

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
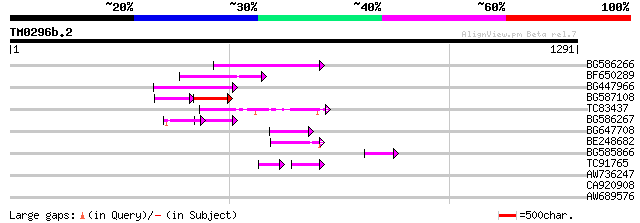
Query= TM0296b.2
(1291 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-li... 157 3e-38
BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgari... 132 7e-31
BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [impo... 124 3e-28
BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - ... 73 2e-20
TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imp... 70 7e-12
BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At... 59 1e-08
BG647708 weakly similar to GP|13786450|gb| putative reverse tran... 57 4e-08
BE248682 similar to GP|18568269|gb putative gag-pol polyprotein ... 57 5e-08
BG585866 54 5e-07
TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative ret... 45 2e-05
AW736247 weakly similar to PIR|T14619|T146 reverse transcriptase... 39 0.010
CA920908 homologue to PIR|T06377|T06 SAR DNA-binding protein-1 -... 30 6.1
AW689576 similar to GP|7940277|gb| F2401.8 {Arabidopsis thaliana... 30 7.9
>BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 789
Score = 157 bits (397), Expect = 3e-38
Identities = 93/253 (36%), Positives = 132/253 (51%), Gaps = 1/253 (0%)
Frame = -3
Query: 465 IKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLPGRGTMDNAFLA*EIIHQM 524
+ E+R I+ CN YK+I K+L R++P L I+ P Q++F+PGR DN + +I+H +
Sbjct: 778 VSEYRTIAPCNTQYKIIAKILSKRMQPLLRSIISPSQSAFVPGRAISDNVLITHKILHYL 599
Query: 525 HRSMARKG-NLAFKIDLEKAYDSISWSFLRETLELYEFPLVIIDLIMCYVSSSQISLLWN 583
+S A+K ++A K D+ KAYD I+W+FLRE L F + I IM VS+ S L N
Sbjct: 598 RQSGAKKHVSMAVKTDMTKAYDRIAWNFLREVLTRLGFHGIWISWIMECVSTVSYSFLIN 419
Query: 584 GAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPGRWKPLHRTAGGTGISHLF 643
G P P RGLRQGDP SPY F+LC E LS Q + G + I+HL
Sbjct: 418 GGPQGRVLPSRGLRQGDPLSPYLFILCTEVLSGLCQQALRKGTLPGVKVARNCPPINHLL 239
Query: 644 FAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKAITSIGVAPEVRTEIGLRAL 703
FA D + F K++ + +L+ + S IN KS S + + +
Sbjct: 238 FADDTMFFGKSNASSCAILLSIMDKYRAASGRCIN*TKSAITFSSKTSQAIIDRVKGELK 59
Query: 704 IPFVNDLGKYLGF 716
I GKYLG+
Sbjct: 58 IAKEGGTGKYLGY 20
>BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgaris}, partial
(69%)
Length = 616
Score = 132 bits (333), Expect = 7e-31
Identities = 73/201 (36%), Positives = 116/201 (57%), Gaps = 3/201 (1%)
Frame = +3
Query: 387 LLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDE 446
LL + EV+ AL SM S APG DG+ FFK W +GD+V + + F++G + +
Sbjct: 9 LLCSEFTAVEVKNALFSMDSSKAPGIDGYNVHFFKCSWNIIGDSVIDAILDFFKTGFMPK 188
Query: 447 RLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLP 506
+ V L+PK + TS+K FRPI+ C+V YK+I+K+L +R++ LN +V Q++F+
Sbjct: 189 IINCTYVTLLPKEVNVTSVKNFRPIACCSVIYKIISKILTSRMQGVLNSVVSENQSAFVK 368
Query: 507 GRGTMDNAFLA*EIIHQMHRSMARKG---NLAFKIDLEKAYDSISWSFLRETLELYEFPL 563
GR DN L+ H++ +S +RKG KIDL KAYDS W F++ + FP
Sbjct: 369 GRVIFDNIILS----HELVKSYSRKGISPRCMVKIDLXKAYDSXEWPFIKHLMLELGFPY 536
