

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0296a.5
(754 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
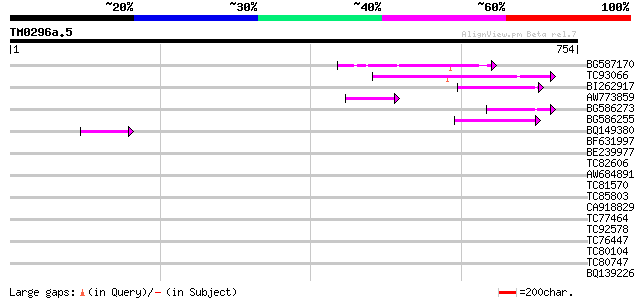
Score E
Sequences producing significant alignments: (bits) Value
BG587170 similar to PIR|F86470|F8 probable retroelement polyprot... 134 1e-31
TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative ret... 109 4e-24
BI262917 weakly similar to GP|19920130|g Putative retroelement {... 73 5e-13
AW773859 68 1e-11
BG586273 weakly similar to PIR|F86470|F8 probable retroelement p... 65 1e-10
BG586255 similar to GP|7682800|gb Hypothetical protein T15F17.l ... 59 7e-09
BQ149380 53 4e-07
BF631997 weakly similar to GP|18542925|gb Putative pol polyprote... 36 0.048
BE239977 weakly similar to GP|23237899|db polyprotein-like {Oryz... 36 0.063
TC82606 similar to GP|10177319|dbj|BAB10645. gb|AAF26800.1~gene_... 35 0.14
AW684891 34 0.18
TC81570 similar to GP|18766642|gb|AAL79042.1 NIMA-related protei... 33 0.41
TC85803 similar to GP|13785471|dbj|BAB43909. phosphoenolpyruvate... 32 1.2
CA918829 similar to GP|15450341|gb AT4g11980/F16J13_50 {Arabidop... 32 1.2
TC77464 similar to GP|13272433|gb|AAK17155.1 unknown protein {Ar... 30 2.6
TC92578 similar to GP|10177072|dbj|BAB10514. contains similarity... 30 3.4
TC76447 homologue to GP|11121502|emb|CAC14888. putative extensin... 30 3.4
TC80104 similar to GP|20197699|gb|AAD20910.3 putative LRR recept... 30 4.5
TC80747 weakly similar to GP|15217295|gb|AAK92639.1 Putative pho... 29 7.6
BQ139226 similar to GP|6625788|gb| dynamin homolog {Astragalus s... 29 7.6
>BG587170 similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 718
Score = 134 bits (338), Expect = 1e-31
Identities = 79/214 (36%), Positives = 116/214 (53%), Gaps = 3/214 (1%)
Frame = -3
Query: 437 TVSSSNKLPNIWHHRLGHPSQPCYISLQQMYPPITTSPCKNCDTCHFAKQKRLSFSLRNT 496
T SSS +WH RLGHP +L M P + KNC+ C K + F +T
Sbjct: 614 TSSSSLNKDALWHARLGHPHGR---ALNLMLPGVVFEN-KNCEACILGKHCKNVFPRTST 447
Query: 497 IYGTVLELLHVDIWGPYSVASITGAKYFHIIVYDKSRFTWIKLMKAKSEARAALQTFILY 556
+Y +L++ D+W S+ S KYF + +KS++TW+ L+ +K A + F Y
Sbjct: 446 VYENCFDLIYTDLWTAPSL-SRDNHKYFVTFIDEKSKYTWLTLIPSKDRVIDAFKNFQAY 270
Query: 557 SQRQYGSLVKIVRLDNGVEFAMTDFYSQ---HGVQHQLSCVKTPQQNGVVERKHQHILNI 613
Y + +KI+R DNG E+ F S HG+ HQ SC TPQQNGV +RK++H++ +
Sbjct: 269 VTNHYHAKIKILRSDNGGEYTSYAFKSHLDHHGILHQTSCPYTPQQNGVAKRKNKHLMEV 90
