

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0284.4
(247 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
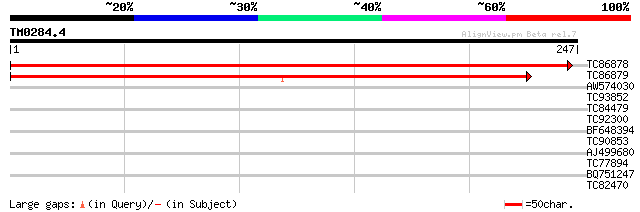
Score E
Sequences producing significant alignments: (bits) Value
TC86878 similar to SP|O80840|PMM_ARATH Probable phosphomannomuta... 461 e-131
TC86879 similar to SP|O80840|PMM_ARATH Probable phosphomannomuta... 413 e-116
AW574030 similar to GP|3851530|gb| nodulin {Glycine max}, partia... 33 0.10
TC93852 31 0.39
TC84479 ENBP1 31 0.39
TC92300 similar to GP|22136838|gb|AAM91763.1 unknown protein {Ar... 28 2.5
BF648394 28 3.3
TC90853 similar to GP|2342427|dbj|BAA21857.1 NPK1-related protei... 28 4.3
AJ499680 27 7.3
TC77894 similar to GP|12583793|gb|AAG59645.1 hypothetical protei... 27 7.3
BQ751247 similar to GP|21110151|gb| conserved hypothetical prote... 27 9.5
TC82470 homologue to GP|23497460|gb|AAN37003.1 hypothetical prot... 27 9.5
>TC86878 similar to SP|O80840|PMM_ARATH Probable phosphomannomutase (EC
5.4.2.8) (PMM). [Mouse-ear cress] {Arabidopsis
thaliana}, complete
Length = 1188
Score = 461 bits (1187), Expect = e-131
Identities = 226/245 (92%), Positives = 232/245 (94%)
Frame = +2
Query: 1 MAVAKPGVIALFDVDGTLTAPRKVANSEMLGFMQELRKVVTVGVVGGSDLIKISEQLGST 60
MA PGV+ALFDVDGTLTAPRK EML FMQ+LRK VTVGVVGGSDL+KISEQLG T
Sbjct: 284 MASLNPGVLALFDVDGTLTAPRKGVTPEMLKFMQDLRKFVTVGVVGGSDLVKISEQLGQT 463
Query: 61 VTHDYDYVFSENGLVAHKQGKLIGTESLKTFLGDEKLKEFINFTLHYIADLDIPIKRGTF 120
VT DYDYVFSENGLVAHKQGKLIGT+SLKTF+GDEKLKEFINFTLHYIADLDIPIKRGTF
Sbjct: 464 VTTDYDYVFSENGLVAHKQGKLIGTQSLKTFVGDEKLKEFINFTLHYIADLDIPIKRGTF 643
Query: 121 IEFRSGMLNVSPIGRNCSQEERDEFEKYDKVQNIRPKMVSVLREKFAHLNLTFSIGGQIS 180
IEFRSGMLNVSPIGRNCSQEERDEFEKYDKVQNIR KMVS+LREKFAHLNLTFSIGGQIS
Sbjct: 644 IEFRSGMLNVSPIGRNCSQEERDEFEKYDKVQNIRSKMVSILREKFAHLNLTFSIGGQIS 823
Query: 181 FDVFPQGWDKTYCLRYLDGFNEIHFFGDKTYKGGNDHEIYESERTIGHTVTSPEDTIKQC 240
FDVFPQGWDKTYCLRYLDGFNEIHFFGDKTYKGGNDHEIYESERTIGHTVTSPEDTIKQC
Sbjct: 824 FDVFPQGWDKTYCLRYLDGFNEIHFFGDKTYKGGNDHEIYESERTIGHTVTSPEDTIKQC 1003
Query: 241 KSLFL 245
SLFL
Sbjct: 1004TSLFL 1018
>TC86879 similar to SP|O80840|PMM_ARATH Probable phosphomannomutase (EC
5.4.2.8) (PMM). [Mouse-ear cress] {Arabidopsis
thaliana}, partial (92%)
Length = 760
Score = 413 bits (1062), Expect = e-116
Identities = 209/252 (82%), Positives = 215/252 (84%), Gaps = 25/252 (9%)
Frame = +1
Query: 1 MAVAKPGVIALFDVDGTLTAPRKVANSEMLGFMQELRKVVTVGVVGGSDLIKISEQLGST 60
MA PGV+ALFDVDGTLTAPRK EML FMQ+LRK VTVGVVGGSDL+KISEQLG T
