

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0279a.6
(1582 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
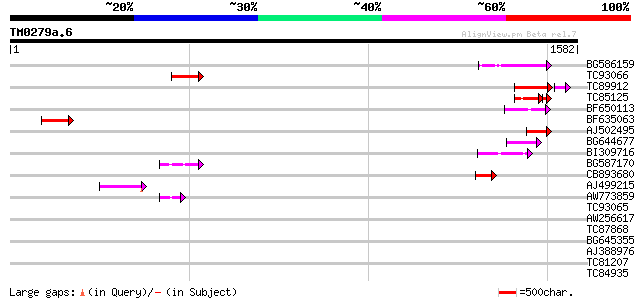
Score E
Sequences producing significant alignments: (bits) Value
BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2... 147 3e-35
TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative ret... 115 1e-25
TC89912 weakly similar to PIR|B84512|B84512 probable retroelemen... 107 7e-25
TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-relate... 94 2e-20
BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vu... 96 1e-19
BF635063 weakly similar to PIR|F84486|F84 probable retroelement ... 73 8e-13
AJ502495 weakly similar to GP|18071369|g putative gag-pol polypr... 65 3e-10
BG644677 weakly similar to GP|7682800|gb| Hypothetical protein T... 64 5e-10
BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F2... 62 2e-09
BG587170 similar to PIR|F86470|F8 probable retroelement polyprot... 62 2e-09
CB893680 weakly similar to GP|1167523|db ORF(AA 1-1338) {Nicotia... 52 2e-06
AJ499215 weakly similar to GP|6642775|gb| gag-pol polyprotein {V... 49 1e-05
AW773859 44 4e-04
TC93065 42 0.001
AW256617 40 0.009
TC87868 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 39 0.021
BG645355 similar to PIR|G96590|G965 hypothetical protein T24C10.... 37 0.061
AJ388976 similar to PIR|E84638|E84 probable RSZp22 splicing fact... 37 0.061
TC81207 similar to GP|21322752|dbj|BAB78536. cold shock protein-... 37 0.079
TC84935 similar to PIR|G96631|G96631 probable RNA-binding protei... 36 0.14
>BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2O9.150 -
Arabidopsis thaliana, partial (11%)
Length = 732
Score = 147 bits (371), Expect = 3e-35
Identities = 82/205 (40%), Positives = 120/205 (58%)
Frame = +1
Query: 1308 IHREKGAKKLWLSQNSYVEGVLSRFDMSKANPVSTPLANHFKLSLEQCPKTASEIEGMSK 1367
I E+G +++ Q YV +L RF M K+N P+A KL ++ +++
Sbjct: 10 IQNEEG---IYICQRKYVTDLLERFGMEKSNLSRNPIAPRCKLIKDE---NGVKVDATK- 168
Query: 1368 ISHASAVGCLMYVMVCTRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYLKGTADRGIMF 1427
+ VGCLMY + TRPDL S + +FM+ P + H AVK +LRYL GT + GIM+
Sbjct: 169 --YKQIVGCLMY-LAATRPDLMYVLSLISRFMNCPTELHMHAVKRVLRYLNGTINLGIMY 339
Query: 1428 SREQGVVPLVVGYVDSDYAGDLDDRRSTTGYVFTLAGGPICWKSSVQSIVAMSTTEAEYM 1487
R + Y DSDYAGDLDDR+ST+GYVF L+ G + W S Q +V +STT+AE++
Sbjct: 340 KRNGS--EKLEAYTDSDYAGDLDDRKSTSGYVFMLSSGAVSWSSKKQPVVTLSTTKAEFI 513
Query: 1488 AVAEAVKEALWLTGLVKKLGVEQGG 1512
A A +++W+ +++KLG Q G
Sbjct: 514 AAAFCACQSVWMRRVLEKLGYTQSG 588
>TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative retroelement
{Oryza sativa} [Oryza sativa (japonica cultivar-group)],
partial (10%)
Length = 823
Score = 115 bits (288), Expect = 1e-25
Identities = 51/90 (56%), Positives = 65/90 (71%), Gaps = 1/90 (1%)
Frame = +1
Query: 451 CKYCVL-GKQCRVRFKTGQHKTKGILDYVHSDVWGPTKEPSVGGFRYFVTFTDDFSRKVW 509
CK+ + G + +V F T H+TKGILDY+HSD+WGP+K S GG RY +T DDF RKVW
Sbjct: 22 CKHLLFFGNRKKVSFSTATHRTKGILDYIHSDLWGPSKVTSYGGRRYMMTIIDDFPRKVW 201
Query: 510 VYFLKYKSEVFAKFKLWKAEVENQTGRKIK 539
VYFL+YK+E F FK W+ VE QTG+ +K
Sbjct: 202 VYFLRYKNETFPTFKKWRILVETQTGKNVK 291
>TC89912 weakly similar to PIR|B84512|B84512 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(10%)
Length = 814
Score = 107 bits (267), Expect(2) = 7e-25
Identities = 47/108 (43%), Positives = 77/108 (70%)
Frame = +1
Query: 1408 EAVKWILRYLKGTADRGIMFSREQGVVPLVVGYVDSDYAGDLDDRRSTTGYVFTLAGGPI 1467
+A+KW+L+YL + + +++ + GYVD+DYAG++D R+S +G+VFTL G I
Sbjct: 1 QALKWVLKYLNESLKSSLKYTKAAQEEDALEGYVDADYAGNVDTRKSLSGFVFTLYGTTI 180
Query: 1468 CWKSSVQSIVAMSTTEAEYMAVAEAVKEALWLTGLVKKLGVEQGGVQL 1515
WK++ QS+V +STT+AEY+A E VK+A+WL G++ +LG+ Q V++
Sbjct: 181 SWKANQQSVVTLSTTQAEYIAFVEGVKDAIWLKGMIGELGITQEYVKI 324
Score = 26.6 bits (57), Expect(2) = 7e-25
Identities = 14/47 (29%), Positives = 24/47 (50%)
Frame = +2
Query: 1519 WRTIRCIMLEPSILM*GSTRSGSCWRPNRYYFRRFTHQRMQPTC*PS 1565
WR I+CIM S L T + R +++++ +R++ C PS
Sbjct: 353 WRIIKCIMRGLSTLTFACTLLET*LNQKRLWWKKWHRKRIRRMCLPS 493
>TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
(EC 3.4.23.-);, partial (7%)
Length = 705
Score = 94.4 bits (233), Expect(2) = 2e-20
Identities = 48/79 (60%), Positives = 59/79 (73%)
Frame = +1
Query: 1410 VKWILRYLKGTADRGIMFSREQGVVPLVVGYVDSDYAGDLDDRRSTTGYVFTLAGGPICW 1469
VK I+RY+KGT+ + F G V GYVDSD+AGD D R+STTGYVFTLAGG + W
Sbjct: 1 VKRIMRYIKGTSGVAVCFG---GSELTVRGYVDSDFAGDHDKRKSTTGYVFTLAGGAVSW 171
Query: 1470 KSSVQSIVAMSTTEAEYMA 1488
S +Q++VA+STTEAEYMA
Sbjct: 172 LSKLQTVVALSTTEAEYMA 228
Score = 24.6 bits (52), Expect(2) = 2e-20
Identities = 9/23 (39%), Positives = 18/23 (78%)
Frame = +3
Query: 1488 AVAEAVKEALWLTGLVKKLGVEQ 1510
++ +A KEA+W+ L+++LG +Q
Sbjct: 228 SLPQACKEAIWMQRLMEELGHKQ 296
>BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vulgaris},
partial (13%)
Length = 494
Score = 95.5 bits (236), Expect = 1e-19
Identities = 54/131 (41%), Positives = 77/131 (58%), Gaps = 1/131 (0%)
Frame = +1
Query: 1380 VMVCTRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYLKGTADRGIMFSR-EQGVVPLVV 1438
