +F
Sbjct: 554 SKPQQVHYQAAIRVLQYLKTAPAKGLFYSATSNLKLSSFADSDWATCPTTRKSVTGYWVF 733
Query: 1353 LG 1354
LG
Sbjct: 734 LG 739
>BG587156 similar to PIR|G85055|G8 probable polyprotein [imported] -
Arabidopsis thaliana, partial (17%)
Length = 618
Score = 204 bits (520), Expect = 2e-52
Identities = 95/193 (49%), Positives = 137/193 (70%)
Frame = -1
Query: 962 HQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRKIVEKPNDKRTVGCKWI 1021
H +F+ ++ +P S +E ++DK+W +++ E A+ KN T E P K+ V +WI
Sbjct: 597 HCAFMVNLNENHIPRSYEEAMEDKEWKESVGAEAGAMIKNDTWYESELPKGKKAVSSRWI 418
Query: 1022 YTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTVRIILSLAAHFDWELQQ 1081
+T+KY++DG+++R K RLVA+G+T TYG DY ETFAPVAK++T+RI+LSLA + W L Q
Sbjct: 417 FTIKYKADGSIERKKTRLVARGFTLTYGEDYIETFAPVAKLHTIRIVLSLAVNLGWGLWQ 238
Query: 1082 FDVKNAFLHGDLEEEVYMEIPPGYDPTSGRNKVCKLKKALYGLKQSPRAWFGRFTNAMVS 1141
DVKNAFL G+LE+EVYM PPG + R V +LKKA+YGLKQSPRAW+ + + +
Sbjct: 237 MDVKNAFLQGELEDEVYMYPPPGLEHLVKRGNVLRLKKAIYGLKQSPRAWYNKLSTTLNG 58
Query: 1142 LGYKQSQGDHTLF 1154
G+++S+ DHTLF
Sbjct: 57 RGFRKSELDHTLF 19
>AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsis thaliana},
partial (9%)
Length = 675
Score = 202 bits (513), Expect = 1e-51
Identities = 95/197 (48%), Positives = 139/197 (70%)
Frame = +1
Query: 982 LKDKKWIQAMNEEMHALDKNGTRKIVEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVA 1041
L D +W+QAM E AL N T +V P K+ +GCKW+Y VK DG+++++KARLVA
Sbjct: 82 LSDPRWLQAMKTEYKALIDNKTWDLVPLPPHKKAIGCKWVYRVKENPDGSVNKFKARLVA 261
Query: 1042 KGYTQTYGIDYDETFAPVAKMNTVRIILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEI 1101
KG++QT G DY ETF+PV K T+R+IL++A + WE+QQ D+ NAFL+G L+EEVYM
Sbjct: 262 KGFSQTLGCDYTETFSPVIKPVTIRLILTIAITYKWEIQQIDINNAFLNGFLQEEVYMSQ 441
Query: 1102 PPGYDPTSGRNKVCKLKKALYGLKQSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNG 1161
P G++ + ++ VCKL K+LYGLKQ+PRAW+ T+A + G+ +S+ D +L I + +NG
Sbjct: 442 PQGFE-AANKSLVCKLNKSLYGLKQAPRAWYEXLTSAQIQFGFTKSRCDPSLLI-YNQNG 615
Query: 1162 KLTLLLVYVNDMIIAGN 1178
L +YV+D++I G+
Sbjct: 616 ACIYLXIYVDDILITGS 666
>CB893805 similar to GP|10177935|d copia-type polyprotein {Arabidopsis
thaliana}, partial (14%)
Length = 778
Score = 192 bits (487), Expect = 1e-48
Identities = 95/248 (38%), Positives = 159/248 (63%), Gaps = 3/248 (1%)
Frame = +3
Query: 975 PTSVQEVLKDKKWIQAMNEEMHALDKNGTRKIVEKPNDKRTVGCKWIYTVKYRSDGTLDR 1034
PT+ +E +K +KW +MN EM A ++N T ++ + + +T+G KWI+ K +G +++
Sbjct: 42 PTTFEEAVKSEKWRASMNNEMEATERNNTWELTDLRSGAKTIGLKWIFKTKLNENGEIEK 221
Query: 1035 YKARLVAKGYTQTYGIDYDETFAPVAKMNTVRIILSLAAHFDWELQQFDVKNAFL---HG 1091
