

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0283.6
(561 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
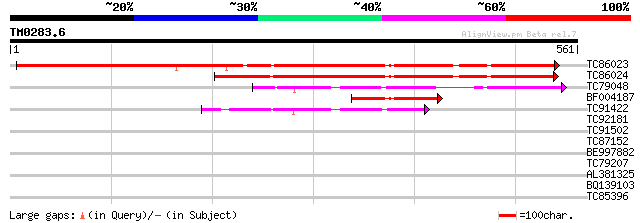
TC86023 weakly similar to GP|18057156|gb|AAL58179.1 putative CEO... 444 e-125
TC86024 similar to GP|16604703|gb|AAL24144.1 unknown protein {Ar... 293 9e-80
TC79048 weakly similar to GP|15450639|gb|AAK96591.1 AT5g62520/K1... 120 1e-27
BF004187 similar to PIR|H86446|H864 unknown protein [imported] -... 116 2e-26
TC91422 weakly similar to PIR|C86369|C86369 protein F5O8.11 [imp... 103 2e-22
TC92181 similar to GP|14517476|gb|AAK62628.1 At2g43320/T1O24.6 {... 33 0.29
TC91502 similar to GP|2564500|gb|AAB81756.1| olfactory receptor ... 32 0.50
TC87152 similar to GP|9280661|gb|AAF86530.1| F21B7.20 {Arabidops... 29 4.2
BE997882 similar to PIR|E84433|E84 hypothetical protein At2g0215... 28 7.2
TC79207 similar to GP|10086489|gb|AAG12549.1 Unknown Protein {Ar... 28 7.2
AL381325 similar to GP|4156245|dbj| ATHP3 {Arabidopsis thaliana}... 28 7.2
BQ139103 homologue to GP|2655420|gb| heat shock cognate protein ... 28 9.3
TC85396 homologue to GP|6911551|emb|CAB72129.1 heat shock protei... 28 9.3
>TC86023 weakly similar to GP|18057156|gb|AAL58179.1 putative CEO protein
{Oryza sativa}, partial (16%)
Length = 2368
Score = 444 bits (1142), Expect = e-125
Identities = 255/559 (45%), Positives = 349/559 (61%), Gaps = 21/559 (3%)
Frame = +3
Query: 7 LKRNETTRCGVHIRGASQQELCQQHFCASPTINFVNRMRLGDYKRKMNILGSGSGQSYLR 66
LKR +R GA + L Q SP V +M+L YK K + +S++R
Sbjct: 444 LKRKHASRHATCPNGALRPVLPQWASLVSPRNKVVKQMKLNSYKSKPTKSETHIRRSWVR 623
Query: 67 FYLNYKKSARPERMMFYNHGEWVDYPKDIVDLVKKDFEIKNAAIGIRLNGRDLMLDFLHA 126
+Y+NYKKS RPER+M Y +GEW D+P+++VDLV++DF K ++ + L+G L+L+FLH
Sbjct: 624 YYMNYKKSGRPERLMVYQNGEWKDFPRNVVDLVREDFNAKKGSVEVELHGHHLVLNFLHM 803
Query: 127 CQMDLKTGFIQPIAWIDKVGGYFFPEVDVASGEESYN-------------FGEQEGMKFH 173
QM+LK+G +PIAWID+ G +FPE+ S E Y+ F + +K H
Sbjct: 804 YQMNLKSGLQRPIAWIDETGCCYFPEIYDDSDEGPYDPCNQDSGESQESLFQDPNDIKLH 983
Query: 174 VEIKMNMVDEFQLRECIGESNALVKDAQVKVR-NMMSIMDC------GYIGEEIAKNEEL 226
+EI++N VD +L E GESN +VK Q+ + N + I D G +GE + +N+ +
Sbjct: 984 LEIEINGVDASKLGESTGESNVIVKHTQIDAKQNDLEIEDSSNNMGNGNVGEAVEQNKNI 1163
Query: 227 GLSAYTGYVHGNLVLDSIRDLFCNGMSIIGNNDFEIVEIYRCSCASMQVQSKLFLTQAEI 286
SA V+ L LD+++ +F G S G+ D IVEIY S M + +LF QAEI