Query: 614 ARALMLQSNLTLEFWSFAVIHAVCLMNLLPTKVL 647
AR+LM Q+N++ A L+N +PTKVL
Sbjct: 89 ARSLMFQANVST---------ACYLINWIPTKVL 15
>TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative retroelement
{Oryza sativa} [Oryza sativa (japonica cultivar-group)],
partial (10%)
Length = 823
Score = 109 bits (272), Expect = 4e-24
Identities = 77/250 (30%), Positives = 121/250 (47%), Gaps = 7/250 (2%)
Frame = +1
Query: 483 FAKQKRLSFSLRNTIYGTVLELLHVDIWGPYSVASITGAKYFHIIVYDKSRFTWIKLMKA 542
F +K++SFS +L+ +H D+WGP V S G +Y I+ D R W+ ++
Sbjct: 40 FGNRKKVSFSTATHRTKGILDYIHSDLWGPSKVTSYGGRRYMMTIIDDFPRKVWVYFLRY 219
Query: 543 KSEARAALQTFILYSQRQYGSLVKIVRLDNGVEFAMTD---FYSQHGVQHQLSCVKTPQQ 599
K+E + + + + Q G VK + DN +EF +D F + HG+ + + PQQ
Sbjct: 220 KNETFPTFKKWRILVETQTGKNVKKLITDN*LEFCSSDFNEFCTNHGIARHKTIPRNPQQ 399
Query: 600 NGVVERKHQHILNIARALMLQSNL--TLEFWSFAVIHAVCLMNLLPTKVLSYSSPFIILH 657
NGV ER + +L AR ++ + L + W A A L+N P L + P I
Sbjct: 400 NGVAERMIRTLLERARCMLSNAGL*N*RDLWVEAASTACHLVNRSPHSALDFKVPEDIWS 579
Query: 658 NLKPDIEHLKVFGSLCYATSLTAHRTKFPLRARKCIVLGQKEGGVKGYLMF--DLKTREV 715
D +L++FG YA + K RA +CI L KGY ++ D K++++
Sbjct: 580 GNLVDYSNLRIFGCPAYA---LVNDGKLAPRAGECIFLSYASES-KGYRLWCSDPKSQKL 747
Query: 716 FLSRNVIFYE 725
LSR+V F E
Sbjct: 748 ILSRDVTFNE 777
>BI262917 weakly similar to GP|19920130|g Putative retroelement {Oryza
sativa} [Oryza sativa (japonica cultivar-group)],
partial (8%)
Length = 426
Score = 72.8 bits (177), Expect = 5e-13
Identities = 41/114 (35%), Positives = 61/114 (52%)
Frame = +1
Query: 596 TPQQNGVVERKHQHILNIARALMLQSNLTLEFWSFAVIHAVCLMNLLPTKVLSYSSPFII 655
TPQQNGV ER ++ +L RA++ + + FW+ AV A ++N P+ V+ +P +
Sbjct: 85 TPQQNGVAERMNRTLLERTRAMLKTAGMAKSFWAEAVKTACYVINRSPSTVIDLKTPMEM 264
Query: 656 LHNLKPDIEHLKVFGSLCYATSLTAHRTKFPLRARKCIVLGQKEGGVKGYLMFD 709
D L VFG Y + RTK ++RKCI LG + VKGY ++D
Sbjct: 265 WKGKPVDYSSLHVFGCPVYVMYNSQERTKLDPKSRKCIFLGYAD-NVKGYXLWD 423
>AW773859
Length = 538
Score = 68.2 bits (165), Expect = 1e-11
Identities = 31/72 (43%), Positives = 39/72 (54%)
Frame = -3
Query: 447 IWHHRLGHPSQPCYISLQQMYPPITTSPCKNCDTCHFAKQKRLSFSLRNTIYGTVLELLH 506
+WH RLGH S +SL +P IT CD CH+++ K+L F L EL H
Sbjct: 230 LWHFRLGHLSNRKLLSLHSNFPFITIDQNSVCDICHYSRHKKLPFQLSTNRASKCYELFH 51
Query: 507 VDIWGPYSVASI 518
DIWGP+S SI
Sbjct: 50 FDIWGPFSTQSI 15
>BG586273 weakly similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 705
Score = 65.1 bits (157), Expect = 1e-10
