

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
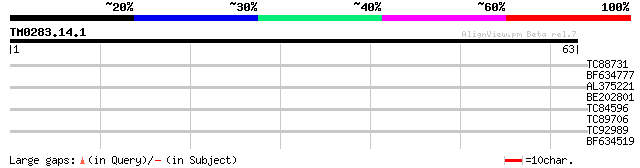
Query= TM0283.14.1
(63 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC88731 similar to GP|21593927|gb|AAM65892.1 unknown {Arabidopsi... 30 0.24
BF634777 homologue to PIR|E86221|E862 hypothetical protein [impo... 25 4.5
AL375221 similar to GP|3128181|gb|A unknown protein {Arabidopsis... 25 5.9
BE202801 homologue to GP|12322744|gb| hypothetical protein; 7880... 25 5.9
TC84596 weakly similar to GP|17064866|gb|AAL32587.1 Unknown prot... 25 5.9
TC89706 similar to GP|3128181|gb|AAC16085.1| unknown protein {Ar... 25 5.9
TC92989 similar to GP|20466294|gb|AAM20464.1 unknown protein {Ar... 25 7.7
BF634519 homologue to GP|18146782|dbj PBng143 {Vigna radiata}, p... 25 7.7
>TC88731 similar to GP|21593927|gb|AAM65892.1 unknown {Arabidopsis
thaliana}, partial (97%)
Length = 1189
Score = 29.6 bits (65), Expect = 0.24
Identities = 14/30 (46%), Positives = 17/30 (56%)
Frame = +3
Query: 17 LAKNMKEQLGDGATFQGIKKAGHLVHLERP 46
+A MKE LG +T Q + GHL HL P
Sbjct: 729 VATYMKEHLGGKSTVQWLDTEGHLPHLSAP 818
>BF634777 homologue to PIR|E86221|E862 hypothetical protein [imported] -
Arabidopsis thaliana, partial (22%)
Length = 677
Score = 25.4 bits (54), Expect = 4.5
Identities = 10/20 (50%), Positives = 12/20 (60%)
Frame = -1
Query: 42 HLERPCVYNRCLKRFIASFF 61
H RPC YNR L ++ FF
Sbjct: 578 HFTRPCKYNRRLX*YLEFFF 519
>AL375221 similar to GP|3128181|gb|A unknown protein {Arabidopsis
thaliana}, partial (17%)
Length = 368
Score = 25.0 bits (53), Expect = 5.9
Identities = 10/23 (43%), Positives = 15/23 (64%)
Frame = -3
Query: 25 LGDGATFQGIKKAGHLVHLERPC 47
L +G ++ ++AGH V L RPC
Sbjct: 81 LPEGEDWREAERAGHRVSLPRPC 13
>BE202801 homologue to GP|12322744|gb| hypothetical protein; 78804-81924
{Arabidopsis thaliana}, partial (3%)
Length = 506
Score = 25.0 bits (53), Expect = 5.9
Identities = 8/16 (50%), Positives = 12/16 (75%)
Frame = +2
Query: 39 HLVHLERPCVYNRCLK 54
H V L PC++NRC++
Sbjct: 143 HKVVLRLPCIHNRCME 190
>TC84596 weakly similar to GP|17064866|gb|AAL32587.1 Unknown protein
{Arabidopsis thaliana}, partial (40%)
Length = 863
Score = 25.0 bits (53), Expect = 5.9
Identities = 8/14 (57%), Positives = 10/14 (71%)
Frame = +3
Query: 39 HLVHLERPCVYNRC 52
H HLE PC ++RC
Sbjct: 27 HKTHLENPCHHHRC 68
>TC89706 similar to GP|3128181|gb|AAC16085.1| unknown protein {Arabidopsis
thaliana}, partial (83%)
Length = 582
Score = 25.0 bits (53), Expect = 5.9
Identities = 10/23 (43%), Positives = 15/23 (64%)
Frame = -2
Query: 25 LGDGATFQGIKKAGHLVHLERPC 47
L +G ++ ++AGH V L RPC
Sbjct: 104 LPEGEDWREAERAGHRVSLPRPC 36
>TC92989 similar to GP|20466294|gb|AAM20464.1 unknown protein {Arabidopsis
thaliana}, partial (39%)
Length = 671
Score = 24.6 bits (52), Expect = 7.7
Identities = 10/26 (38%), Positives = 14/26 (53%)
Frame = -2
Query: 21 MKEQLGDGATFQGIKKAGHLVHLERP 46
M E L + +QG HL H+E+P
Sbjct: 481 MMEDLQEAQVYQGTSSQLHL*HIEKP 404
>BF634519 homologue to GP|18146782|dbj PBng143 {Vigna radiata}, partial (44%)
Length = 692
Score = 24.6 bits (52), Expect = 7.7
Identities = 15/53 (28%), Positives = 25/53 (46%)
Frame = +2
Query: 2 IHLLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLK 54
I LLW E ++ EL+ M ++LGD + L ++ R Y R ++
Sbjct: 404 IKLLWEEFMLFYQMELSSPMSKRLGDYMNKLVLVGFTPLQNMNRTTTYGRNIR 562
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.328 0.142 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,180,372
Number of Sequences: 36976
Number of extensions: 19977
Number of successful extensions: 134
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 134
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 134
length of query: 63
length of database: 9,014,727
effective HSP length: 39
effective length of query: 24
effective length of database: 7,572,663
effective search space: 181743912
effective search space used: 181743912
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0283.14.1


