

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0302a.8
(578 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
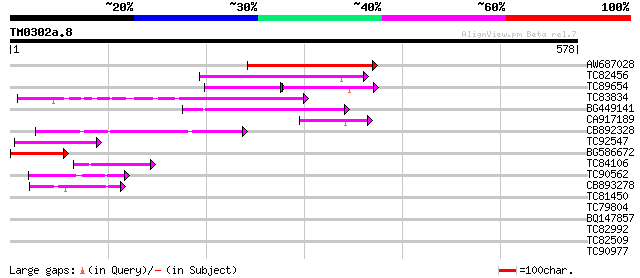
Sequences producing significant alignments: (bits) Value
AW687028 weakly similar to GP|9909176|dbj| hypothetical protein~... 143 2e-34
TC82456 weakly similar to GP|18057159|gb|AAL58182.1 putative far... 125 6e-29
TC89654 weakly similar to PIR|T05644|T05644 hypothetical protein... 73 1e-25
TC83834 similar to PIR|G96565|G96565 F6D8.26 [imported] - Arabid... 90 3e-18
BG449141 similar to GP|15983442|gb At1g10240/F14N23_12 {Arabidop... 89 5e-18
CA917189 similar to PIR|T05645|T05 hypothetical protein F20D10.3... 64 1e-10
CB892328 GP|15075873|em PROBABLE GLYCOSYL HYDROLASE PROTEIN {Sin... 64 2e-10
TC92547 similar to PIR|A56235|A56235 transcription activator Maf... 64 2e-10
BG586672 54 2e-07
TC84106 similar to GP|9502168|gb|AAF88018.1| contains simlarity ... 53 3e-07
TC90562 similar to GP|16604382|gb|AAL24197.1 AT3g59470/T16L24_20... 49 4e-06
CB893278 weakly similar to PIR|T05645|T056 hypothetical protein ... 47 2e-05
TC81450 similar to GP|16604382|gb|AAL24197.1 AT3g59470/T16L24_20... 40 0.002
TC79804 weakly similar to GP|6175165|gb|AAF04891.1| Mutator-like... 39 0.005
BQ147857 weakly similar to GP|15983442|gb At1g10240/F14N23_12 {A... 37 0.021
TC82992 homologue to PIR|T05645|T05645 hypothetical protein F20D... 32 0.67
TC82509 similar to GP|15293233|gb|AAK93727.1 unknown protein {Ar... 28 7.4
TC90977 similar to GP|10178227|dbj|BAB11607. ZW10-like protein {... 28 9.7
>AW687028 weakly similar to GP|9909176|dbj| hypothetical protein~similar to
Arabidopsis thaliana F20D10.300, partial (11%)
Length = 662
Score = 143 bits (360), Expect = 2e-34
Identities = 64/133 (48%), Positives = 89/133 (66%)
Frame = +1
Query: 243 DVVAFDATYGKNRYKCPFVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFLEAMKGK 302
DV+AFD TY KN+Y P +F G NHH+++ IF AL+ +E IE+Y W+L RFLE M+ K
Sbjct: 1 DVLAFDTTYKKNKYNYPLCIFSGCNHHSQTIIFGVALLEDETIESYKWVLNRFLECMENK 180
Query: 303 APLFVITDGDKAMKAAIKQVFPNAHHRLCAWHILDNARKHVHIRAFNTELRKCMFMELDV 362
P V+TDGD +M+ AIKQVFP+A HRLCAWH+ NA++++ F RK M+
Sbjct: 181 FPKAVVTDGDGSMREAIKQVFPDASHRLCAWHLHKNAQENIKKTPFLEGFRKAMYSNFTP 360
Query: 363 SEFEEHWAATVAK 375
+FE+ W+ + K
Sbjct: 361 EQFEDFWSELIQK 399