Query: 564 VIIDLIMCYVSSSQISLLWNG 584
++ +M ++++ + NG
Sbjct: 537 KFVNWVMAXLTTASYTFNXNG 599
>BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [imported] -
Arabidopsis thaliana, partial (7%)
Length = 687
Score = 124 bits (310), Expect = 3e-28
Identities = 66/191 (34%), Positives = 102/191 (52%)
Frame = +2
Query: 327 RKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQEASQTPFPGDNVPKLKEVVRF 386
RK N + +LK G+WC ++ + YF LF + + KL
Sbjct: 62 RKVNEIKKLKDETGNWCKGEENVERLLITYFNNLFTSSNPTAIEETCEVVKGKLSHEHIV 241
Query: 387 LLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDE 446
+ ++EEV +A+ M APGPDG FF++YW VG V ++V + + E
Sbjct: 242 WCEKEFTEEEVLEAINQMHPVKAPGPDGLPALFFQKYWHIVGKEVQQMVLQVLNNSMETE 421
Query: 447 RLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLP 506
L + +VLIPK +P + K++RPISLCNV K+ITK++ NR++ L D++ Q++F+
Sbjct: 422 ELNKTFIVLIPKGKNPNTPKDYRPISLCNVVMKIITKVIANRVKQTLPDVIDVEQSAFVQ 601
Query: 507 GRGTMDNAFLA 517
GR DNA +A
Sbjct: 602 GRLITDNALIA 634
>BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - Arabidopsis
thaliana, partial (17%)
Length = 677
Score = 73.2 bits (178), Expect(2) = 2e-20
Identities = 36/89 (40%), Positives = 54/89 (60%)
Frame = +3
Query: 419 FFKQYWEQVGDTVWKVVSEAFESGKVDERLLEILVVLIPKVNHPTSIKEFRPISLCNVTY 478
FF+ W + + K+V+ SGK+D RL + LIPK PT + E RPISLCNV Y
Sbjct: 411 FFQHSWHIIKMDLLKMVNSFLASGKLDTRLNTTNICLIPKKKRPTRMTELRPISLCNVGY 590
Query: 479 KLITKLLVNRIRPFLNDIVGPMQNSFLPG 507
K+I+K+L R++ L ++ Q++F+ G
Sbjct: 591 KIISKVLCQRLKVCLPSLISETQSAFVHG 677
Score = 45.4 bits (106), Expect(2) = 2e-20
Identities = 27/92 (29%), Positives = 43/92 (46%)
Frame = +2
Query: 329 RNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQEASQTPFPGDNVPKLKEVVRFLL 388
RN + L DG+W ++ +++ A+ YF+ LF F + + + L
Sbjct: 140 RNRIVGLYDYDGNWITEEQGVEKVAVDYFEDLFQRTTPTGFDGFLDEITSSITPQMNQRL 319
Query: 389 VQPVSKEEVRQALMSMKSFTAPGPDGFQPFFF 420
++ ++EEVR AL M APGPDG F
Sbjct: 320 LRLATEEEVRLALFIMHPEKAPGPDGMTTLLF 415
>TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imported] -
Arabidopsis thaliana, partial (6%)
Length = 951
Score = 69.7 bits (169), Expect = 7e-12
Identities = 82/315 (26%), Positives = 132/315 (41%), Gaps = 16/315 (5%)
Frame = +2
Query: 432 WKVVSEAFESGKVDERLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRP 491
++ VSE + K+ + + + LIPKV++P + +FRPISL YK++ KLL NR+R
Sbjct: 2 FRFVSEFHRNRKLFKGINSTFIALIPKVDNPQRLNDFRPISLVGSLYKILGKLLANRLRV 181
Query: 492 FLNDIVGPMQNSFLPGRGTMDNAFLA*EIIHQMHRSMARKGNLAFKIDLEKAYDSISWSF 551
+ ++ Q++F+ R ++ FL QM M + + K + + W
Sbjct: 182 VIGSVISDAQSAFVKNRQILEMVFL-----*QMRLWMRLR-------N*RKIFCCLRWIL 325
Query: 552 LR-ETLEL----------YEFPLVIIDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGD 600
R TL + F ++ I VS++ S+L NG+P + L Q
Sbjct: 326 KRLITLSIGLIWILF*VGMSFLVLWRKWIKECVSTATTSVLVNGSPTNVLM--KSLVQTQ 499
Query: 601 PKSPYFFVLCMERLSIHIQNLVEPGRWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVT 660
+ Y F +V P +SHL FA D LL + A +
