

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0288.8
(846 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
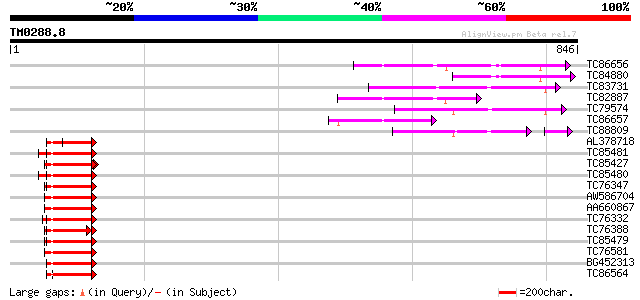
Score E
Sequences producing significant alignments: (bits) Value
TC86656 similar to GP|7108521|gb|AAF36454.1| ubiquitin-protein l... 177 1e-44
TC84880 similar to PIR|T01491|T01491 ubiquitin--protein ligase h... 116 3e-26
TC83731 similar to PIR|T46155|T46155 hypothetical protein T4D2.2... 112 8e-25
TC82887 similar to GP|7108521|gb|AAF36454.1| ubiquitin-protein l... 104 1e-22
TC79574 similar to PIR|T05688|T05688 hypothetical protein F20M13... 86 5e-17
TC86657 similar to GP|7108521|gb|AAF36454.1| ubiquitin-protein l... 78 1e-14
TC88809 similar to PIR|T05688|T05688 hypothetical protein F20M13... 60 4e-11
AL378718 homologue to GP|6138833|emb Polyubiquitin {Plasmodium f... 64 2e-10
TC85481 homologue to PIR|S17435|S17435 polyubiquitin 6 - common ... 64 2e-10
TC85427 chitinase 64 2e-10
TC85480 GP|170352|gb|AAA34123.1|| hexameric polyubiquitin {Nicot... 64 2e-10
TC76347 PIR|S17435|S17435 polyubiquitin 6 - common sunflower, pa... 64 2e-10
AW586704 PIR|S17435|S17 polyubiquitin 6 - common sunflower, part... 64 2e-10
AA660867 homologue to PIR|S17435|S17 polyubiquitin 6 - common su... 64 2e-10
TC76332 GP|15450587|gb|AAK96565.1 AT4g05050/T32N4_13 {Arabidopsi... 64 3e-10
TC76388 homologue to PIR|S17435|S17435 polyubiquitin 6 - common ... 63 4e-10
TC85479 PIR|S20925|S20925 polyubiquitin - maize, complete 63 4e-10
TC76581 homologue to PIR|S40240|S40240 ubiquitin/ribosomal prote... 63 5e-10
BG452313 homologue to PIR|T40261|T40 ubi4 protein - fission yeas... 62 7e-10
TC86564 PIR|C86439|C86439 protein T19E23.13 [imported] - Arabido... 62 9e-10
>TC86656 similar to GP|7108521|gb|AAF36454.1| ubiquitin-protein ligase 1
{Arabidopsis thaliana}, partial (12%)
Length = 1939
Score = 177 bits (450), Expect = 1e-44
Identities = 114/338 (33%), Positives = 176/338 (51%), Gaps = 13/338 (3%)
Frame = +1
Query: 513 GLFMEFKNEEATGPGVL-REWFLLVCQALFNPQNALFVACPNDRRRFYPNHASKVHPLHL 571
GL + F+ EE G L REW+ L+ + +F+ LF N+ F PN S HL
Sbjct: 760 GLTVHFQGEEGIDAGGLTREWYQLLSRVIFDKGALLFTTVGNEST-FQPNPNSVYQTEHL 936
Query: 572 EYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKHITLEDVKDADPFLYSSCKQILEMD 631
YF F GRV+ AL + + R F+ + G +T D++ DP + + K +LE D
Sbjct: 937 SYFKFIGRVVGKALFDGQLLDVHFTRSFYKHILGVKVTYHDIEAIDPDYFKNLKWMLEND 1116
Query: 632 SDFIDSDALGLTFVREVEE-----LGDTKVV--ELCPGGKNLIVNSKNREKYIDLLIKDR 684
SD L LTF + +E T+V EL PGG+N+ V +N+ +Y+DL+ + R
Sbjct: 1117 I----SDVLDLTFSIDADEEKLILYERTEVTDYELIPGGRNIKVTEENKHQYVDLVAEHR 1284
Query: 685 FVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLHGSEDTISVEDWKAHTEY 744
T+I Q++ F +GF++++ F + LEL ++ G D I ++D +A+TEY
Sbjct: 1285 LTTAIRPQINAFLEGFSELIPRELISIFNDKELEL-----LISGLPD-IDLDDLRANTEY 1446
Query: 745 NGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGFCGL-----ASRLYIYKS 799
+GY I WFWE+V ++ E + LL F T +P+EGF L + + I+K+
Sbjct: 1447 SGYSAASPVIQWFWEVVQGLSKEDKARLLQFVTGTSKVPLEGFSALQGISGSQKFQIHKA 1626
Query: 800 PEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQE 837
D LPS+HTCF ++ P Y S ++ERL + E
Sbjct: 1627 YGSPDHLPSAHTCFNQLDLPEYPSKQHLEERLLLAIHE 1740
>TC84880 similar to PIR|T01491|T01491 ubiquitin--protein ligase homolog
F17O7.15 - Arabidopsis thaliana, partial (17%)
Length = 801
Score = 116 bits (291), Expect = 3e-26
Identities = 67/190 (35%), Positives = 105/190 (55%), Gaps = 6/190 (3%)
Frame = +1
Query: 661 PGGKNLIVNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELE 720