YKARLVAKGY+Q YG+DY E FAPVA+ +T+R++++LAA ++++ V + + H
Sbjct: 222 YKARLVAKGYSQQYGVDYTEVFAPVARWDTIRMVIALAA----QIKRDGVCIS*M*KAHS 389
Query: 1092 DLEEEVYMEIPPGYDPTSGRNKVCKLKKALYGLKQSPRAWFGRFTNAMVSLGYKQSQGDH 1151
+E + + + R ++K+ALYGLKQ+PRAW+ R G+++ +H
Sbjct: 390 CMEN*MRKFLLINHRVM*RRVIS*RVKRALYGLKQAPRAWYSRIEAYFTKEGFEKCPYEH 569
Query: 1152 TLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQNLRKELAVQFEMKDLGKLKYFLGMEVA 1211
TLF+K + GK+ ++ +YV+D+I GNDE + +K + +F M DLGK+ YFLG+EV
Sbjct: 570 TLFVKLSEGGKILIISLYVDDLIFIGNDENMFEEFKKSMKKEFNMSDLGKMHYFLGVEVT 749
Query: 1212 YSKQGIFI 1219
+++GI+I
Sbjct: 750 QNEKGIYI 773
>BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2O9.150 -
Arabidopsis thaliana, partial (11%)
Length = 732
Score = 174 bits (441), Expect = 2e-43
Identities = 87/217 (40%), Positives = 135/217 (62%)
Frame = +1
Query: 1208 MEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPIEQNHKIGSSEEDLRVHKAQYQRL 1267
+EV +++GI+I QRKYV DLL+ G PI K+ E ++V +Y+++
Sbjct: 1 VEVIQNEEGIYICQRKYVTDLLERFGMEKSNLSRNPIAPRCKLIKDENGVKVDATKYKQI 180
Query: 1268 VGKLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQAVDRILQYLKASPGRGLLFKKSEQLS 1327
VG L+YL+ TRPD+ Y +S++S+FM+ P E H+ AV R+L+YL + G+++K++
Sbjct: 181 VGCLMYLAATRPDLMYVLSLISRFMNCPTELHMHAVKRVLRYLNGTINLGIMYKRNGSEK 360
Query: 1328 MEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQNVVARSSAEAEFRAMAQGVCEL 1387
+E YTD+DYAG + DR+ST+GY L V+W SKKQ VV S+ +AEF A A C+
Sbjct: 361 LEAYTDSDYAGDLDDRKSTSGYVFMLSSGAVSWSSKKQPVVTLSTTKAEFIAAAFCACQS 540
Query: 1388 LRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQH 1424
+ M+ +L+ L + ++CDN S I ++ NPV H
Sbjct: 541 VWMRRVLEKLGYTQSGSITMYCDNNSTIKLSKNPVLH 651
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 137 bits (346), Expect(2) = 3e-41
Identities = 65/142 (45%), Positives = 102/142 (71%)
Frame = -2
Query: 1032 LDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTVRIILSLAAHFDWELQQFDVKNAFLHG 1091
+ R K++LV +GY Q GIDYDE F+PVA+M +RI+++ AA ++L Q DVK+AF++G
Sbjct: 424 ITRNKSKLVVQGYNQKEGIDYDEAFSPVARMEAIRILIAFAAFMGFKLYQMDVKSAFING 245
Query: 1092 DLEEEVYMEIPPGYDPTSGRNKVCKLKKALYGLKQSPRAWFGRFTNAMVSLGYKQSQGDH 1151
DL+EEV+++ PPG++ N V +L K LYGLKQ+PRAW+ R + ++ G+K+ + D+
Sbjct: 244 DLKEEVFVKQPPGFEDAEVPNHVFRLNKTLYGLKQAPRAWYERLSKFLLKNGFKRGKIDN 65
Query: 1152 TLFIKHTRNGKLTLLLVYVNDM 1173
TLF+ R +L ++ VYV+D+
Sbjct: 64 TLFLL-KRE*ELLIIQVYVDDI 2
Score = 50.8 bits (120), Expect(2) = 3e-41
Identities = 22/66 (33%), Positives = 39/66 (58%)
Frame = -3
Query: 960 VQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRKIVEKPNDKRTVGCK 1019
