

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0306.6
(510 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
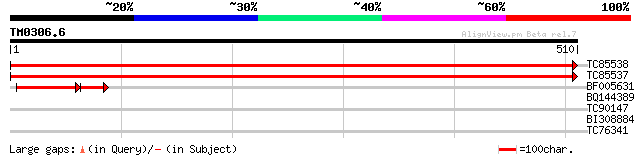
TC85538 homologue to GP|14764466|gb|AAK72098.1 myo-inositol-1-ph... 975 0.0
TC85537 homologue to GP|14764466|gb|AAK72098.1 myo-inositol-1-ph... 965 0.0
BF005631 similar to GP|14582467|gb| 1L-myo-inositol-1-phosphate ... 111 5e-34
BQ144389 weakly similar to GP|21322711|em pherophorin-dz1 protei... 33 0.26
TC90147 similar to GP|21537059|gb|AAM61400.1 pectate lyase-like ... 30 2.9
BI308884 weakly similar to GP|22136008|gb unknown protein {Arabi... 29 3.8
TC76341 similar to GP|7528270|gb|AAF63202.1| poly(A)-binding pro... 28 6.4
>TC85538 homologue to GP|14764466|gb|AAK72098.1 myo-inositol-1-phosphate
synthase {Glycine max}, complete
Length = 1862
Score = 975 bits (2521), Expect = 0.0
Identities = 483/510 (94%), Positives = 501/510 (97%)
Frame = +1
Query: 1 MFIENFKVESPNVKYTETEIQSVYNYETTELVHENRNGSYQWIVKPKSVKYEFKTSTNVP 60
MFIENFKVESPNVKYTETEIQSVYNYETTELVHENRNG+YQWIVKPK+VKYEFKT +VP
Sbjct: 100 MFIENFKVESPNVKYTETEIQSVYNYETTELVHENRNGTYQWIVKPKTVKYEFKTDIHVP 279
Query: 61 KLGVMLVGWGGNNGSTLTGGVIANKEGISWATKDKIERANYFGSLTQASAIRVGSFQGEE 120
KLGVMLVGWGGNNGSTLTGGVIAN+EGISWATKDKI++ANYFGSLTQASAIRVGSFQGEE
Sbjct: 280 KLGVMLVGWGGNNGSTLTGGVIANREGISWATKDKIQQANYFGSLTQASAIRVGSFQGEE 459
Query: 121 IYAPFKSLLPMVNPDDVVFGGWDISDMNLADAMGRARVFDIDLQKQLRPYMESMVPLPGI 180
I+APFKSLLPMVNPDD+VFGGWDISDMNLADAM RARVFDIDLQKQLRPYMESMVPLPGI
Sbjct: 460 IHAPFKSLLPMVNPDDIVFGGWDISDMNLADAMARARVFDIDLQKQLRPYMESMVPLPGI 639
Query: 181 YDPDFIAANQGDRANNVIKGTKKEQIQQIIKDIKEFKEASKVDRVVVLWTANTERYSNLV 240
YDPDFIAANQG+RANNVIKGTK+EQI QIIKDIKEFKEA+KVDRVVVLWTANTERYSNLV
Sbjct: 640 YDPDFIAANQGERANNVIKGTKREQINQIIKDIKEFKEANKVDRVVVLWTANTERYSNLV 819
Query: 241 VGLNDTMENLLASVDRNEAEISPSTLYAIACVMENIPFINGSPQNTFVPGLIDLAIKKNS 300
VGLNDTMENL A+VDRNE+EISPSTL+AIACVMEN+PFINGSPQNTFVPGLIDLAIK N
Sbjct: 820 VGLNDTMENLFAAVDRNESEISPSTLFAIACVMENVPFINGSPQNTFVPGLIDLAIKNNC 999
Query: 301 LIGGDDFKSGQTKMKSVLVDFLVGAGIKPTSIVSYNHLGNNDGMNLSAPQTFRSKEISKS 360
LIGGDDFKSGQTKMKSVLVDFLVGAGIKPTSIVSYNHLGNNDGMNLSAPQTFRSKEISKS
Sbjct: 1000LIGGDDFKSGQTKMKSVLVDFLVGAGIKPTSIVSYNHLGNNDGMNLSAPQTFRSKEISKS 1179
Query: 361 NVVDDMVNSNAILYEPGEHPDHVVVIKYVPYVADSKRAMDEYTSEIFMGGKNTIVLHNTC 420
NVVDDMVNSNAILY PGEHPDHVVVIKYVPYV DSKRAMDEYTSEIFMGGKNTIVLHNTC
Sbjct: 1180NVVDDMVNSNAILYAPGEHPDHVVVIKYVPYVGDSKRAMDEYTSEIFMGGKNTIVLHNTC 1359
Query: 421 EDSLLAAPIILDLVLLAELSTRIQFKSEAEDKFHSFHPVATILSYLTKAPLVPPGTPVVN 480
EDSLLAAPIILDLVLLAELSTRIQFKSEAE+KFH+FHPVATILSYLTKAPLVPPGTPVVN