PGG+N + +N+ +Y+DL+ + R T+I Q++ F +GF +I+ F + LEL
Sbjct: 43 PGGRNTKITEENKHQYVDLVAEHRLTTAIRPQINAFLEGFGEIIPKELISIFNDKELEL- 219
Query: 721 DLDWMLHGSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVK 780
++ G D I ++D +A+TEY+GY I WFWE+V + E + LL F T
Sbjct: 220 ----LISGLPD-IDLDDLRANTEYSGYSAASPVIQWFWEVVQGFSKEDKARLLQFVTGTS 384
Query: 781 YLPVEGFCGL-----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEV-I 834
+P+EGF L A + I+K+ D LPS+HTCF ++ P Y S ++ERL + I
Sbjct: 385 KVPLEGFSALQGISGAQKFQIHKAYGSSDHLPSAHTCFNQLDLPEYPSKQHLEERLLLAI 564
Query: 835 TQEHIGCSFG 844
+ + G FG
Sbjct: 565 HEANEGFGFG 594
>TC83731 similar to PIR|T46155|T46155 hypothetical protein T4D2.20 -
Arabidopsis thaliana, partial (27%)
Length = 954
Score = 112 bits (279), Expect = 8e-25
Identities = 82/298 (27%), Positives = 134/298 (44%), Gaps = 12/298 (4%)
Frame = +2
Query: 536 VCQALFNPQNALFVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVL 595
+ + F+P+ LF PN +++ L+ F GRV+ AL + +
Sbjct: 8 ISKEAFSPEYGLFSQTSTSDSLLIPNASARFLDNGLQMIEFLGRVVGKALYEGILLDYSF 187
Query: 596 DRVFFLQLAGKHITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTK 655
VF +L G++ L+++ DP LY S + D D + L L F E G
Sbjct: 188 SHVFVQKLLGRYSFLDELSTLDPELYRSLMYVKNYDGDVKE---LSLDFTVTEESFGKRH 358
Query: 656 VVELCPGGKNLIVNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQ 715
VVEL GGK++ V ++N+ +YI + + I + F +G TD++S S + F
Sbjct: 359 VVELKSGGKDISVTNENKMQYIHAMADYKLNQQILLFSNAFYRGLTDLISPSWLKLF--- 529
Query: 716 SLELEDLDWMLHGSEDTISVEDWKAHTEY-NGYKETDIQITWFWEIVGRMTAEQRKVLLF 774
+ + +L G I ++D+K++T Y GY E I FWE++ ++R +LL
Sbjct: 530 --NASEFNQLLSGGNYDIDIDDFKSNTRYTGGYNEGSRTIKIFWEVIKGFEPKERCMLLK 703
Query: 775 FWTSVKYLPVEGFCGLASRLYIYK-----------SPEPEDRLPSSHTCFYRMCFPAY 821
F TS P+ GF L I+K + +RL S+ TC+ + P Y
Sbjct: 704 FVTSCSRGPLLGFKYLQPPFTIHKVACDVPLWATIGGQDAERLSSASTCYNTLKLPTY 877
>TC82887 similar to GP|7108521|gb|AAF36454.1| ubiquitin-protein ligase 1
{Arabidopsis thaliana}, partial (8%)
Length = 937
Score = 104 bits (260), Expect = 1e-22
Identities = 66/224 (29%), Positives = 110/224 (48%), Gaps = 8/224 (3%)
Frame = +2
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATGPGVL-REWFLLVCQALFNPQNAL 547
+ R+ +L +S+ + T L + L ++F+ EE G L REW+ L+ + +F+ L
Sbjct: 245 VRRAYILEDSYNQLRMRPTQDLKSRLNVQFQGEEGIDAGGLTREWYQLLSRVIFDKGALL 424
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F N+ F PN S HL YF F GRV+ AL + + R F+ + G
Sbjct: 425 FTTVGNNAT-FQPNPNSVYQTEHLSYFKFVGRVVGKALFDGQLLDVYFTRSFYKHILGVK 601
Query: 608 ITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVE-------ELGDTKVVELC 660
+T D++ DP Y + K +LE D SD LTF + + E + EL
Sbjct: 602 VTYHDIEAVDPDYYKNLKWMLENDV----SDIPDLTFNMDADEEKLILYEKNEVTDYELK 769
Query: 661 PGGKNLIVNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDIL 704
PGG+N+ V + + +Y+DL+ + +I Q++ F +GF +++
Sbjct: 770 PGGRNIKVTEETKHEYVDLVAEHLLTNAIRAQINSFLEGFNEMV 901
>TC79574 similar to PIR|T05688|T05688 hypothetical protein F20M13.160 -
Arabidopsis thaliana, partial (38%)
Length = 1412
Score = 86.3 bits (212), Expect = 5e-17
Identities = 72/278 (25%), Positives = 121/278 (42%), Gaps = 20/278 (7%)
Frame = +2
Query: 574 FGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKHITLEDVKDADPFLYSSCK--QILEMD 631
F G+V+A AL + + + F+ + GK + L D++ DP L Q L
Sbjct: 218 FFLLGQVVAKALQDGRVLDLHFSKAFYKLILGKELYLYDIQSLDPELGRVLHEFQALVNR 397
Query: 632 SDFIDSDALGLTFVREVEELGDTKVVELC--------------PGGKNLIVNSKNREKYI 677
++S G + + + D+++ +LC G + +VN +N E Y+
Sbjct: 398 KFCLESVCEGNSELEQGLSFRDSRIEDLCLDFTLPGYPDIVLASGSDHTMVNMRNLEDYV 577