Identities = 35/91 (38%), Positives = 52/91 (56%)
Frame = -2
Query: 635 AVCLMNLLPTKVLSYSSPFIILHNLKPDIEHLKVFGSLCYATSLTAHRTKFPLRARKCIV 694
A L+N +PT+VL +PF +L+ KP + +++VFG LCY R K R+RK +
Sbjct: 704 ACYLINRIPTRVLKDQAPFEVLNQRKPSLTYMRVFGCLCYVLVPGELRNKLEARSRKAMF 525
Query: 695 LGQKEGGVKGYLMFDLKTREVFLSRNVIFYE 725
+G KGY +D + R V +SR+V F E
Sbjct: 524 IGYST-TQKGYKCYDPEARRVLVSRDVKFIE 435
>BG586255 similar to GP|7682800|gb Hypothetical protein T15F17.l {Arabidopsis
thaliana}, partial (12%)
Length = 436
Score = 58.9 bits (141), Expect = 7e-09
Identities = 38/115 (33%), Positives = 54/115 (46%)
Frame = -1
Query: 592 SCVKTPQQNGVVERKHQHILNIARALMLQSNLTLEFWSFAVIHAVCLMNLLPTKVLSYSS 651
S V QNG+ E + I I R L+++S + W AV+HA L+ + P+ Y S
Sbjct: 397 SVVHVHTQNGLAESFIKRIQLITRPLLMRSKQPVSAWGHAVLHAAELIRIRPSSEHKY-S 221
Query: 652 PFIILHNLKPDIEHLKVFGSLCYATSLTAHRTKFPLRARKCIVLGQKEGGVKGYL 706
P +L PD+ H+K FG Y HRTK + R I +G + YL
Sbjct: 220 PSQLLSGHVPDVSHIKTFGYPVYVPIAPPHRTKMGPQRRMGIYVGFDSPTIIKYL 56
>BQ149380
Length = 419
Score = 53.1 bits (126), Expect = 4e-07
Identities = 26/70 (37%), Positives = 40/70 (57%)
Frame = +3
Query: 95 PHACCCEVVKIFKQYRENDCVLCFLRGLNDNFAAARS*ILLMEPLPTLSKICSMIIQQER 154
P C C ++ K D ++ FL GLND++ S +LLM PLP ++ + SM++QQER
Sbjct: 129 PIQCTCFAMRNNKALTAEDRIIKFLMGLNDDYQGVTSKVLLMYPLPQINMVFSMVMQQER 308
Query: 155 HLGIGVLHEP 164
+ V+ P
Sbjct: 309 KMQ*NVVSSP 338
>BF631997 weakly similar to GP|18542925|gb Putative pol polyprotein {Oryza
sativa}, partial (6%)
Length = 650
Score = 36.2 bits (82), Expect = 0.048
Identities = 22/70 (31%), Positives = 36/70 (51%), Gaps = 3/70 (4%)
Frame = -2
Query: 539 LMKAKSEARAALQTFILYSQRQYGSLVKIVRLDNGVEFAMTDF---YSQHGVQHQLSCVK 595
+MK KSEA + +S+ + G + +L+NG+E +F + + Q +
Sbjct: 640 MMKHKSEAFYFSRNGRFWSRVKQGRR*NVFKLNNGLEICSAEFNELCKEEHITRQYTVRN 461
Query: 596 TPQQNGVVER 605
TPQ+NGV ER
Sbjct: 460 TPQKNGVAER 431
>BE239977 weakly similar to GP|23237899|db polyprotein-like {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 514
Score = 35.8 bits (81), Expect = 0.063
Identities = 22/69 (31%), Positives = 37/69 (52%)
Frame = -2
Query: 657 HNLKPDIEHLKVFGSLCYATSLTAHRTKFPLRARKCIVLGQKEGGVKGYLMFDLKTREVF 716
+NL P ++FG + + R+KF RA KC+ + + KGY + +R+ F
Sbjct: 471 NNLTP-----RIFGCTSFVHIHSDGRSKFDHRALKCVFI-RYSSTQKGYRCYHPPSRKYF 310
Query: 717 LSRNVIFYE 725
+SR+V F+E
Sbjct: 309 VSRDVTFHE 283
>TC82606 similar to GP|10177319|dbj|BAB10645.