V +FIS+I+ P +V+E L+D WI +M EE+H +++ +V +P K +G +
Sbjct: 618 VSFSAFISSIE----PKNVKEALRDADWINSMQEELHQFERSKVWYLVPRPEGKTVIGTR 451
Query: 1020 WIYTVK 1025
W++ K
Sbjct: 450 WVFRNK 433
>TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative retroelement
{Oryza sativa} [Oryza sativa (japonica cultivar-group)],
partial (10%)
Length = 823
Score = 163 bits (412), Expect = 5e-40
Identities = 95/254 (37%), Positives = 143/254 (55%), Gaps = 4/254 (1%)
Frame = +1
Query: 555 FSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVTFIDDCTRITWVFLMKD 614
F + +F + ++ D +HSD+WGPS +++ G ++ +T IDD R WV+ ++
Sbjct: 40 FGNRKKVSFSTATHRTKGILDYIHSDLWGPSKVTSYGGRRYMMTIIDDFPRKVWVYFLRY 219
Query: 615 KSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLTKNGVVHELTCVDAPQQ 674
K+E F F + ++ TQ GK +K+L +DN E + +F+EF T +G+ T PQQ
Sbjct: 220 KNETFPTFKKWRILVETQTGKNVKKLITDN*LEFCSSDFNEFCTNHGIARHKTIPRNPQQ 399
Query: 675 NGVAERKNRHLLEITRALLFQMNV--PKFYWGEAVLTAAYLINRLPSRVLSNVSPVLVMT 732
NGVAER R LLE R +L + + W EA TA +L+NR P L P + +
Sbjct: 400 NGVAERMIRTLLERARCMLSNAGL*N*RDLWVEAASTACHLVNRSPHSALDFKVPEDIWS 579
Query: 733 SFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYASDKKGYK--CYHPP 790
++ S L R+FGC A+ V+ GKL PRA +CIF+ YAS+ KGY+ C P
Sbjct: 580 G---NLVDYSNL--RIFGCPAYALVND---GKLAPRAGECIFLSYASESKGYRLWCSDPK 735
Query: 791 SRRVYISMDVTFQE 804
S+++ +S DVTF E
Sbjct: 736 SQKLILSRDVTFNE 777
>BG587170 similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 718
Score = 161 bits (408), Expect = 2e-39
Identities = 94/248 (37%), Positives = 136/248 (53%), Gaps = 4/248 (1%)
Frame = -3
Query: 479 LATGQIIGIAKEQDGLYCLQHEES----KCHQTSSESWNTSQIWLRHKRLGHPPFSILRT 534
+ + Q+IG + LY L+ + KC TSS S N +W H RLGHP L
Sbjct: 710 IESSQLIGEGVTKGDLYMLEKLDPVSNYKCSFTSSSSLNKDALW--HARLGHPHGRALNL 537
Query: 535 MFPHLFSKVSVESFHCDVCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAK 594
M P V E+ +C+ C KH ++ F ++ FDL+++D+W S+S K
Sbjct: 536 MLPG----VVFENKNCEACILGKHCKNVFPRTSTVYENCFDLIYTDLWTAPSLSR-DNHK 372
Query: 595 WFVTFIDDCTRITWVFLMKDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFS 654
+FVTFID+ ++ TW+ L+ K V F NF + + IK LRSDNG E + F
Sbjct: 371 YFVTFIDEKSKYTWLTLIPSKDRVIDAFKNFQAYVTNHYHAKIKILRSDNGGEYTSYAFK 192
Query: 655 EFLTKNGVVHELTCVDAPQQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLI 714
L +G++H+ +C PQQNGVA+RKN+HL+E+ R+L+FQ NV TA YLI
Sbjct: 191 SHLDHHGILHQTSCPYTPQQNGVAKRKNKHLMEVARSLMFQANVS---------TACYLI 39
Query: 715 NRLPSRVL 722
N +P++VL
Sbjct: 38 NWIPTKVL 15
>BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 503
Score = 127 bits (318), Expect = 4e-29