Query: 678 DLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLHGSEDTISVED 737
L++ + IS QV F GF + Q F+ E+L+ +L G +D+ ++ +
Sbjct: 578 SLIVDATVKSGISRQVEAFKSGFNQVFPIENLQIFYE-----EELERILCGEDDSWAINE 742
Query: 738 WKAHTEYN-GYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGFCGLASRLYI 796
H +++ GY + I EI+ QR+ L F T LP G L +L I
Sbjct: 743 LADHIKFDHGYTASSPPIVNLLEIIREFDHGQRRAFLQFVTGTPRLPPGGLASLNPKLTI 922
Query: 797 YK---SPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERL 831
+ S + + LPS TC + P YSS M+E+L
Sbjct: 923 VRKHCSNQADTDLPSVMTCANYLKLPPYSSKERMKEKL 1036
>TC86657 similar to GP|7108521|gb|AAF36454.1| ubiquitin-protein ligase 1
{Arabidopsis thaliana}, partial (6%)
Length = 1302
Score = 78.2 bits (191), Expect = 1e-14
Identities = 50/166 (30%), Positives = 80/166 (48%), Gaps = 5/166 (3%)
Frame = +3
Query: 476 EVKEDYEELHEML---IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATGPGVL-RE 531
++K ++ H L + R+ +L +S+ + T L L + F+ EE G L RE
Sbjct: 804 KIKHQHDHHHSPLRISVRRAYVLEDSYNQLRMRSTQDLKGRLTVHFQGEEGIDAGGLTRE 983
Query: 532 WFLLVCQALFNPQNALFVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQV 591
W+ L+ + +F+ LF N+ F PN S HL YF F GRV+ AL +
Sbjct: 984 WYQLLSRVIFDKGALLFTTVGNEST-FQPNPNSVYQTEHLSYFKFVGRVVGKALFDGQLL 1160
Query: 592 GIVLDRVFFLQLAGKHITLEDVKDADPFLYSSCKQILEMD-SDFID 636
+ R F+ + G +T D++ DP + + K +LE D SD +D
Sbjct: 1161DVHFTRSFYKHILGAKVTYHDIEAIDPAYFKNLKWLLENDISDVLD 1298
>TC88809 similar to PIR|T05688|T05688 hypothetical protein F20M13.160 -
Arabidopsis thaliana, partial (40%)
Length = 1406
Score = 59.7 bits (143), Expect(2) = 4e-11
Identities = 58/227 (25%), Positives = 94/227 (40%), Gaps = 19/227 (8%)
Frame = +2
Query: 571 LEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKHITLEDVKDADPFLYSSCKQI--L 628
+EYF GRV+A AL + + L F+ + G+ + L D+ D L + +++ L
Sbjct: 191 IEYFRLLGRVVAKALQDGRLLDLPLSVAFYKLVLGQDLDLHDILYVDAELGKTLQELNAL 370
Query: 629 EMDSDFIDSDALGLTFVREVEELGDTKVVELC--------------PGGKNLIVNSKNRE 674
I+S G T + +LC PG + IV+ N E
Sbjct: 371 VCRKHNIESIGGGNTGTVSNLHYRGAPIADLCLDFTLPGYPEYTLKPGDE--IVDLNNLE 544
Query: 675 KYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLHGSEDTIS 734
YI +++ T I+ Q+ F GF + S Q F +LD++L G +
Sbjct: 545 DYISMVVDATVKTGITRQLEAFRAGFNQVFDISSLQIF-----TPHELDYLLCGRRELWK 709
Query: 735 VEDWKAHTEY-NGYKETDIQITWFWEIVGRMTAEQRKVLL--FFWTS 778
E H ++ +GY I EI+G T EQ+ LL +W++
Sbjct: 710 TETLADHIKFDHGYTAKSPAIVNLLEIMGEFTPEQQPCLLSICYWST 850
Score = 26.6 bits (57), Expect(2) = 4e-11
Identities = 14/42 (33%), Positives = 20/42 (47%)
Frame = +3
Query: 799 SPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQEHIG 840
S +D LPS TC + P YS+ +M ++L E G
Sbjct: 954 SETADDDLPSVMTCANYLKLPPYSTKEIMHKKLMYAINEGQG 1079
>AL378718 homologue to GP|6138833|emb Polyubiquitin {Plasmodium falciparum
3D7}, partial (33%)
Length = 432
Score = 63.9 bits (154), Expect = 2e-10
Identities = 32/75 (42%), Positives = 51/75 (67%)
Frame = +1
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI + P D+++++ +IQ +GIP +QRLI+ GKQL +TLA+
Sbjct: 157 MQIFVKTL-TGKTITLDVEPSDSIENVKAKIQDKEGIPPDQQRLIFAGKQLDDGRTLADY 333
Query: 115 SIGKDASLQLVGRMR 129
+I K+A+L LV R+R
Sbjct: 334 NIQKEATLHLVLRLR 378
Score = 47.0 bits (110), Expect = 3e-05
Identities = 24/50 (48%), Positives = 36/50 (72%)
Frame = +1
Query: 80 SIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAECSIGKDASLQLVGRMR 129
++ +IQ +GIP +QRLI+ GKQL +TLA+ +I K+A+L LV R+R
Sbjct: 1 NVKAKIQDKEGIPPDQQRLIFAGKQLDDGRTLADYNIQKEATLHLVLRLR 150
>TC85481 homologue to PIR|S17435|S17435 polyubiquitin 6 - common sunflower,