>TC82456 weakly similar to GP|18057159|gb|AAL58182.1 putative far-red
impaired response protein {Oryza sativa}, partial (7%)
Length = 1244
Score = 125 bits (313), Expect = 6e-29
Identities = 62/176 (35%), Positives = 95/176 (53%), Gaps = 4/176 (2%)
Frame = +2
Query: 194 DAEFAISYIRGLGSKDPLLYCRHVANDDGNLDRLFWSDGISQLNYQVFGDVVAFDATYGK 253
D + S+ + ++ YC +D N+ LFW+D + Y+ FG+V+ D TY
Sbjct: 716 DVDAIHSFFHRMQKQNSQFYCAMDMDDKRNIQNLFWADARCRAAYEYFGEVITLDTTYLT 895
Query: 254 NRYKCPFVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFLEAMKGKAPLFVITDGDK 313
++Y P V F GVNHH ++ + A++SN +T WL R+LE M G AP +IT+ DK
Sbjct: 896 SKYDLPLVPFVGVNHHGQTILLGCAILSNLDAKTLTWLFTRWLECMHGHAPNGIITEEDK 1075
Query: 314 AMKAAIKQVFPNAHHRLCAWHIL----DNARKHVHIRAFNTELRKCMFMELDVSEF 365
AMK AI+ FP A HR C WHI+ + K+ + T L+ ++ + S+F
Sbjct: 1076AMKNAIEVAFPKARHRWCLWHIMKKVPEMLGKYSRNESIKTLLQDVVYDSMSKSDF 1243
>TC89654 weakly similar to PIR|T05644|T05644 hypothetical protein F20D10.290
- Arabidopsis thaliana, partial (65%)
Length = 1358
Score = 73.2 bits (178), Expect(2) = 1e-25
Identities = 33/103 (32%), Positives = 56/103 (54%), Gaps = 5/103 (4%)
Frame = +3
Query: 279 LVSNEKIETYVWLLERFLEAMKGKAPLFVITDGDKAMKAAIKQVFPNAHHRLCAWHIL-D 337
L+ NE +Y+WL + +L A+ G+ P+ + TD D ++ A+ QV P HR C W I +
Sbjct: 375 LILNESEPSYIWLFKTWLRAVSGRPPVSITTDLDPVIQVAVAQVLPPTRHRFCKWSIFRE 554
Query: 338 NARKHVHI----RAFNTELRKCMFMELDVSEFEEHWAATVAKF 376
N K H+ F+ E +KC+ + + EFE W + + ++
Sbjct: 555 NRSKLAHLYQSNPTFDNEFKKCVHESVTIDEFESCWRSLLERY 683
Score = 62.0 bits (149), Expect(2) = 1e-25
Identities = 27/83 (32%), Positives = 44/83 (52%)
Frame = +2
Query: 199 ISYIRGLGSKDPLLYCRHVANDDGNLDRLFWSDGISQLNYQVFGDVVAFDATYGKNRYKC 258
+ Y++ + +++ + ++ D +FW+D + NY FGD V FD TY N+Y+
Sbjct: 134 LDYLKRMRAENHAFFYAVQSDVDNAGGNIFWADETCRTNYSYFGDTVIFDTTYKTNQYRV 313
Query: 259 PFVVFYGVNHHNKSTIFSTALVS 281
PF F G NHH + +F A S
Sbjct: 314 PFASFTGFNHHGQPVLFGCAFDS 382
>TC83834 similar to PIR|G96565|G96565 F6D8.26 [imported] - Arabidopsis
thaliana, partial (24%)
Length = 1032
Score = 89.7 bits (221), Expect = 3e-18
Identities = 72/301 (23%), Positives = 133/301 (43%), Gaps = 5/301 (1%)
Frame = +3
Query: 9 KKLVPLDLMKYDFLDSEVAYLFYHWYG*VNGFAVR---KYIKIHNKE--GLVIQRNFVCH 63
+K P + +F + AY +Y Y GF VR + K ++KE G V+ C
Sbjct: 195 RKEFPAPALAMEFESYDDAYSYYICYAKEVGFCVRVKNSWFKRNSKEKYGAVL----CCS 362
