

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0301a.2
(80 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
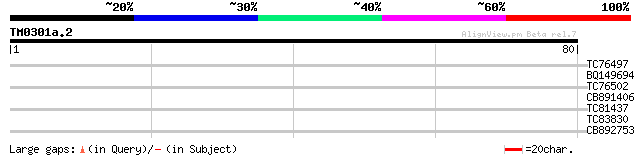
TC76497 homologue to SP|P06411|PSBB_TOBAC Photosystem II P680 ch... 28 0.64
BQ149694 26 2.4
TC76502 weakly similar to GP|1311386|pdb|1CBG| Cyanogenic Beta-G... 25 5.4
CB891406 weakly similar to GP|10176783|dbj contains similarity t... 25 5.4
TC81437 similar to GP|10177523|dbj|BAB10918. gene_id:K1F13.7~sim... 25 7.1
TC83830 similar to GP|18146966|dbj|BAB82502. cig3 {Nicotiana tab... 25 7.1
CB892753 similar to GP|10172595|d multidrug resistance-associate... 24 9.3
>TC76497 homologue to SP|P06411|PSBB_TOBAC Photosystem II P680 chlorophyll A
apoprotein (CP-47 protein). [Common tobacco] {Nicotiana
tabacum}, complete
Length = 4045
Score = 28.1 bits (61), Expect = 0.64
Identities = 14/20 (70%), Positives = 16/20 (80%)
Frame = +2
Query: 45 LDNTKMK*IIENSRGR*GSR 64
+ KM+*IIE SRGR*GSR
Sbjct: 3017 IQTKKMR*IIEYSRGR*GSR 3076
>BQ149694
Length = 610
Score = 26.2 bits (56), Expect = 2.4
Identities = 15/41 (36%), Positives = 22/41 (53%)
Frame = -2
Query: 10 WLRPNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLDNTKM 50
+L+P +LS G E S+MFL+ GTLL + K+
Sbjct: 543 FLKPMMNLSMGMKPQTMPKEPNDKKSSMFLNKGTLL*DNKL 421
>TC76502 weakly similar to GP|1311386|pdb|1CBG| Cyanogenic Beta-Glucosidase
Mol_id: 1; Molecule: Cyanogenic Beta-Glucosidase; Chain:
Null; Ec: 3.2., partial (32%)
Length = 1738
Score = 25.0 bits (53), Expect = 5.4
Identities = 13/16 (81%), Positives = 14/16 (87%)
Frame = +1
Query: 49 KMK*IIENSRGR*GSR 64
KM+*IIE SR R*GSR
Sbjct: 1627 KMR*IIEYSRXR*GSR 1674
>CB891406 weakly similar to GP|10176783|dbj contains similarity to protein
kinase~gene_id:MIK19.26 {Arabidopsis thaliana}, partial
(5%)
Length = 826
Score = 25.0 bits (53), Expect = 5.4
Identities = 11/23 (47%), Positives = 14/23 (60%)
Frame = +2
Query: 3 LLIHRQRWLRPNNSLSNGFFRSN 25
L++HRQ P N LS +RSN
Sbjct: 437 LVLHRQTPNNPRNKLSESVWRSN 505
>TC81437 similar to GP|10177523|dbj|BAB10918. gene_id:K1F13.7~similar to
unknown protein~sp|P55606 {Arabidopsis thaliana},
partial (47%)
Length = 1426
Score = 24.6 bits (52), Expect = 7.1
Identities = 20/62 (32%), Positives = 29/62 (46%), Gaps = 8/62 (12%)
Frame = +1
Query: 2 LLLIHRQRWLRPNNSLSNGFFRSN-TLSESYQYLSNMFLSNGTL-------LDNTKMK*I 53
+LLIH+Q +L+ + FR N L +L+N + N L N K+K*I
Sbjct: 268 ILLIHQQPFLKLSKMFMRVLFRKNQALGPLSMHLANSQMQNQKLWRKHS*YCSNLKIK*I 447
Query: 54 IE 55
E
Sbjct: 448 KE 453
>TC83830 similar to GP|18146966|dbj|BAB82502. cig3 {Nicotiana tabacum},
partial (4%)
Length = 790
Score = 24.6 bits (52), Expect = 7.1
Identities = 12/31 (38%), Positives = 17/31 (54%)
Frame = +3
Query: 2 LLLIHRQRWLRPNNSLSNGFFRSNTLSESYQ 32
LL H Q+ L+P L+NG +L E +Q
Sbjct: 429 LLKAHHQQQLQPPQPLTNGNQHFTSLPEQFQ 521
>CB892753 similar to GP|10172595|d multidrug resistance-associated protein
(MRP); ABC-transoprter {Arabidopsis thaliana}, partial
(14%)
Length = 783
Score = 24.3 bits (51), Expect = 9.3
Identities = 13/33 (39%), Positives = 19/33 (57%), Gaps = 1/33 (3%)
Frame = -3
Query: 13 PNNSLSNGFFRSNTLSESYQYLS-NMFLSNGTL 44
PN+ +S FF+ + S ++ YLS MF N L
Sbjct: 493 PNDKISKSFFKEHA-SSTFSYLSLTMFSPNNIL 398
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.350 0.154 0.482
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,093,525
Number of Sequences: 36976
Number of extensions: 20324
Number of successful extensions: 296
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 295
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 296
length of query: 80
length of database: 9,014,727
effective HSP length: 56
effective length of query: 24
effective length of database: 6,944,071
effective search space: 166657704
effective search space used: 166657704
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0301a.2