Sbjct: 4 MASLNPGVLALFDVDGTLTAPRKGVTPEMLKFMQDLRKFVTVGVVGGSDLVKISEQLGQT 183
Query: 61 VTHDYDYVFSENGLVAHKQGKLIGTESLKTFLGDEKLKEFINFTLHYIADLDIPIKR--- 117
VT DYDYVFSENGLVAHKQGKLIGT+SLKTF+GDEKLKEFINFTLHYIADLDIPIKR
Sbjct: 184 VTTDYDYVFSENGLVAHKQGKLIGTQSLKTFVGDEKLKEFINFTLHYIADLDIPIKRLVR 363
Query: 118 ----------------------GTFIEFRSGMLNVSPIGRNCSQEERDEFEKYDKVQNIR 155
GTFIEFRSGMLNVSPIGRNCSQEERDEFEKYDKVQNIR
Sbjct: 364 FSG*SHCYIKLLNPNLCKKKSRGTFIEFRSGMLNVSPIGRNCSQEERDEFEKYDKVQNIR 543
Query: 156 PKMVSVLREKFAHLNLTFSIGGQISFDVFPQGWDKTYCLRYLDGFNEIHFFGDKTYKGGN 215
KMVS+LREKFAHLNLTFSIGGQISFDVFPQGWDKTYCLRYLDGFNEIHFFGDKTYKGGN
Sbjct: 544 SKMVSILREKFAHLNLTFSIGGQISFDVFPQGWDKTYCLRYLDGFNEIHFFGDKTYKGGN 723
Query: 216 DHEIYESERTIG 227
DHEIYESERTIG
Sbjct: 724 DHEIYESERTIG 759
>AW574030 similar to GP|3851530|gb| nodulin {Glycine max}, partial (39%)
Length = 707
Score = 33.1 bits (74), Expect = 0.10
Identities = 18/60 (30%), Positives = 29/60 (48%)
Frame = +3
Query: 15 DGTLTAPRKVANSEMLGFMQELRKVVTVGVVGGSDLIKISEQLGSTVTHDYDYVFSENGL 74
D T A ++ A +E+ +E +VTVG G + K+ LG+ T DY+ G+
Sbjct: 462 DATFYASKQAAEAELYAKKKEAEGIVTVGQAQGVYVSKLLNALGNDYTAVRDYLMINGGM 641
>TC93852
Length = 490
Score = 31.2 bits (69), Expect = 0.39
Identities = 22/82 (26%), Positives = 39/82 (46%)
Frame = -2
Query: 115 IKRGTFIEFRSGMLNVSPIGRNCSQEERDEFEKYDKVQNIRPKMVSVLREKFAHLNLTFS 174
I R F +F S + ++P C+ + ++ NI+ K+V++L L ++S
Sbjct: 468 ITRSQFFQFISSLHTITPFAEKCTHFSQ-------QIYNIQFKVVTMLE---F*LIFSYS 319
Query: 175 IGGQISFDVFPQGWDKTYCLRY 196
IG I + P G T C+R+
Sbjct: 318 IGNIIQNSLNPNGILNTDCVRF 253
>TC84479 ENBP1
Length = 5108
Score = 31.2 bits (69), Expect = 0.39
Identities = 14/47 (29%), Positives = 25/47 (52%)
Frame = +1
Query: 149 DKVQNIRPKMVSVLREKFAHLNLTFSIGGQISFDVFPQGWDKTYCLR 195
D+VQN++PK+ K N+ G ++ + +GW K +CL+
Sbjct: 1843 DEVQNVKPKLGRPKGSKNKKKNIAGEDGNKLHKEKKRRGWPKGFCLK 1983
>TC92300 similar to GP|22136838|gb|AAM91763.1 unknown protein {Arabidopsis
thaliana}, partial (7%)
Length = 989
Score = 28.5 bits (62), Expect = 2.5
Identities = 8/17 (47%), Positives = 13/17 (76%)
Frame = -2
Query: 177 GQISFDVFPQGWDKTYC 193
GQ+ F VFP+ W++ +C
Sbjct: 685 GQVPFTVFPKSWERVHC 635
>BF648394
Length = 650
Score = 28.1 bits (61), Expect = 3.3
Identities = 16/54 (29%), Positives = 30/54 (54%), Gaps = 1/54 (1%)
Frame = -3
Query: 151 VQNIRPKMVSVLREKFAHLNLTFSIGGQIS-FDVFPQGWDKTYCLRYLDGFNEI 203
+Q + VSVL +F ++ L IGG +S + + ++T+C + L +N+I
Sbjct: 540 IQTTFGQSVSVLEHRFKYVTLYSGIGGHVSRYALDNIALEETHCRKTLCMYNDI 379