Query: 64 KEGLREDRGLTMDKRQRESRTDFRCGCTAKFRVHIDKTNGRWYAKLFSDEHNHELEDDKM 123
+G + + + + ++E+RT GC A R+ + ++ RW + EHNH L
Sbjct: 363 SQGFKRTKDV--NNLRKETRT----GCPAMIRMKLVESQ-RWRICEVTLEHNHVLG---- 509
Query: 124 CGMIAAHRKMNESDIMHMNNMSRSGISTPQIHNTFASQTGGYQKVRFSKGNMYNVQGRQR 183
H+ + ++ + + + T ++++ T G + N + + +
Sbjct: 510 ---AKIHKSIKKNSLPSSDAEGK----TIKVYHALVIDTEGNDNLN---SNARDDRAFSK 659
Query: 184 RMKKKEPEKTDAEFAISYIRGLGSKDPLLYCRHVANDDGNLDRLFWSDGISQLNYQVFGD 243
K K D + +++ + +P + ND+G+L W D S+ F D
Sbjct: 660 YSNKLNLRKGDTQAIYNFLCRMQLTNPNFFYLMDFNDEGHLRNALWVDAKSRAACGYFSD 839
Query: 244 VVAFDATYGKNRYKCPFVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFLEAMKGKA 303
V+ FD TY N+Y+ P V G+NHH +S + L++ E IE+Y WL +++ + +
Sbjct: 840 VIYFDNTYLVNKYEIPLVALVGINHHGQSVLLGCGLLAGEIIESYKWLFRTWIKCIPRCS 1019
Query: 304 P 304
P
Sbjct: 1020P 1022
>BG449141 similar to GP|15983442|gb At1g10240/F14N23_12 {Arabidopsis
thaliana}, partial (33%)
Length = 690
Score = 89.0 bits (219), Expect = 5e-18
Identities = 45/170 (26%), Positives = 81/170 (47%)
Frame = +3
Query: 177 NVQGRQRRMKKKEPEKTDAEFAISYIRGLGSKDPLLYCRHVANDDGNLDRLFWSDGISQL 236
+V+ + +K +PE+ + + R + KDP + + + L+ + WS S
Sbjct: 183 DVRNLLQSFRKLDPEEETLDL-LRMCRNIKDKDPNFKFEYTLDANNRLENIAWSYASSIQ 359
Query: 237 NYQVFGDVVAFDATYGKNRYKCPFVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFL 296
Y +FGD V FD T+ + P ++ G+N++ F L+ +E + ++ W ++ FL
Sbjct: 360 LYDIFGDAVVFDTTHRLTAFDMPLGLWVGINNYGMPCFFGCVLLRDETVRSFSWAIKAFL 539
Query: 297 EAMKGKAPLFVITDGDKAMKAAIKQVFPNAHHRLCAWHILDNARKHVHIR 346
M GKAP ++TD + +K A+ P H C W I+ V+ R
Sbjct: 540 GFMNGKAPQTILTDQNICLKEALSAEMPMTKHAFCIWMIVAKFPSWVNAR 689
>CA917189 similar to PIR|T05645|T05 hypothetical protein F20D10.300 -
Arabidopsis thaliana, partial (12%)
Length = 312
Score = 64.3 bits (155), Expect = 1e-10
Identities = 30/80 (37%), Positives = 45/80 (55%), Gaps = 5/80 (6%)
Frame = +1
Query: 296 LEAMKGKAPLFVITDGDKAMKAAIKQVFPNAHHRLCAWHILDNARK---HVHIR--AFNT 350
L AM G+ PL + TD D +++AI QVFP+ HR C WHI ++ H+ ++ F
Sbjct: 1 LTAMSGRPPLSITTDHDSVIQSAIMQVFPDTRHRFCKWHIFKQCQEKLSHIFLQFPNFEA 180
Query: 351 ELRKCMFMELDVSEFEEHWA 370
E KC+ + + EFE W+
Sbjct: 181 EFHKCVNLTDSIDEFESCWS 240
>CB892328 GP|15075873|em PROBABLE GLYCOSYL HYDROLASE PROTEIN {Sinorhizobium
meliloti}, partial (1%)
Length = 828
Score = 63.9 bits (154), Expect = 2e-10