partial (41%)
Length = 620
Score = 63.9 bits (154), Expect = 2e-10
Identities = 32/87 (36%), Positives = 57/87 (64%)
Frame = +2
Query: 43 AVYSAVHHRSIMLQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNG 102
+++S+ + +Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ G
Sbjct: 23 SLFSSSFYLRFKMQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAG 199
Query: 103 KQLQSEQTLAECSIGKDASLQLVGRMR 129
KQL+ +TLA+ +I K+++L LV R+R
Sbjct: 200 KQLEDGRTLADYNIQKESTLHLVLRLR 280
Score = 62.0 bits (149), Expect = 9e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +2
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 287 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 463
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 464 NIQKESTLHLVLRLR 508
>TC85427 chitinase
Length = 1090
Score = 63.9 bits (154), Expect = 2e-10
Identities = 32/77 (41%), Positives = 53/77 (68%)
Frame = +3
Query: 53 IMLQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLA 112
I++Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA
Sbjct: 264 IIMQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLA 440
Query: 113 ECSIGKDASLQLVGRMR 129
+ +I K+++L LV R+R
Sbjct: 441 DYNIQKESTLHLVLRLR 491
Score = 63.5 bits (153), Expect = 3e-10
Identities = 32/78 (41%), Positives = 52/78 (66%)
Frame = +3
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 726 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 902
Query: 115 SIGKDASLQLVGRMRSTG 132
+I K+++L LV R+R G
Sbjct: 903 NIQKESTLHLVLRLRGGG 956
Score = 62.0 bits (149), Expect = 9e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +3
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 498 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 674
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 675 NIQKESTLHLVLRLR 719
>TC85480 GP|170352|gb|AAA34123.1|| hexameric polyubiquitin {Nicotiana
sylvestris}, complete
Length = 1695
Score = 63.9 bits (154), Expect = 2e-10
Identities = 32/87 (36%), Positives = 57/87 (64%)
Frame = +3
Query: 43 AVYSAVHHRSIMLQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNG 102
+++S+ + +Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ G
Sbjct: 33 SLFSSSFYLRFKMQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAG 209
Query: 103 KQLQSEQTLAECSIGKDASLQLVGRMR 129
KQL+ +TLA+ +I K+++L LV R+R
Sbjct: 210 KQLEDGRTLADYNIQKESTLHLVLRLR 290
Score = 62.0 bits (149), Expect = 9e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +3
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 525 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 701
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 702 NIQKESTLHLVLRLR 746
Score = 62.0 bits (149), Expect = 9e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +3
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 753 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 929
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 930 NIQKESTLHLVLRLR 974
Score = 62.0 bits (149), Expect = 9e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +3
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 981 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 1157
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 1158 NIQKESTLHLVLRLR 1202
Score = 62.0 bits (149), Expect = 9e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +3
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 1209 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 1385
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 1386 NIQKESTLHLVLRLR 1430