Sbjct: 1164VGSAV---VYVRLDLDTVQKMFLKGTSSFGSAD--IVEIYPFSSTMMLSRLELFEKQAEI 1328
Query: 287 TKELHGDANVRYAWLSFSKGELSTMMEHGLGHCELFTTKCTYGVGVHLAAISCPYASARY 346
TK GDAN++YAWL+ SKGELSTMM+ GLGHC + T+KCT+G GVHLAA + P+ASA +
Sbjct: 1329TKNCRGDANIQYAWLASSKGELSTMMKCGLGHCGISTSKCTHGFGVHLAAATFPFASASH 1508
Query: 347 CDIDGNGVRHLILCRIIMGNMELLCPSIGTSNVQFQPSSSEYDNGVDDIQCPRYYTIWNM 406
CDID NGVRHL+ CR+IMGNMELL P GT QF+PSSS+YD+GVDDI PRYY +WNM
Sbjct: 1509CDIDENGVRHLVFCRVIMGNMELLRP--GTR--QFRPSSSDYDSGVDDIHNPRYYIVWNM 1676
Query: 407 NISTYICPAFVVSFKVSRGVEGYFCGTVDKNNVSRDNSSGANSDSHCSGDPVQSASSAYN 466
N +T+ICP +VVSFKVS EG G+ KNN+ G NS S +S SS +
Sbjct: 1677NTNTHICPEYVVSFKVSSATEGLLLGSKTKNNI-----FGVNSASRSPKILSRSESSTVD 1841
Query: 467 EIAGNGVACTQKIPKSPWMPLHMLIAAINDKVPPNDMSLIKENFELYKAKQISRDDFVKE 526
+G +++ P SP MP +LI A+ +KV DM +I ++ ++AK+++RDDF K+
Sbjct: 1842TGIASG---SRRGPTSPSMPFPVLIDALKNKVSAKDMEVINMHYLQFRAKRMTRDDFAKK 2012
Query: 527 MRLIV-GDTLLRDTITTLQ 544
+RLIV D LLR TI LQ
Sbjct: 2013LRLIVRDDALLRATIVGLQ 2069
>TC86024 similar to GP|16604703|gb|AAL24144.1 unknown protein {Arabidopsis
thaliana}, partial (23%)
Length = 1431
Score = 293 bits (751), Expect = 9e-80
Identities = 164/341 (48%), Positives = 216/341 (63%)
Frame = +1
Query: 203 KVRNMMSIMDCGYIGEEIAKNEELGLSAYTGYVHGNLVLDSIRDLFCNGMSIIGNNDFEI 262
K+ + + M G +GE + +N+ +G +AY V+G L LD+++ +F G G+ I
Sbjct: 82 KIEDSSNNMGNGSVGEAVEQNKSIGFNAYNEIVYGKLDLDTVQKMFLKGTGSFGSA--YI 255
Query: 263 VEIYRCSCASMQVQSKLFLTQAEITKELHGDANVRYAWLSFSKGELSTMMEHGLGHCELF 322
VEIY S +Q + +LF QAEI K GDANVRYAWL+ SKGEL TMM+ GLGHC
Sbjct: 256 VEIYPFSSTLVQSRLELFEKQAEIAKNCRGDANVRYAWLASSKGELPTMMKCGLGHCVPS 435
Query: 323 TTKCTYGVGVHLAAISCPYASARYCDIDGNGVRHLILCRIIMGNMELLCPSIGTSNVQFQ 382
KC +G+GVHLAA +CP+ASA YCD+D NGVRH++ CR+IMGNMELL P GT Q++
Sbjct: 436 APKCMHGIGVHLAAATCPFASASYCDVDENGVRHIVFCRVIMGNMELLHP--GTR--QYR 603
Query: 383 PSSSEYDNGVDDIQCPRYYTIWNMNISTYICPAFVVSFKVSRGVEGYFCGTVDKNNVSRD 442
PSSS+YD+GVDDI PR + +WNMN++T+I P FVVSFKVS EG G KNN+
Sbjct: 604 PSSSDYDSGVDDIHNPRNFIVWNMNMNTHIYPEFVVSFKVSSDAEGLLLGCERKNNI--- 774
Query: 443 NSSGANSDSHCSGDPVQSASSAYNEIAGNGVACTQKIPKSPWMPLHMLIAAINDKVPPND 502
G NS S+ +S SS + G + + SP P L AAI +KVP D
Sbjct: 775 --FGVNSASYGPKILPRSESSRVSTGIPTG---SPEASTSPRFPFSTLFAAIINKVPAED 939
Query: 503 MSLIKENFELYKAKQISRDDFVKEMRLIVGDTLLRDTITTL 543
M LI +KAK+++RDDF+K +R +VGD LLR T+ L