Identities = 50/218 (22%), Positives = 92/218 (41%), Gaps = 2/218 (0%)
Frame = +1
Query: 27 AYLFYHWYG*VNGFAVRKYIKIH--NKEGLVIQRNFVCHKEGLREDRGLTMDKRQRESRT 84
AY Y + GF+VRK +++ N++ ++F C K+G + + D R
Sbjct: 190 AYDLYQEHAFKTGFSVRKGKELYYDNEKKKTRLKDFYCSKQGFKNNEP---DGEVAYKRA 360
Query: 85 DFRCGCTAKFRVHIDKTNGRWYAKLFSDEHNHELEDDKMCGMIAAHRKMNESDIMHMNNM 144
D R C A R ++ K G W +HNHE + ++ + R M+ + ++
Sbjct: 361 DSRTNCLAMVRFNVTK-EGVWKVTKLILDHNHEFVPLQQRYLLRSMRNMSNLKEDRIKSL 537
Query: 145 SRSGISTPQIHNTFASQTGGYQKVRFSKGNMYNVQGRQRRMKKKEPEKTDAEFAISYIRG 204
I + + GG K +M+N K K E DA+ +++++
Sbjct: 538 VNDAIRVTNVGGYLGKEVGGVDKRGVMLKDMHNYVSTG---KLKFIEAGDAQSLLNHLQS 708
Query: 205 LGSKDPLLYCRHVANDDGNLDRLFWSDGISQLNYQVFG 242
++D + Y + + L+ +FW DG S ++Y G
Sbjct: 709 KQAQDSMFYYSVQLDHESRLNNVFWRDGKSVVDYIATG 822
>TC92547 similar to PIR|A56235|A56235 transcription activator MafB -
chicken, partial (6%)
Length = 1158
Score = 63.5 bits (153), Expect = 2e-10
Identities = 29/88 (32%), Positives = 52/88 (58%)
Frame = +3
Query: 6 IDLKKLVPLDLMKYDFLDSEVAYLFYHWYG*VNGFAVRKYIKIHNKEGLVIQRNFVCHKE 65
+++ + +++K +F ++ AY FY+ YG GF++RK N G++ R FVC+K
Sbjct: 891 VNVNSIASDEILKLEFGPADEAYEFYYRYGKCKGFSIRKGDVRRNSSGIITMREFVCNKN 1070
Query: 66 GLREDRGLTMDKRQRESRTDFRCGCTAK 93
GLR+ + L+ + R+R+ R R C A+
Sbjct: 1071GLRDKKHLSRNDRKRDHRRLTRTNCEAR 1154
>BG586672
Length = 667
Score = 53.5 bits (127), Expect = 2e-07
Identities = 27/60 (45%), Positives = 37/60 (61%)
Frame = +3
Query: 1 QDILGIDLKKLVPLDLMKYDFLDSEVAYLFYHWYG*VNGFAVRKYIKIHNKEGLVIQRNF 60
+DI ID KKL D+MK+ F + VAY FY+WY +NGF+ R+ NK +IQ+ F
Sbjct: 462 EDIGKIDFKKLNASDVMKFHFPNIAVAYTFYNWYARMNGFSARRSKVRRNKNNEIIQQIF 641
>TC84106 similar to GP|9502168|gb|AAF88018.1| contains simlarity to
Arabidopsis thaliana far-red impaired response protein
(GB:AAD51282.1), partial (25%)
Length = 673
Score = 53.1 bits (126), Expect = 3e-07
Identities = 31/85 (36%), Positives = 47/85 (54%), Gaps = 2/85 (2%)
Frame = +2
Query: 66 GLREDRGLTMDKRQRESRTDFRCGCTAKFRVHIDKTNG--RWYAKLFSDEHNHELEDDKM 123
GL RG M R+ R RCGC AK + D +G +W FS+ HNHEL +D
Sbjct: 353 GLPRLRGRQMVMHHRD-RKSVRCGCDAKMYLSKDVVDGVSQWTVLQFSNVHNHELLEDDQ 529
Query: 124 CGMIAAHRKMNESDIMHMNNMSRSG 148
++ A+RK++E+D + +S++G
Sbjct: 530 VRLLPAYRKIHEADQERILLLSKAG 604
>TC90562 similar to GP|16604382|gb|AAL24197.1 AT3g59470/T16L24_20
{Arabidopsis thaliana}, partial (66%)