Identities = 67/130 (51%), Positives = 86/130 (65%)
Frame = +1
Query: 1126 QSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQN 1185
QSPR WF RFT + GY Q Q DH +FIKH+ K +L+VYV+D+ + G+ +
Sbjct: 1 QSPRDWFDRFT*VVKKFGYIQCQTDHAMFIKHSSTVKKAILIVYVDDIFLTGDHGK*IKR 180
Query: 1186 LRKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPIE 1245
L+ LA +FE+KDLG LKYFLGMEVA K+G ISQRKYVLDLLKET +GCK + P
Sbjct: 181 LKNLLAEEFEIKDLGNLKYFLGMEVARWKKGSSISQRKYVLDLLKETRMIGCKTIRDPYG 360
Query: 1246 QNHKIGSSEE 1255
N + +S +
Sbjct: 361 CNCEARNSRQ 390
Score = 64.7 bits (156), Expect = 3e-10
Identities = 31/58 (53%), Positives = 40/58 (68%)
Frame = +2
Query: 1235 LGCKPMGVPIEQNHKIGSSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFM 1292
L KP P++ K+G+ + V K +YQRLVGKLIYLSHTRPDI++ V +SQFM
Sbjct: 329 LDVKPSETPMDATVKLGTLDNGTLVDKGRYQRLVGKLIYLSHTRPDISFVVCTMSQFM 502
>BG453259 homologue to GP|21434|emb|CA ORF4 {Solanum tuberosum}, partial (5%)
Length = 657
Score = 125 bits (313), Expect = 2e-28
Identities = 75/143 (52%), Positives = 95/143 (65%), Gaps = 2/143 (1%)
Frame = -2
Query: 1306 ILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQ 1365
+++ L +S + ++FK+ +LSMEVY + AG IVDR ST+GY MFLGGN+V KQ
Sbjct: 566 LIEGLXSSGTQMIVFKRE*KLSMEVYXNTVCAGWIVDRGSTSGY*MFLGGNMV---E*KQ 396
Query: 1366 NVVARSSAEAEFRAMAQGVCELL--RMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQ 1423
NVVAR F +QG+ ++ +KI LDDL I Y+ PM LF +N IAHNPVQ
Sbjct: 395 NVVAR*VQRHNFELCSQGL*RVMDEELKIKLDDLIINYKDPMTLF*NNNFVSRIAHNPVQ 216
Query: 1424 HDRTKHIEIDRHFIKEKLNSSLI 1446
H RTKHIEID+HFI EKL S LI
Sbjct: 215 HYRTKHIEIDQHFIIEKLYSGLI 147
>AJ502495 weakly similar to GP|18071369|g putative gag-pol polyprotein {Oryza
sativa}, partial (9%)
Length = 542
Score = 117 bits (294), Expect = 3e-26
Identities = 59/147 (40%), Positives = 93/147 (63%)
Frame = +2
Query: 1334 ADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQNVVARSSAEAEFRAMAQGVCELLRMKII 1393
+D+AG R+ST+GY LG ++W SKKQ VVA S+AEAE+ A + + ++ I
Sbjct: 2 SDWAGDTETRKSTSGYAFHLGTGAISWSSKKQPVVAFSTAEAEYIASTSCATQTVWLRRI 181
Query: 1394 LDDLKIKYEAPMRLFCDNKSAISIAHNPVQHDRTKHIEIDRHFIKEKLNSSLISTQYVPS 1453
L+ + + P +++CDNKSAI+++ NPV H R+KHI+I H I+E + + +Y P+
Sbjct: 182 LEVMHHEQNTPTKIYCDNKSAIALSKNPVFHGRSKHIDIQFHKIRELIAEKEVVIEYCPT 361
Query: 1454 GLQLADVLTKGLPLERFRELTCKLGMI 1480
++AD+ TK L +E F +L LGM+
Sbjct: 362 EEKIADIFTKPLKIESFYKLKKMLGMM 442
>BE239977 weakly similar to GP|23237899|db polyprotein-like {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 514
Score = 113 bits (282), Expect = 6e-25
Identities = 50/96 (52%), Positives = 68/96 (70%), Gaps = 1/96 (1%)