Sbjct: 940 MKLIHMYHLHFKAKKMTRDDFLKNLRSVVGDDLLRATLPGL 1062
>TC79048 weakly similar to GP|15450639|gb|AAK96591.1 AT5g62520/K19B1_13
{Arabidopsis thaliana}, partial (19%)
Length = 1709
Score = 120 bits (302), Expect = 1e-27
Identities = 82/316 (25%), Positives = 154/316 (47%), Gaps = 5/316 (1%)
Frame = +3
Query: 241 LDSIRDLFCNGMSIIGNNDFEIVEIYRCSCASMQVQSKLF---LTQAEITKELHGDANVR 297
++ IR F G+S G + + I+R C ++ Q+++ + + K G+ANV+
Sbjct: 570 INLIRSRFVCGLSAQGLKPYGVA-IFRNPCKTVMAQARIQSFEIFARAVAKMRGGNANVK 746
Query: 298 YAWL-SFSKGELSTMMEHGLGHCELFTTKCTYGVGVHLAAISCPYASARYCDIDGNGVRH 356
+ W + S+ E+ +++HG G+ + G+ L+ P S + +D G RH
Sbjct: 747 HVWYGASSREEIVDIIQHGFGY--------VHSNGLRLSPNDTPLESVKRTVVDKQGFRH 902
Query: 357 LILCRIIMGNMELLCPSIGTSNVQFQPSSSEYDNGVDDIQCPRYYTIWNMNISTYICPAF 416
L+LCR+IMG +E ++ + Q +PSS E+D+GVD P+ + +W+ I+T++ P +
Sbjct: 903 LLLCRVIMGKLE----AVPAGSDQRRPSSEEFDSGVDSFSSPKEFIVWSNKINTHVLPEY 1070
Query: 417 VVSFKVSRGVEGYFCGTVDKNNVSRDNSSGANSDSHCSGDPVQSASSAYNEIAGNGVACT 476
V+SFK+ ++ + ++ GV
Sbjct: 1071VLSFKL-------------------------------------ASDKGHEKV---GVGGQ 1130
Query: 477 QKIPKSPWMPLHMLIAAINDKVPPNDMSLIKENFELYKAKQISRDDFVKEMRLIVGDTLL 536
P SP+M LI+A++ +PP+D+ I + + Y+ K+ISR + ++++R + GD LL
Sbjct: 1131HMRPSSPFMQFPTLISALSKILPPSDIVSIAKFHKEYRDKKISRHEMIQKVRKVAGDKLL 1310
Query: 537 RDTITTLQF-MVHNIF 551
I + + +H IF
Sbjct: 1311FSVIKSFRAKKMHAIF 1358
>BF004187 similar to PIR|H86446|H864 unknown protein [imported] - Arabidopsis
thaliana, partial (10%)
Length = 387
Score = 116 bits (291), Expect = 2e-26
Identities = 56/90 (62%), Positives = 69/90 (76%)
Frame = +3
Query: 339 CPYASARYCDIDGNGVRHLILCRIIMGNMELLCPSIGTSNVQFQPSSSEYDNGVDDIQCP 398
C SA YCD+D NGVRH++ CR+IMGNMELL P GT Q++PSSS+YD+GVDDI P
Sbjct: 78 CRIYSASYCDVDENGVRHIVFCRVIMGNMELLHP--GTR--QYRPSSSDYDSGVDDIHNP 245
Query: 399 RYYTIWNMNISTYICPAFVVSFKVSRGVEG 428
R + +WNMN++T+I P FVVSFKVS EG
Sbjct: 246 RNFIVWNMNMNTHIYPEFVVSFKVSSDAEG 335
>TC91422 weakly similar to PIR|C86369|C86369 protein F5O8.11 [imported] -
Arabidopsis thaliana, partial (16%)
Length = 754
Score = 103 bits (256), Expect = 2e-22
Identities = 69/230 (30%), Positives = 114/230 (49%), Gaps = 4/230 (1%)
Frame = +2
Query: 190 IGESNALVKDAQVKVRNMMSIMDCGYIGEEIAKNEELGLSAYTGYVHGNLVLDSIRDLFC 249
IGE ++ V D + V + GEE + + S G+ V D ++ F
Sbjct: 125 IGEDSSTVSDCESGVSATTT-------GEEQRQRRQNSDSFLVWLGEGDSVYDLLKTRFL 283