Length = 1169
Score = 49.3 bits (116), Expect = 4e-06
Identities = 35/106 (33%), Positives = 48/106 (45%), Gaps = 3/106 (2%)
Frame = +2
Query: 20 DFLDSEVAYLFYHWYG*VNGFAVR-KYIKIHNKEGLVIQRNFVCHKEGLREDRGLTMDKR 78
+F+ A+ FY+ Y GF +R + ++G VI R VC+KEG R DKR
Sbjct: 338 EFVSEAEAHAFYNSYATRVGFVIRVSKLSRSRRDGSVIGRALVCNKEGFR-----MPDKR 502
Query: 79 QR--ESRTDFRCGCTAKFRVHIDKTNGRWYAKLFSDEHNHELEDDK 122
++ R + R GC A V +G W F EH H L K
Sbjct: 503 EKIVRQRAETRVGCRAMIMVR-KLNSGLWSITKFVKEHTHPLTPGK 637
>CB893278 weakly similar to PIR|T05645|T056 hypothetical protein F20D10.300 -
Arabidopsis thaliana, partial (5%)
Length = 821
Score = 47.0 bits (110), Expect = 2e-05
Identities = 35/102 (34%), Positives = 46/102 (44%), Gaps = 4/102 (3%)
Frame = +3
Query: 21 FLDSEVAYLFYHWYG*VNGFAVRKYIKIHNKEGLV----IQRNFVCHKEGLREDRGLTMD 76
F E A FY YG GF VR + HN+ V I ++FVC KEG R + +
Sbjct: 333 FNSREEARGFYDGYGRRIGFTVRIH---HNRRSRVNNELIGQDFVCSKEGFRAKKYVHRK 503
Query: 77 KRQRESRTDFRCGCTAKFRVHIDKTNGRWYAKLFSDEHNHEL 118
R R GC A R+ + + G+W F EH H+L
Sbjct: 504 DRVLPPPPATREGCQAMIRLAL-RDEGKWVVTKFVKEHTHKL 626
>TC81450 similar to GP|16604382|gb|AAL24197.1 AT3g59470/T16L24_20
{Arabidopsis thaliana}, partial (59%)
Length = 832
Score = 40.0 bits (92), Expect = 0.002
Identities = 29/90 (32%), Positives = 38/90 (42%), Gaps = 3/90 (3%)
Frame = +2
Query: 51 KEGLVIQRNFVCHKEGLREDRGLTMDKRQR--ESRTDFRCGCTAKFRVHIDKTNGRWYAK 108
K+G I R VC+KEG R DKR++ R + R GC A + +G+W
Sbjct: 149 KDGTAIGRALVCNKEGYR-----MPDKREKIVRQRAETRVGCRAMIMMR-KINSGKWVIT 310
Query: 109 LFSDEHNHELE-DDKMCGMIAAHRKMNESD 137
F EH H L M + NE D
Sbjct: 311 KFVKEHTHPLNPGSSRRDMFEIYPSQNEHD 400
>TC79804 weakly similar to GP|6175165|gb|AAF04891.1| Mutator-like
transposase {Arabidopsis thaliana}, partial (57%)
Length = 2129
Score = 38.9 bits (89), Expect = 0.005
Identities = 24/102 (23%), Positives = 44/102 (42%), Gaps = 4/102 (3%)
Frame = +1
Query: 244 VVAFDATYGKNRYKCPFVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFLEAMKGKA 303
+VA K++Y F+ + + A+V E E++ W L A++
Sbjct: 757 IVALGGIQLKSKYLSTFLSATSFDADGGLFPLAFAVVDVENDESWTWFLSELHNALEVNT 936
Query: 304 P----LFVITDGDKAMKAAIKQVFPNAHHRLCAWHILDNARK 341
+ ++DG K + AI++ FP + H C H+ +N K
Sbjct: 937 ECMPQIIFLSDGQKGIVDAIRRKFPRSSHAFCMRHLSENIGK 1062
>BQ147857 weakly similar to GP|15983442|gb At1g10240/F14N23_12 {Arabidopsis
thaliana}, partial (26%)
