

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0299.14
(88 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
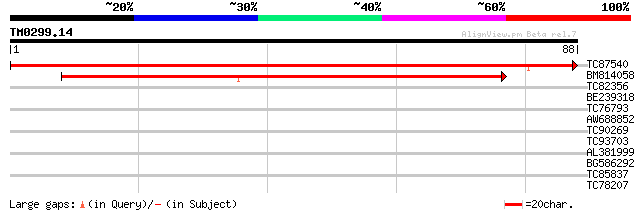
Score E
Sequences producing significant alignments: (bits) Value
TC87540 similar to GP|6714455|gb|AAF26142.1| unknown protein {Ar... 155 2e-39
BM814058 77 2e-15
TC82356 homologue to GP|10177845|dbj|BAB11274. protein kinase-li... 27 1.0
BE239318 weakly similar to GP|2369690|emb| FPF1 protein {Arabido... 26 2.3
TC76793 similar to GP|9711891|dbj|BAB07982.1 contains EST AU0711... 26 2.3
AW688852 weakly similar to PIR|E71443|E714 probable DNA-binding ... 25 4.0
TC90269 similar to GP|15451577|gb|AAK98701.1 Putative ABC transp... 25 4.0
TC93703 similar to GP|9294064|dbj|BAB02021.1 DNA-dependent RNA p... 25 5.2
AL381999 similar to GP|18033111|gb functional candidate resistan... 25 5.2
BG586292 similar to PIR|T46163|T46 nodulin / glutamate-ammonia l... 25 6.8
TC85837 similar to GP|21539471|gb|AAM53288.1 putative protein {A... 25 6.8
TC78207 similar to SP|Q02921|NO93_SOYBN Early nodulin 93 (N-93).... 25 6.8
>TC87540 similar to GP|6714455|gb|AAF26142.1| unknown protein {Arabidopsis
thaliana}, partial (77%)
Length = 734
Score = 155 bits (393), Expect = 2e-39
Identities = 75/90 (83%), Positives = 85/90 (94%), Gaps = 2/90 (2%)
Frame = +1
Query: 1 MAEAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQAL 60
M++AKTQ+ES+RKWVV+HKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQAL
Sbjct: 148 MSDAKTQIESLRKWVVEHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQAL 327
Query: 61 TLAALAGAAVVEYYDHRAE--AKRAKNSEQ 88
TL ALAGAAVVEYYDH++E AK A++S +
Sbjct: 328 TLGALAGAAVVEYYDHKSEEKAKAARDSRR 417
>BM814058
Length = 574
Score = 76.6 bits (187), Expect = 2e-15
Identities = 37/70 (52%), Positives = 52/70 (73%), Gaps = 1/70 (1%)
Frame = +1
Query: 9 ESIRKWVVDHKLRTVGCLWLSGITGS-IAYNWSRPNMKTSVKIIHARLHAQALTLAALAG 67
E+I+ WV +KL ++G LW SGI S +AY+ + MK S+++IHAR+HAQALTLA L+G
Sbjct: 25 EAIQSWVSKNKLASIGTLWASGIGASLVAYSRVKSPMKPSLRLIHARMHAQALTLAVLSG 204
Query: 68 AAVVEYYDHR 77
AA YY++R
Sbjct: 205AAAYHYYENR 234
>TC82356 homologue to GP|10177845|dbj|BAB11274. protein kinase-like protein
{Arabidopsis thaliana}, partial (37%)
Length = 1108
Score = 27.3 bits (59), Expect = 1.0
Identities = 8/23 (34%), Positives = 13/23 (55%)
Frame = +2
Query: 25 CLWLSGITGSIAYNWSRPNMKTS 47
C+W+ GI G+ Y W + + S
Sbjct: 467 CIWIFGIMGNCCYKWDSSDYRVS 535
>BE239318 weakly similar to GP|2369690|emb| FPF1 protein {Arabidopsis
thaliana}, partial (43%)
Length = 515
Score = 26.2 bits (56), Expect = 2.3
Identities = 9/24 (37%), Positives = 12/24 (49%)
Frame = +2
Query: 38 NWSRPNMKTSVKIIHARLHAQALT 61
NW PN KT+ I +H +T
Sbjct: 38 NWENPNFKTTTNIYRTFIHMNTVT 109
>TC76793 similar to GP|9711891|dbj|BAB07982.1 contains EST
AU071162(R10843)~unknown protein {Oryza sativa (japonica
cultivar-group)}, partial (86%)
Length = 781
Score = 26.2 bits (56), Expect = 2.3
Identities = 9/24 (37%), Positives = 12/24 (49%)
Frame = -3
Query: 38 NWSRPNMKTSVKIIHARLHAQALT 61
NW PN KT+ I +H +T
Sbjct: 707 NWENPNFKTTTNIYRTFIHMNTVT 636
>AW688852 weakly similar to PIR|E71443|E714 probable DNA-binding protein -
