

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0283.2
(142 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
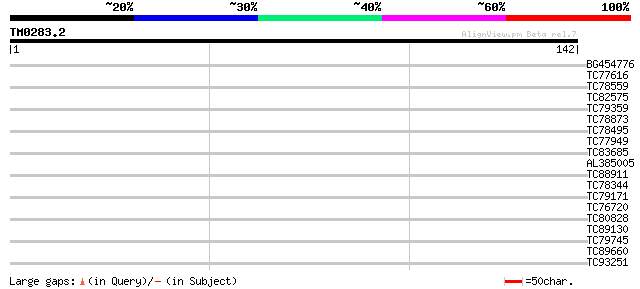
Score E
Sequences producing significant alignments: (bits) Value
BG454776 weakly similar to PIR|T49896|T4 glycine/proline-rich pr... 38 0.001
TC77616 similar to GP|11994422|dbj|BAB02424. oxidoreductase sho... 34 0.022
TC78559 weakly similar to GP|5733686|gb|AAD49719.1| maturation p... 32 0.065
TC82575 weakly similar to GP|6665807|gb|AAF23018.1| putative ABA... 32 0.065
TC79359 weakly similar to PIR|S49915|S49915 extensin-like protei... 31 0.19
TC78873 similar to GP|21740530|emb|CAD41509. OSJNBa0029H02.25 {O... 30 0.25
TC78495 homologue to PIR|S53504|S53504 extensin-like protein S3 ... 30 0.32
TC77949 weakly similar to GP|11762218|gb|AAG40387.1 AT4g27520 {A... 30 0.42
TC83685 29 0.55
AL385005 weakly similar to GP|21438547|emb Ratten cDNA {Rattus n... 28 0.94
TC88911 weakly similar to GP|13926280|gb|AAK49610.1 AT5g14920/F2... 28 0.94
TC78344 similar to GP|13540405|gb|AAK29456.1 histone H1 {Lens cu... 28 0.94
TC79171 similar to GP|19310435|gb|AAL84954.1 F17H15.1/F17H15.1 {... 28 1.2
TC76720 similar to GP|11121502|emb|CAC14888. putative extensin {... 28 1.6
TC80828 28 1.6
TC89130 similar to GP|12003396|gb|AAG43555.1 Avr9/Cf-9 rapidly e... 27 3.6
TC79745 similar to GP|6714475|gb|AAF26161.1| unknown protein {Ar... 27 3.6
TC89660 similar to GP|11034579|dbj|BAB17103. hypothetical protei... 27 3.6
TC93251 similar to PIR|T39903|T39903 serine-rich protein - fissi... 27 3.6
>BG454776 weakly similar to PIR|T49896|T4 glycine/proline-rich protein -
Arabidopsis thaliana, partial (7%)
Length = 680
Score = 38.1 bits (87), Expect = 0.001
Identities = 27/71 (38%), Positives = 38/71 (53%), Gaps = 3/71 (4%)
Frame = +3
Query: 70 LQTGFGTGLGAPVEAPAPGPFASAPQPSSEAAQQAFSYGKDKAYD-IKDKA-EVAYEDAK 127
L + + APV APA P ASAP+ + + QA+ D+AYD D+A + AY
Sbjct: 180 LNSAVSDSVKAPVPAPAKAP-ASAPKAAVQIKDQAYDQAYDQAYDQAYDQAYDQAYSQTN 356
Query: 128 DKA-ESAYQSA 137
D+A + AY A
Sbjct: 357 DQAYDQAYDQA 389
>TC77616 similar to GP|11994422|dbj|BAB02424. oxidoreductase short-chain
dehydrogenase/reductase family-like protein {Arabidopsis
thaliana}, partial (98%)
Length = 1912
Score = 33.9 bits (76), Expect = 0.022
Identities = 20/59 (33%), Positives = 29/59 (48%), Gaps = 2/59 (3%)
Frame = +3
Query: 66 TTENLQTGFGTGLGAPVEAPAPGPFASAPQPSSEAAQQAFSYGKDK--AYDIKDKAEVA 122
TT NL +GT V APGP P S A ++ S G+D+ Y + +K ++A
Sbjct: 642 TTRNLALEWGTDYDIRVNGIAPGPIGETPGMSKLAPEEIGSRGRDEMPLYKLGEKWDIA 818
>TC78559 weakly similar to GP|5733686|gb|AAD49719.1| maturation protein
pPM32 {Glycine max}, partial (55%)
Length = 1128
Score = 32.3 bits (72), Expect = 0.065
Identities = 18/49 (36%), Positives = 30/49 (60%), Gaps = 1/49 (2%)
Frame = +2
Query: 93 APQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKA-ESAYQSAKKT 140
A + + +AA+ A +D AYD KD+ + A E A +KA + AY + ++T
Sbjct: 449 AKERTKDAAENAGESIRDYAYDAKDRTKEAAESAGEKARDYAYDAKERT 595
Score = 30.4 bits (67), Expect = 0.25
Identities = 32/131 (24%), Positives = 51/131 (38%), Gaps = 23/131 (17%)