Frame = -2
Query: 730 VMTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYASDKKGYKCYHP 789
V+ SF+P VP + L R+FGC++FVH+H R K D RA+KC+FI Y+S +KGY+CYHP
Sbjct: 507 VLPSFYPYVPTSNNLTPRIFGCTSFVHIHSDGRSKFDHRALKCVFIRYSSTQKGYRCYHP 328
Query: 790 PSRRVYISMDVTFQESESFY-PNPQLQGGDTQEAES 824
PSR+ ++S DVTF E ES++ N GG +E ES
Sbjct: 327 PSRKYFVSRDVTFHEQESYFGQN*SSSGGKYKEDES 220
>TC89912 weakly similar to PIR|B84512|B84512 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(10%)
Length = 814
Score = 112 bits (279), Expect = 1e-24
Identities = 61/165 (36%), Positives = 105/165 (62%), Gaps = 2/165 (1%)
Frame = +1
Query: 1301 QAVDRILQYLKASPGRGLLFKKS--EQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLV 1358
QA+ +L+YL S L + K+ E+ ++E Y DADYAG++ R+S +G+ L G +
Sbjct: 1 QALKWVLKYLNESLKSSLKYTKAAQEEDALEGYVDADYAGNVDTRKSLSGFVFTLYGTTI 180
Query: 1359 TWRSKKQNVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIA 1418
+W++ +Q+VV S+ +AE+ A +GV + + +K ++ +L I E +++ CD++SAI +A
Sbjct: 181 SWKANQQSVVTLSTTQAEYIAFVEGVKDAIWLKGMIGELGITQEY-VKIHCDSQSAIHLA 357
Query: 1419 HNPVQHDRTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTK 1463
++ V H+RTKHI+I HFI++ + S I + + S ADV TK
Sbjct: 358 NHQVYHERTKHIDIRLHFIRDMIESKEIVVEKMASEENPADVFTK 492
>TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
(EC 3.4.23.-);, partial (7%)
Length = 705
Score = 66.6 bits (161), Expect(2) = 4e-24
Identities = 36/77 (46%), Positives = 49/77 (62%)
Frame = +1
Query: 1303 VDRILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRS 1362
V RI++Y+K + G + F SE L++ Y D+D+AG R+STTGY L G V+W S
Sbjct: 1 VKRIMRYIKGTSGVAVCFGGSE-LTVRGYVDSDFAGDHDKRKSTTGYVFTLAGGAVSWLS 177
Query: 1363 KKQNVVARSSAEAEFRA 1379
K Q VVA S+ EAE+ A
Sbjct: 178 KLQTVVALSTTEAEYMA 228
Score = 64.7 bits (156), Expect(2) = 4e-24
Identities = 32/103 (31%), Positives = 59/103 (57%)
Frame = +3
Query: 1379 AMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHDRTKHIEIDRHFIK 1438
++ Q E + M+ ++++L K E + ++CD++SA+ IA NP H RTKHI I HF++
Sbjct: 228 SLPQACKEAIWMQRLMEELGHKQEQ-ITVYCDSQSALHIARNPAFHSRTKHIGIQYHFVR 404
Query: 1439 EKLNSSLISTQYVPSGLQLADVLTKGLPLERFRELTCKLGMID 1481
E + + Q + + LAD +TK + ++F G+++
Sbjct: 405 EVVEEGSVDMQKIHTNDNLADAMTKSINTDKFIWCRSSYGLLE 533
>BG648737 weakly similar to GP|21433|emb|CA ORF3 {Solanum tuberosum}, partial
(10%)
Length = 804
Score = 105 bits (261), Expect = 2e-22
Identities = 79/267 (29%), Positives = 129/267 (47%), Gaps = 31/267 (11%)
Frame = +3
Query: 784 YKCYHPPSRRVYISMDVTFQESESFYPNPQLQGGD-TQEAESPELSVIPLLQ------EP 836
YKCY P +++ Y SMDVTF E + FY +QG T+E + ++ V+ + P