Query: 250 NGMSIIGNNDFEIVEIYRCSCASMQVQSKL--FLTQAEITKELHG-DANVRYAWLSFS-K 305
+G+ + + EI+ I R +C+ Q++L FL AE +L G D+NV+YAW S +
Sbjct: 284 HGLGALSSKA-EILAIQRNACSDAVSQARLRSFLVYAEAVSKLRGGDSNVKYAWYGSSGE 460
Query: 306 GELSTMMEHGLGHCELFTTKCTYGVGVHLAAISCPYASARYCDIDGNGVRHLILCRIIMG 365
++ ++ +G H +G + L+ P S + C + +GVRHLILCR+I+G
Sbjct: 461 NDVRGILSNGFSH--------VHGNSICLSPDDSPLQSVKSCAVGRDGVRHLILCRVILG 616
Query: 366 NMELLCPSIGTSNVQFQPSSSEYDNGVDDIQCPRYYTIWNMNISTYICPA 415
E+ + Q PS ++YD+GVD P Y IW+ ++T++ PA
Sbjct: 617 RTEI----VQADTKQCYPSCADYDSGVDSFSAPTKYMIWSSRMNTHVWPA 754
>TC92181 similar to GP|14517476|gb|AAK62628.1 At2g43320/T1O24.6 {Arabidopsis
thaliana}, partial (50%)
Length = 1114
Score = 33.1 bits (74), Expect = 0.29
Identities = 23/76 (30%), Positives = 39/76 (51%), Gaps = 2/76 (2%)
Frame = -1
Query: 270 CASMQVQSK--LFLTQAEITKELHGDANVRYAWLSFSKGELSTMMEHGLGHCELFTTKCT 327
C S+ +SK L+L+ IT L ++RY+ S+S L M +G HC ++TK
Sbjct: 328 CKSLP*RSKGRLWLSTLPITSLLEITKDIRYSCASYSFSILILKMHNGSSHC-FYSTKRL 152
Query: 328 YGVGVHLAAISCPYAS 343
+ + A + C +A+
Sbjct: 151 *KISLLAAELVCQFAN 104
>TC91502 similar to GP|2564500|gb|AAB81756.1| olfactory receptor protein
{Necturus maculosus}, partial (9%)
Length = 589
Score = 32.3 bits (72), Expect = 0.50
Identities = 25/74 (33%), Positives = 32/74 (42%), Gaps = 1/74 (1%)
Frame = -2
Query: 7 LKRNETTRCGVHIRGASQQELCQQHFCASPTINFVNRMRLGDYKRKMNILGSGSGQ-SYL 65
LKR +R GA + L Q +P V MRL YK K + G +L
Sbjct: 225 LKRKRASRHATCSNGALRPVLPQCTSLVTPRNKVVKHMRLKGYKSKPMKSENHIGTIRWL 46
Query: 66 RFYLNYKKSARPER 79
+NYKKS P+R
Sbjct: 45 GIIMNYKKSGSPKR 4
>TC87152 similar to GP|9280661|gb|AAF86530.1| F21B7.20 {Arabidopsis
thaliana}, partial (69%)
Length = 1464
Score = 29.3 bits (64), Expect = 4.2
Identities = 14/43 (32%), Positives = 25/43 (57%)
Frame = -2
Query: 152 EVDVASGEESYNFGEQEGMKFHVEIKMNMVDEFQLRECIGESN 194
E ++S E+S++ Q G+ F ++ ++ + F REC ESN
Sbjct: 773 EFSLSSAEQSFSDSPQSGVMFRLKQVLSDLFCFDSRECQNESN 645
>BE997882 similar to PIR|E84433|E84 hypothetical protein At2g02150 [imported]
- Arabidopsis thaliana, partial (3%)
Length = 625
Score = 28.5 bits (62), Expect = 7.2
Identities = 14/37 (37%), Positives = 22/37 (58%)
Frame = +2
Query: 49 YKRKMNILGSGSGQSYLRFYLNYKKSARPERMMFYNH 85
+++K GSG+ QS L +L+ K S RP+ + NH
Sbjct: 173 FEKKNEH*GSGTLQSQLSHFLHRKPSPRPQLRRYPNH 283