Sbjct: 1360EDSLLAAPIILDLVLLAELSTRIQFKSEAENKFHTFHPVATILSYLTKAPLVPPGTPVVN 1539
Query: 481 ALSKQRAMLENILRACVGLAPENNMILEYK 510
ALSKQRAMLENI+RACVGLAPENNMILEYK
Sbjct: 1540ALSKQRAMLENIMRACVGLAPENNMILEYK 1629
>TC85537 homologue to GP|14764466|gb|AAK72098.1 myo-inositol-1-phosphate
synthase {Glycine max}, complete
Length = 1817
Score = 965 bits (2495), Expect = 0.0
Identities = 473/510 (92%), Positives = 502/510 (97%)
Frame = +1
Query: 1 MFIENFKVESPNVKYTETEIQSVYNYETTELVHENRNGSYQWIVKPKSVKYEFKTSTNVP 60
MFIENFKVESPNVKYTETEIQSVY+YETTELVHENRN +Y+W+VKPK+VKYEFKT TNVP
Sbjct: 85 MFIENFKVESPNVKYTETEIQSVYSYETTELVHENRNNTYEWVVKPKTVKYEFKTQTNVP 264
Query: 61 KLGVMLVGWGGNNGSTLTGGVIANKEGISWATKDKIERANYFGSLTQASAIRVGSFQGEE 120
KLGVMLVGWGGNNGSTLTGGVIAN+EGISWATKDKI++ANYFGSLTQASAIRVGSFQGEE
Sbjct: 265 KLGVMLVGWGGNNGSTLTGGVIANREGISWATKDKIQQANYFGSLTQASAIRVGSFQGEE 444
Query: 121 IYAPFKSLLPMVNPDDVVFGGWDISDMNLADAMGRARVFDIDLQKQLRPYMESMVPLPGI 180
I+APFKSLLPMVNP+D+V+GGWDIS++NLADAMGRARVFDIDLQKQLRPYMESMVPLPGI
Sbjct: 445 IHAPFKSLLPMVNPEDIVWGGWDISNLNLADAMGRARVFDIDLQKQLRPYMESMVPLPGI 624
Query: 181 YDPDFIAANQGDRANNVIKGTKKEQIQQIIKDIKEFKEASKVDRVVVLWTANTERYSNLV 240
YDPDFIAANQGDRANN+IKGTKKEQ+QQ+IKDIKEFKE +KVD+VVVLWTANTERYSN+V
Sbjct: 625 YDPDFIAANQGDRANNIIKGTKKEQLQQVIKDIKEFKETTKVDKVVVLWTANTERYSNIV 804
Query: 241 VGLNDTMENLLASVDRNEAEISPSTLYAIACVMENIPFINGSPQNTFVPGLIDLAIKKNS 300
VGLNDT ENLLASVD+NEAEISPSTLYA+ACV+EN+PFINGSPQNTFVPGLIDLAIK+NS
Sbjct: 805 VGLNDTTENLLASVDKNEAEISPSTLYALACVLENVPFINGSPQNTFVPGLIDLAIKRNS 984
Query: 301 LIGGDDFKSGQTKMKSVLVDFLVGAGIKPTSIVSYNHLGNNDGMNLSAPQTFRSKEISKS 360
LIGGDDFKSGQTK+KSVLVDFLVGAGIKPTSIVSYNHLGNNDGMNLSAPQTFRSKEISKS
Sbjct: 985 LIGGDDFKSGQTKIKSVLVDFLVGAGIKPTSIVSYNHLGNNDGMNLSAPQTFRSKEISKS 1164
Query: 361 NVVDDMVNSNAILYEPGEHPDHVVVIKYVPYVADSKRAMDEYTSEIFMGGKNTIVLHNTC 420
NVVDDMVNSNAILY PGEHPDHVVVIKYVPYV DSKRAMDEYTSEIFMGGKNTIV+HNTC
Sbjct: 1165NVVDDMVNSNAILYGPGEHPDHVVVIKYVPYVGDSKRAMDEYTSEIFMGGKNTIVMHNTC 1344
Query: 421 EDSLLAAPIILDLVLLAELSTRIQFKSEAEDKFHSFHPVATILSYLTKAPLVPPGTPVVN 480
EDSLLAAPIILDLVLLAELSTRIQFKSE EDKFHSFH VATILSYL+KAPLVPPGTPVVN
Sbjct: 1345EDSLLAAPIILDLVLLAELSTRIQFKSEQEDKFHSFHAVATILSYLSKAPLVPPGTPVVN 1524
Query: 481 ALSKQRAMLENILRACVGLAPENNMILEYK 510
ALSKQR+MLENILRACVGLAPENNMILEYK
Sbjct: 1525ALSKQRSMLENILRACVGLAPENNMILEYK 1614
>BF005631 similar to GP|14582467|gb| 1L-myo-inositol-1-phosphate synthase
{Phaseolus vulgaris}, partial (16%)
Length = 444
Score = 111 bits (277), Expect(2) = 5e-34
Identities = 50/57 (87%), Positives = 55/57 (95%)
Frame = +2
Query: 7 KVESPNVKYTETEIQSVYNYETTELVHENRNGSYQWIVKPKSVKYEFKTSTNVPKLG 63
KVESPNVKYTETEIQSVY+YETTELVHENRN +Y+W+VKPK+VKYEFKT TNVPKLG