Score = 62.0 bits (149), Expect = 9e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +3
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 297 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 473
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 474 NIQKESTLHLVLRLR 518
>TC76347 PIR|S17435|S17435 polyubiquitin 6 - common sunflower, partial (89%)
Length = 1421
Score = 63.9 bits (154), Expect = 2e-10
Identities = 32/77 (41%), Positives = 52/77 (66%)
Frame = +2
Query: 53 IMLQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLA 112
I +Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA
Sbjct: 188 IKMQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDRRTLA 364
Query: 113 ECSIGKDASLQLVGRMR 129
+ +I K+++L LV R+R
Sbjct: 365 DYNIQKESTLHLVLRLR 415
Score = 62.0 bits (149), Expect = 9e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +2
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 650 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 826
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 827 NIQKESTLHLVLRLR 871
Score = 62.0 bits (149), Expect = 9e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +2
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 878 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 1054
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 1055NIQKESTLHLVLRLR 1099
Score = 62.0 bits (149), Expect = 9e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +2
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 1106 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 1282
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 1283 NIQKESTLHLVLRLR 1327
Score = 62.0 bits (149), Expect = 9e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +2
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 422 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 598
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 599 NIQKESTLHLVLRLR 643
>AW586704 PIR|S17435|S17 polyubiquitin 6 - common sunflower, partial (19%)
Length = 561
Score = 63.9 bits (154), Expect = 2e-10
Identities = 32/77 (41%), Positives = 53/77 (68%)
Frame = +1
Query: 53 IMLQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLA 112
I++Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA
Sbjct: 295 IIMQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLA 471
Query: 113 ECSIGKDASLQLVGRMR 129
+ +I K+++L LV R+R
Sbjct: 472 DYNIQKESTLHLVLRLR 522
>AA660867 homologue to PIR|S17435|S17 polyubiquitin 6 - common sunflower,
partial (29%)
Length = 653
Score = 63.9 bits (154), Expect = 2e-10
Identities = 32/77 (41%), Positives = 52/77 (66%)
Frame = +3
Query: 53 IMLQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLA 112
I +Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA
Sbjct: 15 IKMQIFVKTL-TGKTITLEVESSDTIDNVXAKIQDKEGIPPDQQRLIFAGKQLEDRRTLA 191
Query: 113 ECSIGKDASLQLVGRMR 129
+ +I K+++L LV R+R
Sbjct: 192 DYNIQKESTLHLVLRLR 242
Score = 33.9 bits (76), Expect = 0.27
Identities = 17/49 (34%), Positives = 27/49 (54%)
Frame = +3
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGK 103
+Q FV+ + G TI ++ DT+ ++ I GIP +QRLI+ K
Sbjct: 249 MQIFVKTL-TGKTITLEVESSDTIDNVKAXIXDXXGIPPDQQRLIFAWK 392
>TC76332 GP|15450587|gb|AAK96565.1 AT4g05050/T32N4_13 {Arabidopsis
thaliana}, complete
Length = 963
Score = 63.5 bits (153), Expect = 3e-10
Identities = 34/80 (42%), Positives = 54/80 (67%)
Frame = +3
Query: 50 HRSIMLQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQ 109
HR I +Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +
Sbjct: 24 HR-IKMQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGR 197
Query: 110 TLAECSIGKDASLQLVGRMR 129
TLA+ +I K+++L LV R+R
Sbjct: 198 TLADYNIQKESTLHLVLRLR 257
Score = 62.0 bits (149), Expect = 9e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +3
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 492 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 668
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 669 NIQKESTLHLVLRLR 713
Score = 62.0 bits (149), Expect = 9e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +3
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 264 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 440
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 441 NIQKESTLHLVLRLR 485
>TC76388 homologue to PIR|S17435|S17435 polyubiquitin 6 - common sunflower,
partial (47%)
Length = 757
Score = 63.2 bits (152), Expect = 4e-10
Identities = 32/78 (41%), Positives = 52/78 (66%)
Frame = +1
Query: 52 SIMLQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTL 111
S +Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TL
Sbjct: 88 SFKMQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTL 264
Query: 112 AECSIGKDASLQLVGRMR 129
A+ +I K+++L LV R+R
Sbjct: 265 ADYNIQKESTLHLVLRLR 318
Score = 62.0 bits (149), Expect = 9e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +1
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 325 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 501
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 502 NIQKESTLHLVLRLR 546
Score = 51.6 bits (122), Expect = 1e-06
Identities = 25/68 (36%), Positives = 44/68 (63%)
Frame = +1
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +Q+LI+ GK L +TLA+
Sbjct: 553 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPNQQKLIFAGKXLXXGRTLADY 729
Query: 115 SIGKDASL 122
+I K+++L
Sbjct: 730 NIXKESTL 753
>TC85479 PIR|S20925|S20925 polyubiquitin - maize, complete
Length = 1983
Score = 63.2 bits (152), Expect = 4e-10
Identities = 32/78 (41%), Positives = 52/78 (66%)
Frame = +2
Query: 52 SIMLQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTL 111
S +Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TL
Sbjct: 68 SFKMQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTL 244
Query: 112 AECSIGKDASLQLVGRMR 129
A+ +I K+++L LV R+R
Sbjct: 245 ADYNIQKESTLHLVLRLR 298
Score = 62.0 bits (149), Expect = 9e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +2
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 533 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 709
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 710 NIQKESTLHLVLRLR 754
Score = 62.0 bits (149), Expect = 9e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +2
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 761 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 937
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 938 NIQKESTLHLVLRLR 982
Score = 62.0 bits (149), Expect = 9e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +2
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 989 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 1165
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 1166 NIQKESTLHLVLRLR 1210
Score = 62.0 bits (149), Expect = 9e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +2
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 1217 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 1393
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 1394 NIQKESTLHLVLRLR 1438