>TC90853 similar to GP|2342427|dbj|BAA21857.1 NPK1-related protein kinase 3
{Arabidopsis thaliana}, partial (22%)
Length = 746
Score = 27.7 bits (60), Expect = 4.3
Identities = 22/70 (31%), Positives = 30/70 (42%), Gaps = 5/70 (7%)
Frame = +3
Query: 140 EERDEFEKYDKVQNIRPKMVSVLREKFAHLNLTFSIGGQIS-----FDVFPQGWDKTYCL 194
EE + K K NI + + E ++ L F GG IS F FP+ TY
Sbjct: 414 EEEVKLLKNLKHPNIVRYLGTAREEDSLNILLEFVPGGSISSLLGKFGSFPESGITTYTK 593
Query: 195 RYLDGFNEIH 204
+ LDG +H
Sbjct: 594 QLLDGLEYLH 623
>AJ499680
Length = 527
Score = 26.9 bits (58), Expect = 7.3
Identities = 14/47 (29%), Positives = 23/47 (48%)
Frame = -1
Query: 129 NVSPIGRNCSQEERDEFEKYDKVQNIRPKMVSVLREKFAHLNLTFSI 175
N+SP + + E Y K + ++ LREKF H+N+ S+
Sbjct: 158 NISPSFISKRTKHARERADYLKSTRLMKNIIRCLREKFRHVNIFISL 18
>TC77894 similar to GP|12583793|gb|AAG59645.1 hypothetical protein {Oryza
sativa}, partial (84%)
Length = 837
Score = 26.9 bits (58), Expect = 7.3
Identities = 13/44 (29%), Positives = 21/44 (47%)
Frame = -3
Query: 146 EKYDKVQNIRPKMVSVLREKFAHLNLTFSIGGQISFDVFPQGWD 189
++Y V+N ++V F+HL L S+ + D F WD
Sbjct: 172 DQYQPVRNCETEIVHFFSFSFSHLKLK*SLNW*ENLDEFEVNWD 41
>BQ751247 similar to GP|21110151|gb| conserved hypothetical protein
{Xanthomonas axonopodis pv. citri str. 306}, partial
(5%)
Length = 787
Score = 26.6 bits (57), Expect = 9.5
Identities = 9/28 (32%), Positives = 15/28 (53%)
Frame = -2
Query: 213 GGNDHEIYESERTIGHTVTSPEDTIKQC 240
G DH + + R + HT+ P+ K+C
Sbjct: 660 GSADHGLQNTSRHVSHTIKRPDSCGKRC 577
>TC82470 homologue to GP|23497460|gb|AAN37003.1 hypothetical protein
{Plasmodium falciparum 3D7}, partial (8%)
Length = 1064
Score = 26.6 bits (57), Expect = 9.5
Identities = 16/48 (33%), Positives = 25/48 (51%), Gaps = 4/48 (8%)
Frame = -2
Query: 91 FLGDEKLKEFINFTLHYIADLD----IPIKRGTFIEFRSGMLNVSPIG 134
FL + L F+NF+LH++A D IP + + +LN+S G
Sbjct: 343 FLCYKILDTFVNFSLHFLASFDNSSNIPNFLFLIFKIFAALLNISLYG 200
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.140 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,527,403
Number of Sequences: 36976
Number of extensions: 81371
Number of successful extensions: 333
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 331
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 332
length of query: 247
length of database: 9,014,727
effective HSP length: 94
effective length of query: 153
effective length of database: 5,538,983
effective search space: 847464399
effective search space used: 847464399
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0284.4