>TC79207 similar to GP|10086489|gb|AAG12549.1 Unknown Protein {Arabidopsis
thaliana}, partial (20%)
Length = 1719
Score = 28.5 bits (62), Expect = 7.2
Identities = 26/100 (26%), Positives = 46/100 (46%), Gaps = 7/100 (7%)
Frame = +3
Query: 436 KNNVSRDNSSGANSDSHCS------GDPVQSASSAYNEIAGNGV-ACTQKIPKSPWMPLH 488
+N + +DNSSG + CS DP N + G G A +QK+ + L
Sbjct: 495 ENKLVQDNSSGERYQNGCSRVLNPCSDP--EVPDDGNLVLGYGANAASQKMSSEEFEELK 668
Query: 489 MLIAAINDKVPPNDMSLIKENFELYKAKQISRDDFVKEMR 528
+ +KV D+S+ EN E+ K++ + + + E++
Sbjct: 669 L----EKEKV-TTDLSICTENLEVAKSQLLETEQLLAEVK 773
>AL381325 similar to GP|4156245|dbj| ATHP3 {Arabidopsis thaliana}, partial
(48%)
Length = 520
Score = 28.5 bits (62), Expect = 7.2
Identities = 20/75 (26%), Positives = 35/75 (46%), Gaps = 1/75 (1%)
Frame = +2
Query: 259 DFEIVEIYRCSCASMQVQSKLFLTQAEIT-KELHGDANVRYAWLSFSKGELSTMMEHGLG 317
+F ++ I + + +V S+L T A +T +H Y W SFSK + +T M L
Sbjct: 71 NFAVLSISSFTETTCEVGSRLRQTAARVTTSHIHVHERCFYGWYSFSKLQHATSMRTILE 250
Query: 318 HCELFTTKCTYGVGV 332
F +C + + +
Sbjct: 251 ----FVFECCFSISL 283
>BQ139103 homologue to GP|2655420|gb| heat shock cognate protein HSC70
{Brassica napus}, partial (29%)
Length = 628
Score = 28.1 bits (61), Expect = 9.3
Identities = 13/28 (46%), Positives = 16/28 (56%)
Frame = -1
Query: 219 EIAKNEELGLSAYTGYVHGNLVLDSIRD 246
E N+ LG+ VHGNL+L SI D
Sbjct: 298 ESTTNQPLGIKNGVDRVHGNLILSSITD 215
>TC85396 homologue to GP|6911551|emb|CAB72129.1 heat shock protein 70
{Cucumis sativus}, complete
Length = 2320
Score = 28.1 bits (61), Expect = 9.3
Identities = 11/24 (45%), Positives = 16/24 (65%)
Frame = -1
Query: 223 NEELGLSAYTGYVHGNLVLDSIRD 246
N+ LG+ G++H NL+L SI D
Sbjct: 337 NQPLGIKNGVGWIHRNLILSSITD 266
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.138 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,074,705
Number of Sequences: 36976
Number of extensions: 256801
Number of successful extensions: 1601
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 1571
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1585
length of query: 561
length of database: 9,014,727
effective HSP length: 101
effective length of query: 460
effective length of database: 5,280,151
effective search space: 2428869460
effective search space used: 2428869460
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0283.6