Frame = +2
Query: 32 SQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTG------------------ 73
S +S PS +WAYD H + D + R E + G
Sbjct: 26 SAESTRKPS-TNWAYD---HSSSKTKRDAEIADRARETMSEGVDSEDIKQYSRDANEDIK 193
Query: 74 -FGTGLGAPVEAPAPGPFASAPQPSSEAAQQAFSYG---KDKAYDIKDKAEVAYEDAKDK 129
F + A + A + + AA++A + G +D AY++KD+ A DK
Sbjct: 194 QFARDTNEKTKDAAGSMWEKAKEGTDRAAEKAENAGEKARDYAYEVKDRTNEAAGSVWDK 373
Query: 130 A-ESAYQSAKK 139
A E ++A+K
Sbjct: 374 AREGTNRAAEK 406
Score = 28.5 bits (62), Expect = 0.94
Identities = 17/49 (34%), Positives = 26/49 (52%), Gaps = 1/49 (2%)
Frame = +2
Query: 93 APQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKA-ESAYQSAKKT 140
A + EAA+ A +D AYD K++ + DK E A ++A+KT
Sbjct: 515 AKDRTKEAAESAGEKARDYAYDAKERTKEGASYMADKTKEGAEKTAEKT 661
>TC82575 weakly similar to GP|6665807|gb|AAF23018.1| putative ABA-induced
guard cell protein {Vicia faba}, partial (37%)
Length = 695
Score = 32.3 bits (72), Expect = 0.065
Identities = 17/41 (41%), Positives = 22/41 (53%)
Frame = +2
Query: 99 EAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAKK 139
EA A KD AYD DK + A A DKA+ +++A K
Sbjct: 398 EAVGSAGDKVKDYAYDANDKTKEAIGSATDKAKEGFEAATK 520
Score = 25.4 bits (54), Expect = 8.0
Identities = 18/47 (38%), Positives = 24/47 (50%), Gaps = 1/47 (2%)
Frame = +2
Query: 95 QPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAES-AYQSAKKT 140
Q ++E + A S DKA + DKA A A DK + AY + KT
Sbjct: 320 QDANEKTKDAASSIADKAKEDTDKAVEAVGSAGDKVKDYAYDANDKT 460
>TC79359 weakly similar to PIR|S49915|S49915 extensin-like protein - maize,
partial (3%)
Length = 1170
Score = 30.8 bits (68), Expect = 0.19
Identities = 25/79 (31%), Positives = 36/79 (44%), Gaps = 3/79 (3%)
Frame = +3
Query: 25 PATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLGAPVEA 84
PATSP + PPS S A ++V + P + + T + + +PVEA
Sbjct: 387 PATSPPAKSPAVQPPSSVSPAISPSNNV-----SSTPPVSSPASSPPTAAVSPVSSPVEA 551
Query: 85 P---APGPFASAPQPSSEA 100
P +P +SA PSS A
Sbjct: 552 PSVSSPPEASSAGIPSSSA 608
>TC78873 similar to GP|21740530|emb|CAD41509. OSJNBa0029H02.25 {Oryza
sativa}, partial (7%)
Length = 623
Score = 30.4 bits (67), Expect = 0.25
Identities = 29/110 (26%), Positives = 48/110 (43%), Gaps = 4/110 (3%)
Frame = +3
Query: 1 MGNAKIVLVVLCLAVCVGICRPDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDE 60
M + K+VL+++ A+ V SP++SPS S P+ + A+ + P +N
Sbjct: 114 MASYKVVLMLVA-ALLVSSTFAQSPSSSPSKS------PAISPSAHSPAASPPAPVKNSP 272
Query: 61 PVLPRTTENLQTGFGTGLGAPVEAPAPGPFA----SAPQPSSEAAQQAFS 106
P + + G+P APA P + A PS+ AA F+
Sbjct: 273 SPSPSAINSPPSPPPASSGSPAAAPAVTPSSISTPPAEAPSNGAALNRFT 422
>TC78495 homologue to PIR|S53504|S53504 extensin-like protein S3 - alfalfa,
partial (97%)
Length = 1049
Score = 30.0 bits (66), Expect = 0.32
Identities = 28/103 (27%), Positives = 45/103 (43%), Gaps = 3/103 (2%)
Frame = +3
Query: 1 MGNAKIVLVVLCLAVCV---GICRPDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYR 57
MG IVLVV+ L +CV + +P+TSP+ S +PP+ + S P +
Sbjct: 165 MGTHHIVLVVIGL-ICVVFSSVGAQQAPSTSPNSSPAPPTPPANTPPTTPQASP--PPVQ 335
Query: 58 NDEPVLPRTTENLQTGFGTGLGAPVEAPAPGPFASAPQPSSEA 100
+ P + + LQ+ P P P S+P P ++