Sbjct: 500 LFTRYSF------------GVVNP------------VVVSHLQFANDTLLLETKNWANIR 607
Query: 661 VLVDALQFLCEGSVLRINVQKSKAITSIGVAPEVRTEIG-----LRALIPFVNDLGKYLG 715
L AL S L++N KS + + +AP +E +PF+ YLG
Sbjct: 608 ALRAALVIF*AMSGLKVNFHKS-GLVCVNIAPSWLSEAASVLSWKVGKVPFL-----YLG 769
Query: 716 FPLQGRRLNSERCEY 730
P++G NS R +
Sbjct: 770 MPIEG---NSRRLSF 805
>BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (16%)
Length = 794
Score = 58.9 bits (141), Expect = 1e-08
Identities = 32/102 (31%), Positives = 58/102 (56%), Gaps = 6/102 (5%)
Frame = +3
Query: 351 EEALG-----YFQALFAGNQEASQTPFPGDNVPKL-KEVVRFLLVQPVSKEEVRQALMSM 404
EE LG +F+ L++ +++ T +++P + E L+ +S+EEVR+A+ +
Sbjct: 273 EEDLGRLADSHFKLLYS-SEDVGITLEDWNSIPAIVTEEQNAQLMAQISREEVREAVFDI 449
Query: 405 KSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDE 446
PGPDG FFF+Q+W+ +GD + + E +GK++E
Sbjct: 450 NPHKCPGPDGMNVFFFQQFWDTMGDDLTSMAQEFLRTGKLEE 575
Score = 51.2 bits (121), Expect = 3e-06
Identities = 32/98 (32%), Positives = 49/98 (49%)
Frame = +1
Query: 420 FKQYWEQVGDTVWKVVSEAFESGKVDERLLEILVVLIPKVNHPTSIKEFRPISLCNVTYK 479
F WE + +W+ S E G + + L+PK + EFRPISLCNV YK
Sbjct: 505 FGTQWEMISH-LWRKNSS--EQGNLKRESTKQTSGLVPKKLEAKRLVEFRPISLCNVAYK 675
Query: 480 LITKLLVNRIRPFLNDIVGPMQNSFLPGRGTMDNAFLA 517
+++K+L R++ L I+ Q +F + DN +A
Sbjct: 676 IVSKVLSKRLKSVLPWIITETQAAFGRRQLISDNILIA 789
>BG647708 weakly similar to GP|13786450|gb| putative reverse transcriptase
{Oryza sativa}, partial (9%)
Length = 708
Score = 57.4 bits (137), Expect = 4e-08
Identities = 33/99 (33%), Positives = 48/99 (48%)
Frame = +1
Query: 592 PGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPGRWKPLHRTAGGTGISHLFFAGDVLLF 651
P +GLRQGDP SPY F+LC LS ++ + I+HL FA D LLF
Sbjct: 4 PEKGLRQGDPLSPYLFILCANVLSGLLKREGNKQNLHGIQVARSDPKITHLLFADDSLLF 183
Query: 652 CKASVAQVTVLVDALQFLCEGSVLRINVQKSKAITSIGV 690
+A++ + ++ L S +N +KS+ S V
Sbjct: 184 ARANLTEAATIMQVLHSYQSASGQLVNFEKSEVSYSQNV 300
>BE248682 similar to GP|18568269|gb putative gag-pol polyprotein {Zea mays},
partial (1%)
Length = 441
Score = 57.0 bits (136), Expect = 5e-08
Identities = 46/130 (35%), Positives = 62/130 (47%), Gaps = 6/130 (4%)
Frame = +3
Query: 594 RGLRQGDPKSPYFFVLCMERLSIHIQNLVEPGRWKPLHRTAGGTGISHLFFAGDVLLFCK 653
RGL+QGDP +P+ F+L E +S ++N V ++ GGT +SHL +A D L
Sbjct: 24 RGLKQGDPLAPFLFLLVAEGISGLMKNAVNRNLFQGFDVKRGGTRVSHLQYADDTLCIGM 203
Query: 654 ASVAQVTVLVDALQFLCEGSVLRINVQKSKAITSIGVAPEVRTEIGLRAL------IPFV 707
+V + L LQ S L++N KS I I V P E R L IPF+
Sbjct: 204 PTVDNLWTLKALLQGFEMASGLKVNFHKSSLI-GINV-PRDFMEAACRFLNCREESIPFI 377
Query: 708 NDLGKYLGFP 717
YLG P
Sbjct: 378 -----YLGLP 392
>BG585866
Length = 828
Score = 53.5 bits (127), Expect = 5e-07
Identities = 25/78 (32%), Positives = 35/78 (44%), Gaps = 1/78 (1%)
Frame = +3
Query: 809 DILQHGAWSLNGLYTVCDFDIASRLNRVTPTLSTAQEDQWIWGPGSRGTYLVKEAYSW-L 867
D+ G LYT+ DIA +N + + D +IW S G Y K YSW L
Sbjct: 237 DVFTTGGQHTQSLYTILPTDIAEVINNTHLNFNASIGDAYIWPHNSNGVYTAKSGYSWIL 416