Sbjct: 2 KVESPNVKYTETEIQSVYSYETTELVHENRNNTYEWVVKPKTVKYEFKTQTNVPKLG 172
Score = 51.6 bits (122), Expect(2) = 5e-34
Identities = 23/26 (88%), Positives = 25/26 (95%)
Frame = +3
Query: 64 VMLVGWGGNNGSTLTGGVIANKEGIS 89
VMLVGWGGNNGSTLTGGVIAN+E +S
Sbjct: 279 VMLVGWGGNNGSTLTGGVIANRELVS 356
>BQ144389 weakly similar to GP|21322711|em pherophorin-dz1 protein {Volvox
carteri f. nagariensis}, partial (4%)
Length = 1283
Score = 33.1 bits (74), Expect = 0.26
Identities = 25/85 (29%), Positives = 41/85 (47%)
Frame = -2
Query: 399 MDEYTSEIFMGGKNTIVLHNTCEDSLLAAPIILDLVLLAELSTRIQFKSEAEDKFHSFHP 458
+D S I++ + L+ + SLL A L+L LS Q S H+FH
Sbjct: 1183 LDSLLSRIYLSRRAARPLYVSSALSLLLA-----LLLALSLSIFFQLVSLRITTAHAFHT 1019
Query: 459 VATILSYLTKAPLVPPGTPVVNALS 483
+++ILS ++ A +P P ++A S
Sbjct: 1018 ISSILSLISLAAPLPLPAPTISAHS 944
>TC90147 similar to GP|21537059|gb|AAM61400.1 pectate lyase-like protein
{Arabidopsis thaliana}, partial (55%)
Length = 1093
Score = 29.6 bits (65), Expect = 2.9
Identities = 13/29 (44%), Positives = 16/29 (54%)
Frame = +3
Query: 268 AIACVMENIPFINGSPQNTFVPGLIDLAI 296
A++ V EN PFIN P N P L L +
Sbjct: 66 ALSSVKENFPFINNVPNNLHFPNLTTLLL 152
>BI308884 weakly similar to GP|22136008|gb unknown protein {Arabidopsis
thaliana}, partial (24%)
Length = 659
Score = 29.3 bits (64), Expect = 3.8
Identities = 22/68 (32%), Positives = 32/68 (46%), Gaps = 7/68 (10%)
Frame = -3
Query: 419 TCEDSLLAAPIILDLVLLAELSTRIQFKSEAEDKFHSFHPVATILSYLT-------KAPL 471
TCE+S+ P ILD E S+R+ +A+ F SF LS+ T + L
Sbjct: 405 TCEESISYTPFILD---SEEWSSRLGTTVKADLSFESFRLAFLTLSF*TYFFFSDPRLVL 235
Query: 472 VPPGTPVV 479
+ PG P +
Sbjct: 234 LGPGLPCI 211
>TC76341 similar to GP|7528270|gb|AAF63202.1| poly(A)-binding protein {Cucumis
sativus}, partial (90%)
Length = 2501
Score = 28.5 bits (62), Expect = 6.4
Identities = 12/32 (37%), Positives = 17/32 (52%)
Frame = +3
Query: 446 KSEAEDKFHSFHPVATILSYLTKAPLVPPGTP 477
++ + +F PVA S + PL PPGTP
Sbjct: 1389 RARLQAQFSQMRPVAITPSVAPRMPLYPPGTP 1484
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.134 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,920,612
Number of Sequences: 36976
Number of extensions: 155473
Number of successful extensions: 700
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 698
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 700
length of query: 510
length of database: 9,014,727
effective HSP length: 100
effective length of query: 410
effective length of database: 5,317,127
effective search space: 2180022070
effective search space used: 2180022070
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0306.6