gb|AAF26800.1~gene_id:MSL3.8~similar to unknown protein
{Arabidopsis thaliana}, partial (15%)
Length = 672
Score = 34.7 bits (78), Expect = 0.14
Identities = 33/126 (26%), Positives = 53/126 (41%), Gaps = 12/126 (9%)
Frame = +1
Query: 218 HGYPSNFPNGYKPEKAYIPTSGGE-SHSSSDQDEPPSSSLELTREYLQGLLALL--PQSK 274
H PSNF + + + P S + H SS+ P S+ E + LL P S
Sbjct: 118 HINPSNFSSHFHNRALFTPPSLFQIEHFSSESKPEPVSTTEPDPIAIAISSELLKEPDSD 297
Query: 275 PTAVAQTPNLKPISTSNLVSSHNVIANNIGMVNS---------QWIMDSGATDHIASSLS 325
P++V Q NL S S++ + N+I + + QW+ + H +LS
Sbjct: 298 PSSVTQRLNL---SFSHITPTPNLILETLNLSLEAGRNVLGFHQWLASNPKFTHTDETLS 468
Query: 326 FFSSYY 331
+F Y+
Sbjct: 469 YFVDYF 486
>AW684891
Length = 539
Score = 34.3 bits (77), Expect = 0.18
Identities = 23/91 (25%), Positives = 45/91 (49%)
Frame = -3
Query: 514 SVASITGAKYFHIIVYDKSRFTWIKLMKAKSEARAALQTFILYSQRQYGSLVKIVRLDNG 573
++ I G K+F +++ + T + L K + + F+ +R + K+
Sbjct: 366 TLIQIYGXKWFLAFIHNSTSIT*LFLKKHEIWFQ-----FVKSIERLCSNKGKVY----- 217
Query: 574 VEFAMTDFYSQHGVQHQLSCVKTPQQNGVVE 604
V+ ++ + ++GV +L CV +PQQNGV E
Sbjct: 216 VDQNLSQYLEENGVSQELKCVNSPQQNGVSE 124
>TC81570 similar to GP|18766642|gb|AAL79042.1 NIMA-related protein kinase
{Populus x canescens}, partial (20%)
Length = 895
Score = 33.1 bits (74), Expect = 0.41
Identities = 28/92 (30%), Positives = 41/92 (44%), Gaps = 2/92 (2%)
Frame = +3
Query: 178 HSNSNRGRSNSQYSSSKSNV*RLCTHCGRQNHTVETCFLKHGYPSNFPNGYKPEKAYIPT 237
HS+++ G ++ + K CT + TV T G FP G E + T
Sbjct: 276 HSSTSTGSADCTITKDK------CTILVDKKVTVPTSITDAGTDVRFPKGSASECSNYVT 437
Query: 238 SGGESHSSSD--QDEPPSSSLELTREYLQGLL 267
+G S SSS+ Q +SS + E L+GLL
Sbjct: 438 TGVSSRSSSESRQHRFDTSSYQQRAEALEGLL 533
>TC85803 similar to GP|13785471|dbj|BAB43909. phosphoenolpyruvate
carboxykinase {Flaveria pringlei}, partial (95%)
Length = 2351
Score = 31.6 bits (70), Expect = 1.2
Identities = 21/69 (30%), Positives = 30/69 (43%), Gaps = 4/69 (5%)
Frame = -3
Query: 429 HFHSVNMSTVSSSNKLPNIWHHRLGHPSQPCYISL----QQMYPPITTSPCKNCDTCHFA 484
H ++ + T S + N+ + +PSQP I L YPP SP N HF
Sbjct: 2286 HMNNTHRKTTSKAKTSTNVKTRNM-NPSQPTNILL*IRPHLFYPPNNPSPTHNTTLLHFK 2110
Query: 485 KQKRLSFSL 493
+ +L SL
Sbjct: 2109 RLDQLQESL 2083
>CA918829 similar to GP|15450341|gb AT4g11980/F16J13_50 {Arabidopsis
thaliana}, partial (54%)
Length = 635
Score = 31.6 bits (70), Expect = 1.2
Identities = 13/49 (26%), Positives = 26/49 (52%)
Frame = +2
Query: 251 PPSSSLELTREYLQGLLALLPQSKPTAVAQTPNLKPISTSNLVSSHNVI 299
P S + R + +L+P S T++ P + P+S+SN+ + ++ I
Sbjct: 350 PVEESRKAVRSTMSSTFSLMPVSSSTSLTAVPTISPLSSSNIPAGNSKI 496
>TC77464 similar to GP|13272433|gb|AAK17155.1 unknown protein {Arabidopsis
thaliana}, partial (24%)
Length = 1109
Score = 30.4 bits (67), Expect = 2.6
Identities = 11/36 (30%), Positives = 20/36 (55%)
Frame = +3
Query: 416 YTSWSFHLSLVSNHFHSVNMSTVSSSNKLPNIWHHR 451
++S FHL + H+H+ ++V L +WHH+
Sbjct: 27 FSSLHFHLPFTTTHYHNCYTASVV*PKTLSLLWHHQ 134
>TC92578 similar to GP|10177072|dbj|BAB10514. contains similarity to unknown
protein~dbj|BAA91048.1~gene_id:MKP11.12 {Arabidopsis
thaliana}, partial (36%)
Length = 1126