Score = 62.0 bits (149), Expect = 9e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +2
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 1445 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 1621
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 1622 NIQKESTLHLVLRLR 1666
Score = 62.0 bits (149), Expect = 9e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +2
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 305 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 481
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 482 NIQKESTLHLVLRLR 526
>TC76581 homologue to PIR|S40240|S40240 ubiquitin/ribosomal protein S27a
fusion protein - white lupine, complete
Length = 665
Score = 62.8 bits (151), Expect = 5e-10
Identities = 32/78 (41%), Positives = 52/78 (66%)
Frame = +3
Query: 52 SIMLQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTL 111
S +Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TL
Sbjct: 30 SAKMQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTL 206
Query: 112 AECSIGKDASLQLVGRMR 129
A+ +I K+++L LV R+R
Sbjct: 207 ADYNIQKESTLHLVLRLR 260
>BG452313 homologue to PIR|T40261|T40 ubi4 protein - fission yeast
(Schizosaccharomyces pombe), partial (51%)
Length = 658
Score = 62.4 bits (150), Expect = 7e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +3
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 294 MQIFVKTL-TGKTITLEVESSDTIDNVKSKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 470
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 471 NIQKESTLHLVLRLR 515
Score = 62.4 bits (150), Expect = 7e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +3
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 66 MQIFVKTL-TGKTITLEVESSDTIDNVKSKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 242
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 243 NIQKESTLHLVLRLR 287
Score = 35.4 bits (80), Expect = 0.092
Identities = 16/46 (34%), Positives = 28/46 (60%)
Frame = +3
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIY 100
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +Q LI+
Sbjct: 522 MQIFVKTL-TGKTITLEVESSDTIDNVKSKIQDKEGIPPDQQXLIF 656
>TC86564 PIR|C86439|C86439 protein T19E23.13 [imported] - Arabidopsis
thaliana, partial (97%)
Length = 839
Score = 62.0 bits (149), Expect = 9e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +1
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 115 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 291
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 292 NIQKESTLHLVLRLR 336
Score = 51.6 bits (122), Expect = 1e-06
Identities = 27/65 (41%), Positives = 38/65 (57%)
Frame = +1
Query: 65 GNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAECSIGKDASLQL 124
G I + P DT+ I ER++ +GIP +QRLIY GKQL ++T E +I + L L
Sbjct: 370 GKEIEIDIEPTDTIDRIKERVEEKEGIPPVQQRLIYAGKQLADDKTAKEYNIEGGSVLHL 549
Query: 125 VGRMR 129
V +R
Sbjct: 550 VLALR 564
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.137 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,808,616
Number of Sequences: 36976
Number of extensions: 413554
Number of successful extensions: 2122
Number of sequences better than 10.0: 91
Number of HSP's better than 10.0 without gapping: 2010
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2112
length of query: 846
length of database: 9,014,727
effective HSP length: 104
effective length of query: 742
effective length of database: 5,169,223
effective search space: 3835563466
effective search space used: 3835563466
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0288.8