Sbjct: 336 SSPPPVQSSPPPLQS------SPPPAQSTPPPVQSSPPPVQQS 446
>TC77949 weakly similar to GP|11762218|gb|AAG40387.1 AT4g27520 {Arabidopsis
thaliana}, partial (34%)
Length = 1297
Score = 29.6 bits (65), Expect = 0.42
Identities = 24/80 (30%), Positives = 36/80 (45%), Gaps = 2/80 (2%)
Frame = +3
Query: 19 ICRPDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGL 78
+ P +P +SPSPS SPPS + G + + P ++ +G
Sbjct: 480 VISPRTPKSSPSPSAGGLSPPSPSPTTTTPSPS--GSPPSPVAIPPASSPVPTSGPTASS 653
Query: 79 GAPVEA--PAPGPFASAPQP 96
+PV + PA GP AS+P P
Sbjct: 654 PSPVVSTPPAGGPMASSPSP 713
>TC83685
Length = 424
Score = 29.3 bits (64), Expect = 0.55
Identities = 14/37 (37%), Positives = 20/37 (53%)
Frame = +3
Query: 19 ICRPDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGP 55
+ +SP TS +P+ SDSPP+ + D V GP
Sbjct: 183 VAEAESPKTSEAPTTFSDSPPAQSPVVVDVVPPVSGP 293
>AL385005 weakly similar to GP|21438547|emb Ratten cDNA {Rattus norvegicus},
partial (5%)
Length = 469
Score = 28.5 bits (62), Expect = 0.94
Identities = 14/39 (35%), Positives = 22/39 (55%)
Frame = +2
Query: 68 ENLQTGFGTGLGAPVEAPAPGPFASAPQPSSEAAQQAFS 106
+ L G+G +P A +P P+AS+P PS+ A + S
Sbjct: 23 QKLSINVGSGSNSP--ATSPSPYASSPSPSTGATPPSAS 133
Score = 26.2 bits (56), Expect = 4.7
Identities = 12/21 (57%), Positives = 15/21 (71%)
Frame = +2
Query: 23 DSPATSPSPSQKSDSPPSFAS 43
+SPATSPSP S SP + A+
Sbjct: 56 NSPATSPSPYASSPSPSTGAT 118
>TC88911 weakly similar to GP|13926280|gb|AAK49610.1 AT5g14920/F2G14_40
{Arabidopsis thaliana}, partial (11%)
Length = 1320
Score = 28.5 bits (62), Expect = 0.94
Identities = 22/92 (23%), Positives = 38/92 (40%), Gaps = 12/92 (13%)
Frame = +3
Query: 23 DSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDE--PVLPRTTENLQTGFGTGLGA 80
D+P+++P+PS + D+P + + H ++ + P T + T A
Sbjct: 783 DTPSSAPAPSPEDDAPEPPPPHKHKRRKHKHSKHKKHHALALAPEPTSSSSTIIRRSPPA 962
Query: 81 PV----------EAPAPGPFASAPQPSSEAAQ 102
P+ E P+P AP PS+ AQ
Sbjct: 963 PLADDNTTMSSDEGPSP-----APSPSANGAQ 1043
Score = 26.6 bits (57), Expect = 3.6
Identities = 23/75 (30%), Positives = 28/75 (36%)
Frame = +3
Query: 22 PDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLGAP 81
PD +P+P+ K P K S V P P P T + T P
Sbjct: 666 PDKETPAPAPTHKKKKAP--------KSSPVPSPAIL--PPSPAPTPAIDT--------P 791
Query: 82 VEAPAPGPFASAPQP 96
APAP P AP+P
Sbjct: 792 SSAPAPSPEDDAPEP 836
>TC78344 similar to GP|13540405|gb|AAK29456.1 histone H1 {Lens culinaris},
partial (86%)
Length = 1215
Score = 28.5 bits (62), Expect = 0.94
Identities = 23/61 (37%), Positives = 28/61 (45%)
Frame = +2
Query: 78 LGAPVEAPAPGPFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSA 137
L A APA P A+ P+P +A K A K KA A AK KA++A A
Sbjct: 461 LPAKSAAPAKKPAAAKPKPKPKAKAPV---AKAPAAKSKAKAAPAKAKAKAKAKAAPAKA 631
Query: 138 K 138
K
Sbjct: 632 K 634
>TC79171 similar to GP|19310435|gb|AAL84954.1 F17H15.1/F17H15.1 {Arabidopsis
thaliana}, partial (27%)
Length = 1421
Score = 28.1 bits (61), Expect = 1.2
Identities = 16/40 (40%), Positives = 23/40 (57%), Gaps = 1/40 (2%)
Frame = +2
Query: 71 QTGFGTGLGAPVEAPAPGPFASAP-QPSSEAAQQAFSYGK 109
Q+G+GTG GAP P+ P A+ P S++ A YG+