Arabidopsis thaliana, partial (9%)
Length = 536
Score = 25.4 bits (54), Expect = 4.0
Identities = 16/42 (38%), Positives = 23/42 (54%)
Frame = +1
Query: 37 YNWSRPNMKTSVKIIHARLHAQALTLAALAGAAVVEYYDHRA 78
+N SRP I HARLH + + +A G+ V+Y D R+
Sbjct: 331 WNASRP-------IWHARLHDASYSASAS*GSC*VQYGDERS 435
>TC90269 similar to GP|15451577|gb|AAK98701.1 Putative ABC transporter
{Oryza sativa} [Oryza sativa (japonica cultivar-group)],
partial (19%)
Length = 1225
Score = 25.4 bits (54), Expect = 4.0
Identities = 11/32 (34%), Positives = 18/32 (55%), Gaps = 4/32 (12%)
Frame = +2
Query: 15 VVDHKLRTVGCLWL----SGITGSIAYNWSRP 42
V H L T+ +W+ I+G + +NW+RP
Sbjct: 659 VTGHPLSTL*LIWIW*DQGQISGLLPWNWNRP 754
>TC93703 similar to GP|9294064|dbj|BAB02021.1 DNA-dependent RNA polymerase
II {Arabidopsis thaliana}, partial (17%)
Length = 598
Score = 25.0 bits (53), Expect = 5.2
Identities = 9/21 (42%), Positives = 15/21 (70%)
Frame = +3
Query: 21 RTVGCLWLSGITGSIAYNWSR 41
RT GC +SG++G I++ +R
Sbjct: 264 RTEGCFRISGVSGKISFRKTR 326
>AL381999 similar to GP|18033111|gb functional candidate resistance protein
KR1 {Glycine max}, partial (4%)
Length = 451
Score = 25.0 bits (53), Expect = 5.2
Identities = 10/31 (32%), Positives = 18/31 (57%)
Frame = +3
Query: 50 IIHARLHAQALTLAALAGAAVVEYYDHRAEA 80
+++ +LH T L A + E++DH+ EA
Sbjct: 69 LLNQKLHEAGNTRFCLPRAKIPEWFDHQCEA 161
>BG586292 similar to PIR|T46163|T46 nodulin / glutamate-ammonia ligase-like
protein - Arabidopsis thaliana, partial (50%)
Length = 719
Score = 24.6 bits (52), Expect = 6.8
Identities = 8/21 (38%), Positives = 15/21 (71%)
Frame = -1
Query: 27 WLSGITGSIAYNWSRPNMKTS 47
W +GITGS + WS+ + +++
Sbjct: 242 WRAGITGSSLWEWSQTSPRST 180
>TC85837 similar to GP|21539471|gb|AAM53288.1 putative protein {Arabidopsis
thaliana}, partial (85%)
Length = 1144
Score = 24.6 bits (52), Expect = 6.8
Identities = 18/56 (32%), Positives = 27/56 (48%)
Frame = +2
Query: 20 LRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAALAGAAVVEYYD 75
L +G + + G I+Y S P K K+IH LHA AL L + A + ++
Sbjct: 359 LMLIGLIIIGG-EAIISYK-SLPLKKEVKKLIHLVLHAHALVLGIIGICAAFKNHN 520
>TC78207 similar to SP|Q02921|NO93_SOYBN Early nodulin 93 (N-93). [Soybean]
{Glycine max}, partial (93%)
Length = 617
Score = 24.6 bits (52), Expect = 6.8
Identities = 16/48 (33%), Positives = 22/48 (45%)
Frame = +3
Query: 39 WSRPNMKTSVKIIHARLHAQALTLAALAGAAVVEYYDHRAEAKRAKNS 86
W+R N+ + AQAL ++ +AGAA D A KNS
Sbjct: 264 WARANLNPT---------AQALIISTVAGAAYFVVADKTVLATARKNS 380
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.127 0.376
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,647,335
Number of Sequences: 36976
Number of extensions: 29921
Number of successful extensions: 157
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 156
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 156
length of query: 88
length of database: 9,014,727
effective HSP length: 64
effective length of query: 24
effective length of database: 6,648,263
effective search space: 159558312
effective search space used: 159558312
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0299.14