Score = 30.0 bits (66), Expect = 3.4
Identities = 19/74 (25%), Positives = 42/74 (56%), Gaps = 6/74 (8%)
Frame = +1
Query: 605 RKHQHILNIARALMLQSNLTLEFWSFAV-----IHAVCLMNLLPT-KVLSYSSPFIILHN 658
R+ Q +LN + ++ + +TL++ + V +HA+ + PT +L ++ + +H
Sbjct: 475 RRKQQLLNSSGSVFQKDQMTLDYGAPWVTLPIMMHAMKKLWKFPTIGLLELNALWPAVHT 654
Query: 659 LKPDIEHLKVFGSL 672
++ ++ HLK +GSL
Sbjct: 655 IEENMRHLKFYGSL 696
>TC76447 homologue to GP|11121502|emb|CAC14888. putative extensin {Nicotiana
sylvestris}, partial (58%)
Length = 564
Score = 30.0 bits (66), Expect = 3.4
Identities = 18/67 (26%), Positives = 32/67 (46%)
Frame = +3
Query: 416 YTSWSFHLSLVSNHFHSVNMSTVSSSNKLPNIWHHRLGHPSQPCYISLQQMYPPITTSPC 475
+TS + L L ++H H +++S L HH+ S P ++S ++ +TSP
Sbjct: 24 FTSTNHPLHLFTHHLHL--FTSISHHLHLSTHHHHQFTSTSHPLHLSTHHLHQFTSTSPH 197
Query: 476 KNCDTCH 482
+ T H
Sbjct: 198 LHPSTHH 218
Score = 28.5 bits (62), Expect = 10.0
Identities = 24/84 (28%), Positives = 37/84 (43%), Gaps = 1/84 (1%)
Frame = +3
Query: 416 YTSWSFHLSLVSNHFHSVNMSTVSSSNKLPNIWHH-RLGHPSQPCYISLQQMYPPITTSP 474
+TS S HL ++H H T +S + P+ HH + S + S ++P TSP
Sbjct: 279 FTSTSPHLHPSTHHLHQF---TSTSPHLHPSTHHHHQFTSTSHLLHPSTHHLHPFTNTSP 449
Query: 475 CKNCDTCHFAKQKRLSFSLRNTIY 498
+ T H + S L +IY
Sbjct: 450 HLHPSTHHLPRFISTSHHLLPSIY 521
>TC80104 similar to GP|20197699|gb|AAD20910.3 putative LRR receptor protein
kinase {Arabidopsis thaliana}, partial (28%)
Length = 1549
Score = 29.6 bits (65), Expect = 4.5
Identities = 13/33 (39%), Positives = 14/33 (42%)
Frame = -1
Query: 449 HHRLGHPSQPCYISLQQMYPPITTSPCKNCDTC 481
HH G C +SLQQ PC C TC
Sbjct: 1315 HHPSGEQRLFCQVSLQQAASHQEARPCTQCPTC 1217
>TC80747 weakly similar to GP|15217295|gb|AAK92639.1 Putative
phosphoinositide phosphatase {Oryza sativa} [Oryza
sativa (japonica cultivar-group)], partial (7%)
Length = 779
Score = 28.9 bits (63), Expect = 7.6
Identities = 12/38 (31%), Positives = 24/38 (62%)
Frame = +1
Query: 609 HILNIARALMLQSNLTLEFWSFAVIHAVCLMNLLPTKV 646
++L + + ++ S L EFW F ++HAV ++L P ++
Sbjct: 367 NMLYLKQPRLIMSVLHCEFW*FCLLHAVLAVSLEPPEI 480
>BQ139226 similar to GP|6625788|gb| dynamin homolog {Astragalus sinicus},
partial (12%)
Length = 674
Score = 28.9 bits (63), Expect = 7.6
Identities = 12/29 (41%), Positives = 19/29 (65%)
Frame = +3
Query: 273 SKPTAVAQTPNLKPISTSNLVSSHNVIAN 301
S PT PN P+S+++LVS+ +V+ N
Sbjct: 189 SSPTKTLMNPNALPLSSTSLVSAMSVLEN 275
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.337 0.145 0.465
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,782,007
Number of Sequences: 36976
Number of extensions: 462681
Number of successful extensions: 4414
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 2476
Number of HSP's successfully gapped in prelim test: 253
Number of HSP's that attempted gapping in prelim test: 1853
Number of HSP's gapped (non-prelim): 2872
length of query: 754
length of database: 9,014,727
effective HSP length: 103
effective length of query: 651
effective length of database: 5,206,199
effective search space: 3389235549
effective search space used: 3389235549
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0296a.5