Sbjct: 875 QSGYGTGYGAP---PSQKPSANPPVYGQSQSPSTAAGYGQ 985
>TC76720 similar to GP|11121502|emb|CAC14888. putative extensin {Nicotiana
sylvestris}, partial (51%)
Length = 1290
Score = 27.7 bits (60), Expect = 1.6
Identities = 10/19 (52%), Positives = 14/19 (73%)
Frame = +1
Query: 22 PDSPATSPSPSQKSDSPPS 40
P +P +P+PS SDSPP+
Sbjct: 643 PPAPTPTPTPSPHSDSPPA 699
>TC80828
Length = 721
Score = 27.7 bits (60), Expect = 1.6
Identities = 27/86 (31%), Positives = 34/86 (39%)
Frame = -2
Query: 17 VGICRPDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGT 76
V I P SP+P+ K SPPS HVF P P T +
Sbjct: 432 VEIASPGQEKQSPAPT-KPPSPPSNC--------HVF-------PTTPEDTTGTPSII-P 304
Query: 77 GLGAPVEAPAPGPFASAPQPSSEAAQ 102
G PV P P P +S+P P+ A+
Sbjct: 303 GKAQPV-LPIPNPISSSPSPNPPNAR 229
>TC89130 similar to GP|12003396|gb|AAG43555.1 Avr9/Cf-9 rapidly elicited
protein 137 {Nicotiana tabacum}, partial (5%)
Length = 1159
Score = 26.6 bits (57), Expect = 3.6
Identities = 14/33 (42%), Positives = 19/33 (57%)
Frame = +2
Query: 20 CRPDSPATSPSPSQKSDSPPSFASWAYDKFSHV 52
C SP+T+PSP++K SP S W K H+
Sbjct: 194 CVASSPSTAPSPTKK--SPNS*MRW*NPKV*HI 286
>TC79745 similar to GP|6714475|gb|AAF26161.1| unknown protein {Arabidopsis
thaliana}, partial (98%)
Length = 577
Score = 26.6 bits (57), Expect = 3.6
Identities = 18/51 (35%), Positives = 27/51 (52%), Gaps = 4/51 (7%)
Frame = +3
Query: 84 APAPGPFASAPQPSSEAAQQAFS----YGKDKAYDIKDKAEVAYEDAKDKA 130
AP PGP++ + A AFS YG K +K KA+ +++ A+ KA
Sbjct: 81 APPPGPYSGTSTLALVARASAFSFGIVYGSIKLKYLKSKAK-SHQKAEAKA 230
>TC89660 similar to GP|11034579|dbj|BAB17103. hypothetical protein~similar
to Arabidopsis thaliana chromosome 3 K15M2.15, partial
(16%)
Length = 1035
Score = 26.6 bits (57), Expect = 3.6
Identities = 23/90 (25%), Positives = 31/90 (33%), Gaps = 4/90 (4%)
Frame = +3
Query: 23 DSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTG----L 78
D+ + P + PP + + S+V PY N P G+G G
Sbjct: 309 DNSNVNYHPPNPTMMPPQYGA---PPNSYVQQPYGNQVPAAG--------GYGPGPVPHY 455
Query: 79 GAPVEAPAPGPFASAPQPSSEAAQQAFSYG 108
G PV P P P PQ + YG
Sbjct: 456 GGPVAGPGPVPGPRGPQTAVGGYPDGSQYG 545
>TC93251 similar to PIR|T39903|T39903 serine-rich protein - fission yeast
(Schizosaccharomyces pombe), partial (4%)
Length = 781
Score = 26.6 bits (57), Expect = 3.6
Identities = 13/35 (37%), Positives = 15/35 (42%)
Frame = -3
Query: 21 RPDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGP 55
+P SP+ SPS S P SW FGP
Sbjct: 644 QPSSPSPSPSCRLSSAKPLPIHSWPLSPLDGRFGP 540
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.311 0.128 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,002,739
Number of Sequences: 36976
Number of extensions: 62503
Number of successful extensions: 631
Number of sequences better than 10.0: 63
Number of HSP's better than 10.0 without gapping: 580
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 622
length of query: 142
length of database: 9,014,727
effective HSP length: 87
effective length of query: 55
effective length of database: 5,797,815
effective search space: 318879825
effective search space used: 318879825
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0283.2