Query: 868 HERNRVSAGAGNWRWLWR 885
+ V+ +W W+WR
Sbjct: 417 SQTETVNYNNSSWSWIWR 470
>TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative retroelement
{Oryza sativa (japonica cultivar-group)}, partial (1%)
Length = 625
Score = 44.7 bits (104), Expect(2) = 2e-05
Identities = 26/60 (43%), Positives = 33/60 (54%)
Frame = +2
Query: 566 IDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPG 625
I MC S+ L+ N A P P RGL+QGD SPY F++C+E LS I + E G
Sbjct: 89 IGCFMCVESNDYYVLVNNDAVDPII-PSRGLQQGDHLSPYIFIICVEGLSFLIPHAKERG 265
Score = 23.1 bits (48), Expect(2) = 2e-05
Identities = 20/76 (26%), Positives = 31/76 (40%)
Frame = +3
Query: 642 LFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKAITSIGVAPEVRTEIGLR 701
L F G L + ++ + L E S I+++KS S V ++T I
Sbjct: 279 LLFEGAPLRSLRVEEHHAQIMKNILILYEEDSGKAISLRKS*IYCSRNVPDILKTSITYI 458
Query: 702 ALIPFVNDLGKYLGFP 717
+ F+ KYLG P
Sbjct: 459 LGVQFMLGTCKYLGLP 506
>AW736247 weakly similar to PIR|T14619|T146 reverse transcriptase - beet
retrotransposon (fragment), partial (4%)
Length = 305
Score = 39.3 bits (90), Expect = 0.010
Identities = 22/54 (40%), Positives = 31/54 (56%), Gaps = 2/54 (3%)
Frame = +1
Query: 393 SKEEVRQALMSMKSFTA--PGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKV 444
S+EEVR+A+ S + PGPDG F K+YWE + +VV E +GK+
Sbjct: 124 SEEEVRKAVWDCDSSHS*NPGPDGVNFTFVKEYWELIKVDFLRVVMEFHTNGKL 285
>CA920908 homologue to PIR|T06377|T06 SAR DNA-binding protein-1 - garden pea,
partial (37%)
Length = 748
Score = 30.0 bits (66), Expect = 6.1
Identities = 16/42 (38%), Positives = 23/42 (54%)
Frame = +3
Query: 547 ISWSFLRETLELYEFPLVIIDLIMCYVSSSQISLLWNGAPLP 588
+SW+FL +TL+ IDLI+C SS W +P+P
Sbjct: 291 LSWAFL*QTLQ--------IDLILCLRGSSTALQAWLCSPIP 392
>AW689576 similar to GP|7940277|gb| F2401.8 {Arabidopsis thaliana}, partial
(45%)
Length = 686
Score = 29.6 bits (65), Expect = 7.9
Identities = 25/76 (32%), Positives = 37/76 (47%), Gaps = 6/76 (7%)
Frame = -3
Query: 268 GGFNVSLITDS-----RLTL*CLRLIYNWSTRKF*NRRNSYG-TRKLERTGSDWGTLTNT 321
G +++SL T S R TL CL TR ++R + T + +RTGSDW L++
Sbjct: 552 GRYHISLRTISYNFIERKTLSCLWNSKRTETRSDLSKRETLDRTNRSKRTGSDWSLLSSL 373
Query: 322 QAVIIRKRNSVHRLKL 337
+ NS RL +
Sbjct: 372 VMMRPSLPNSFFRLHI 325
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.340 0.149 0.508
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 41,409,078
Number of Sequences: 36976
Number of extensions: 612026
Number of successful extensions: 4142
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 2190
Number of HSP's successfully gapped in prelim test: 192
Number of HSP's that attempted gapping in prelim test: 1810
Number of HSP's gapped (non-prelim): 2569
length of query: 1291
length of database: 9,014,727
effective HSP length: 107
effective length of query: 1184
effective length of database: 5,058,295
effective search space: 5989021280
effective search space used: 5989021280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0296b.2


