

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0290.7
(47 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
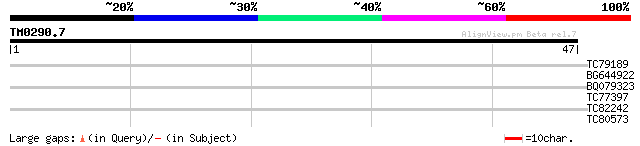
Score E
Sequences producing significant alignments: (bits) Value
TC79189 similar to GP|21593647|gb|AAM65614.1 replication protein... 30 0.15
BG644922 similar to GP|8778232|gb F10B6.19 {Arabidopsis thaliana... 27 2.2
BQ079323 similar to PIR|F81738|F817 hypothetical protein TC0114 ... 25 4.9
TC77397 homologue to PIR|S37159|S37159 NADPH--ferrihemoprotein r... 25 4.9
TC82242 weakly similar to GP|21592781|gb|AAM64730.1 putative mem... 25 6.4
TC80573 similar to GP|9759187|dbj|BAB09724.1 importin beta {Arab... 25 8.3
>TC79189 similar to GP|21593647|gb|AAM65614.1 replication protein A1-like
{Arabidopsis thaliana}, partial (79%)
Length = 1251
Score = 30.4 bits (67), Expect = 0.15
Identities = 15/34 (44%), Positives = 21/34 (61%), Gaps = 1/34 (2%)
Frame = +1
Query: 4 INPDYGFNPTG-WEVQELGFKSKPNVWAYFSVKS 36
++ D FNP ++ QE FKS PN W+Y S K+
Sbjct: 913 VHSDM*FNPASCFDSQESRFKSHPN*WSYESAKA 1014
>BG644922 similar to GP|8778232|gb F10B6.19 {Arabidopsis thaliana}, partial
(20%)
Length = 788
Score = 26.6 bits (57), Expect = 2.2
Identities = 10/30 (33%), Positives = 18/30 (59%), Gaps = 2/30 (6%)
Frame = -1
Query: 5 NPDYGFNPTGWEVQEL--GFKSKPNVWAYF 32
N ++ F P W+VQE+ F + ++W Y+
Sbjct: 680 NHNHIFEPLSWQVQEIFCQFHQRRHIWFYY 591
>BQ079323 similar to PIR|F81738|F817 hypothetical protein TC0114 [imported] -
Chlamydia muridarum (strain Nigg), partial (57%)
Length = 298
Score = 25.4 bits (54), Expect = 4.9
Identities = 13/27 (48%), Positives = 17/27 (62%)
Frame = +1
Query: 8 YGFNPTGWEVQELGFKSKPNVWAYFSV 34
YG +P EV+ELG K + VW+ SV
Sbjct: 139 YGCSPFK-EVRELGLKRRETVWSLSSV 216
>TC77397 homologue to PIR|S37159|S37159 NADPH--ferrihemoprotein reductase (EC
1.6.2.4) - spring vetch, complete
Length = 2519
Score = 25.4 bits (54), Expect = 4.9
Identities = 7/22 (31%), Positives = 15/22 (67%)
Frame = -3
Query: 15 WEVQELGFKSKPNVWAYFSVKS 36
WE + + +P++W++F+V S
Sbjct: 2169 WENHQTSLR*RPSIWSFFTVAS 2104
>TC82242 weakly similar to GP|21592781|gb|AAM64730.1 putative membrane
protein {Arabidopsis thaliana}, partial (9%)
Length = 580
Score = 25.0 bits (53), Expect = 6.4
Identities = 9/17 (52%), Positives = 11/17 (63%)
Frame = -2
Query: 19 ELGFKSKPNVWAYFSVK 35
+LGFK K N+W VK
Sbjct: 90 DLGFKRKANIWIVLKVK 40
>TC80573 similar to GP|9759187|dbj|BAB09724.1 importin beta {Arabidopsis
thaliana}, partial (27%)
Length = 1220
Score = 24.6 bits (52), Expect = 8.3
Identities = 9/11 (81%), Positives = 10/11 (90%)
Frame = -1
Query: 23 KSKPNVWAYFS 33
KSKP+VWAY S
Sbjct: 359 KSKPDVWAYIS 327
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.142 0.463
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,740,457
Number of Sequences: 36976
Number of extensions: 16888
Number of successful extensions: 74
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 74
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 74
length of query: 47
length of database: 9,014,727
effective HSP length: 23
effective length of query: 24
effective length of database: 8,164,279
effective search space: 195942696
effective search space used: 195942696
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0290.7