Sbjct: 3 YKCYSPITKKFYNSMDVTFFEDQPFYTKIGIQGESFTEEYQLWDIKVLESIHGENSDVSP 182
Query: 837 LMPSSIPITED----------SDDDDNGPEQVP--NQ----NDDNGLELVPDQNDDNGPE 880
+ + PI SD + P P NQ N + + + +Q +
Sbjct: 183 QVNQTNPIVSYHLSSKIPPPVSDQPNIQPSIFPMSNQFDLSNSQSTIMPLSEQQPPSQRS 362
Query: 881 LVPDQNNVDRFRI---KYQRKVKPVLIQQQD-QPSDPEVSVPNPEPSNSYEH----NLDD 932
+ FR+ +Y+++ + Q QD S+ + N + + E DD
Sbjct: 363 TSTQPAEICDFRVFTTRYKKQEGKRMQQIQDCSDSNSNFEIENTQDNEICEPIPTVEFDD 542
Query: 933 LPIALRKGKRSCAKYPISQYVSTQNLSVQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMN 992
I+LRK KRSC +PI +++S LS H +F+S +D I++P SV E L + W +A+
Sbjct: 543 RLISLRKDKRSCTSHPICKFISYDKLSPHHLAFVSNLDKIQIPNSVHESLTNP*WKKAIL 722
Query: 993 EEMHALDKNGTRKIVEKPNDKRTVGCK 1019
E+HAL+KN T ++ + P+ K +GCK
Sbjct: 723 GEIHALEKNETWELSDLPSGKHPMGCK 803
>BI262917 weakly similar to GP|19920130|g Putative retroelement {Oryza
sativa} [Oryza sativa (japonica cultivar-group)],
partial (8%)
Length = 426
Score = 103 bits (257), Expect = 5e-22
Identities = 58/138 (42%), Positives = 77/138 (55%)
Frame = +1
Query: 652 NFSEFLTKNGVVHELTCVDAPQQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAA 711
+F F+ K + PQQNGVAER NR LLE TRA+L + K +W EAV TA
Sbjct: 28 SF*HFVNKRVL*GNSXVAHTPQQNGVAERMNRTLLERTRAMLKTAGMAKSFWAEAVKTAC 207
Query: 712 YLINRLPSRVLSNVSPVLVMTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIK 771
Y+INR PS V+ +P+ + P+ VFGC +V + R KLDP++ K
Sbjct: 208 YVINRSPSTVIDLKTPM----EMWKGKPVDYS-SLHVFGCPVYVMYNSQERTKLDPKSRK 372
Query: 772 CIFIGYASDKKGYKCYHP 789
CIF+GYA + KGY + P
Sbjct: 373 CIFLGYADNVKGYXLWDP 426
>BG586273 weakly similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 705
Score = 99.4 bits (246), Expect = 9e-21
Identities = 61/177 (34%), Positives = 92/177 (51%)
Frame = -2
Query: 710 AAYLINRLPSRVLSNVSPVLVMTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRA 769
A YLINR+P+RVL + +P V+ PS+ RVFGC +V V G R KL+ R+
Sbjct: 704 ACYLINRIPTRVLKDQAPFEVLNQRKPSLTYM-----RVFGCLCYVLVPGELRNKLEARS 540
Query: 770 IKCIFIGYASDKKGYKCYHPPSRRVYISMDVTFQESESFYPNPQLQGGDTQEAESPELSV 829
K +FIGY++ +KGYKCY P +RRV +S DV F E +Y + D ++ S + V
Sbjct: 539 RKAMFIGYSTTQKGYKCYDPEARRVLVSRDVKFIEERGYYEEKNQE--DLRDLTSDKAGV 366
Query: 830 IPLLQEPLMPSSIPITEDSDDDDNGPEQVPNQNDDNGLELVPDQNDDNGPELVPDQN 886
+ ++ E L I + +D PE+ + N+ P + + G E +N
Sbjct: 365 LRVILEGL---GIKMNQDQSTRSRQPEE--SSNEPRRAAQTPHLDHEGGSEPETQEN 210
>BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vulgaris},
partial (13%)
Length = 494
Score = 95.5 bits (236), Expect = 1e-19
Identities = 53/124 (42%), Positives = 77/124 (61%), Gaps = 3/124 (2%)
Frame = +1