Length = 560
Score = 37.0 bits (84), Expect = 0.021
Identities = 21/81 (25%), Positives = 36/81 (43%), Gaps = 6/81 (7%)
Frame = +1
Query: 303 APLFVITDGDKAMKAAIKQVFPNAHHRLCAWHILDNARKHVHI------RAFNTELRKCM 356
AP ++TD + +K AI P + H C WHIL ++ + + +
Sbjct: 1 APQTLLTDHNTWLKEAIAVEMPESKHAFCIWHILSKFSDWXYLLLGSQYDEWKADFHRLY 180
Query: 357 FMELDVSEFEEHWAATVAKFG 377
+E+ +FE+ W V K+G
Sbjct: 181 NLEM-XEDFEKSWRQMVDKYG 240
>TC82992 homologue to PIR|T05645|T05645 hypothetical protein F20D10.300 -
Arabidopsis thaliana, partial (7%)
Length = 354
Score = 32.0 bits (71), Expect = 0.67
Identities = 18/48 (37%), Positives = 25/48 (51%), Gaps = 1/48 (2%)
Frame = +1
Query: 20 DFLDSEVAYLFYHWYG*VNGFAVR-KYIKIHNKEGLVIQRNFVCHKEG 66
+F E A FY+ Y GF+ R + ++G +IQR VC KEG
Sbjct: 208 EFESEEAAKAFYNSYARRVGFSTRVSSSRRSRRDGAIIQRQXVCAKEG 351
>TC82509 similar to GP|15293233|gb|AAK93727.1 unknown protein {Arabidopsis
thaliana}, partial (72%)
Length = 662
Score = 28.5 bits (62), Expect = 7.4
Identities = 18/60 (30%), Positives = 29/60 (48%), Gaps = 1/60 (1%)
Frame = +3
Query: 59 NFVCHKEGLREDRGLTMDKRQRESRTDFRCGCTAKFRVHIDKTNG-RWYAKLFSDEHNHE 117
N+ CH+ + + G D+ QRE + + C + H TNG +Y+ LF NH+
Sbjct: 222 NYHCHQCQWQTELGCCHDEGQREEK--YTCCTNIEKHNHGCNTNGNNFYSSLFRLSSNHK 395
>TC90977 similar to GP|10178227|dbj|BAB11607. ZW10-like protein {Arabidopsis
thaliana}, partial (84%)
Length = 779
Score = 28.1 bits (61), Expect = 9.7
Identities = 13/32 (40%), Positives = 17/32 (52%)
Frame = +3
Query: 313 KAMKAAIKQVFPNAHHRLCAWHILDNARKHVH 344
K KA + + +AH W +DN RKHVH
Sbjct: 3 KNYKAYLNPDYFDAHLTPLGWEQVDNLRKHVH 98
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.340 0.149 0.499
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,666,312
Number of Sequences: 36976
Number of extensions: 263036
Number of successful extensions: 1800
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 1285
Number of HSP's successfully gapped in prelim test: 54
Number of HSP's that attempted gapping in prelim test: 499
Number of HSP's gapped (non-prelim): 1357
length of query: 578
length of database: 9,014,727
effective HSP length: 101
effective length of query: 477
effective length of database: 5,280,151
effective search space: 2518632027
effective search space used: 2518632027
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0302a.8