Query: 1278 RPDIAYPVSVVSQFMHDPQERHLQAVDRILQYLKASPGRGLLF---KKSEQLSMEVYTDA 1334
RPDI Y VSV+S+FMHDP++ HL A +RIL+Y++ + GLLF KSE + Y+D+
Sbjct: 121 RPDICYSVSVISKFMHDPRKPHLIAANRILRYVRGTMEYGLLFPYGAKSEVYELICYSDS 300
Query: 1335 DYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQNVVARSSAEAEFRAMAQGVCELLRMKIIL 1394
D+ G DRRST+GY ++W +KKQ + A SS EAE+ A + L + ++
Sbjct: 301 DWCG---DRRSTSGYVFKFNDAAISWCTKKQPITALSSYEAEYIAGTFATFQALWLDSVI 471
Query: 1395 DDLK 1398
+LK
Sbjct: 472 KELK 483
>BG586293 weakly similar to PIR|E84473|E84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(7%)
Length = 763
Score = 94.7 bits (234), Expect = 2e-19
Identities = 45/89 (50%), Positives = 62/89 (69%)
Frame = +2
Query: 1003 TRKIVEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKM 1062
T K+V+KP + +G +WIY +K DGTL +YKARLVAKGY + GID+DE FAPV ++
Sbjct: 56 TLKLVKKPTGVKPIGLRWIYKIKRNEDGTLIKYKARLVAKGYVKQQGIDFDEVFAPVVRI 235
Query: 1063 NTVRIILSLAAHFDWELQQFDVKNAFLHG 1091
T+ ++L+LAA + DVK AFL+G
Sbjct: 236 ETI*LLLALAATNGC*IHHIDVKIAFLNG 322
>AW684891
Length = 539
Score = 73.6 bits (179), Expect(3) = 1e-18
Identities = 41/94 (43%), Positives = 57/94 (60%)
Frame = -3
Query: 586 SISNVSGAKWFVTFIDDCTRITWVFLMKDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNG 645
++ + G KWF+ FI + T IT +FL K H I QF K I+RL S+ G
Sbjct: 366 TLIQIYGXKWFLAFIHNSTSIT*LFLKK-------------HEIWFQFVKSIERLCSNKG 226
Query: 646 KE*VNKNFSEFLTKNGVVHELTCVDAPQQNGVAE 679
K V++N S++L +NGV EL CV++PQQNGV+E
Sbjct: 225 KVYVDQNLSQYLEENGVSQELKCVNSPQQNGVSE 124
Score = 38.5 bits (88), Expect(3) = 1e-18
Identities = 15/30 (50%), Positives = 20/30 (66%)
Frame = -2
Query: 540 FSKVSVESFHCDVCQFSKHHRSTFLPSNNK 569
F+K +E FHCDV Q + HH +F PS +K
Sbjct: 463 FTKELIEYFHCDVFQLAXHHPESFFPSKDK 374
Score = 20.8 bits (42), Expect(3) = 1e-18
Identities = 8/12 (66%), Positives = 10/12 (82%)
Frame = -2
Query: 690 RALLFQMNVPKF 701
R L+FQ+ VPKF
Sbjct: 127 RVLIFQIFVPKF 92
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.135 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 47,375,558
Number of Sequences: 36976
Number of extensions: 721272
Number of successful extensions: 3927
Number of sequences better than 10.0: 83
Number of HSP's better than 10.0 without gapping: 3797
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3888
length of query: 1485
length of database: 9,014,727
effective HSP length: 108
effective length of query: 1377
effective length of database: 5,021,319
effective search space: 6914356263
effective search space used: 6914356263
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0310b.12