+ +C RPD+ + S + KFM P K H A ILRY++GT + G++F + V ++
Sbjct: 106 INLC*RPDICYSVSVISKFMHDPRKPHLIAANRILRYVRGTMEYGLLFPYGAKSEVYELI 285
Query: 1439 GYVDSDYAGDLDDRRSTTGYVFTLAGGPICWKSSVQSIVAMSTTEAEYMAVAEAVKEALW 1498
Y DSD+ GD RRST+GYVF I W + Q I A+S+ EAEY+A A +ALW
Sbjct: 286 CYSDSDWCGD---RRSTSGYVFKFNDAAISWCTKKQPITALSSYEAEYIAGTFATFQALW 456
Query: 1499 LTGLVKKLGVE 1509
L ++K+L E
Sbjct: 457 LDSVIKELKCE 489
>BF635063 weakly similar to PIR|F84486|F84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(4%)
Length = 677
Score = 73.2 bits (178), Expect = 8e-13
Identities = 35/88 (39%), Positives = 58/88 (65%), Gaps = 1/88 (1%)
Frame = -2
Query: 90 MSKTLMNKLFAKQRLYSLKMQEGGDLQAHVYAFNNILADLTRLGVTVDDEDKTIILLCSL 149
M+K+L ++ F KQ+LYS +M E + + FN IL DL + V ++DE+K I+LLC+L
Sbjct: 295 MTKSLAHRQFLKQQLYSFRMVESKAIMEQLTEFNKILDDLENIEVQLEDEEKAILLLCAL 116
Query: 150 PGSYDHLVTTLTYGKD-SITLDSISSTL 176
P S++ T+ YGK+ ++TL+ + + L
Sbjct: 115 PKSFESFKDTMLYGKEGTVTLEEVQAAL 32
>AJ502495 weakly similar to GP|18071369|g putative gag-pol polyprotein {Oryza
sativa}, partial (9%)
Length = 542
Score = 64.7 bits (156), Expect = 3e-10
Identities = 30/68 (44%), Positives = 42/68 (61%)
Frame = +2
Query: 1443 SDYAGDLDDRRSTTGYVFTLAGGPICWKSSVQSIVAMSTTEAEYMAVAEAVKEALWLTGL 1502
SD+AGD + R+ST+GY F L G I W S Q +VA ST EAEY+A + +WL +
Sbjct: 2 SDWAGDTETRKSTSGYAFHLGTGAISWSSKKQPVVAFSTAEAEYIASTSCATQTVWLRRI 181
Query: 1503 VKKLGVEQ 1510
++ + EQ
Sbjct: 182 LEVMHHEQ 205
>BG644677 weakly similar to GP|7682800|gb| Hypothetical protein T15F17.l
{Arabidopsis thaliana}, partial (3%)
Length = 539
Score = 63.9 bits (154), Expect = 5e-10
Identities = 33/99 (33%), Positives = 56/99 (56%)
Frame = -3
Query: 1386 PDLAQAASQVCKFMSKPGKQHWEAVKWILRYLKGTADRGIMFSREQGVVPLVVGYVDSDY 1445
P++ + + + ++ S P +H +K I +YLKG D G+ +S++ P ++GYV++ Y
Sbjct: 510 PNITFSINLLSRYSSAPTMRH*NGIKHICKYLKGIIDMGLFYSKDCS--PDLIGYVNA*Y 337
Query: 1446 AGDLDDRRSTTGYVFTLAGGPICWKSSVQSIVAMSTTEA 1484
D RS TGY+FT I W+S+ S +A S+ A
Sbjct: 336 LSDPHKARS*TGYIFTCGNTVISWRSTK*STIATSSNHA 220
>BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (10%)
Length = 744
Score = 62.0 bits (149), Expect = 2e-09
Identities = 45/155 (29%), Positives = 74/155 (47%)
Frame = +2
Query: 1304 LGMEIHREKGAKKLWLSQNSYVEGVLSRFDMSKANPVSTPLANHFKLSLEQCPKTASEIE 1363
LG+E+ R K + + L+Q Y +L TP KL P E +
Sbjct: 293 LGLEVARSK--QGILLNQRKYTLELLEDSGNLAVKSTLTPYDISLKLHNSDSPLYNDETQ 466
Query: 1364 GMSKISHASAVGCLMYVMVCTRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYLKGTADR 1423
+ +G L+Y + TRPD++ A Q+ +F+SKP + H++A +L+YLK +
Sbjct: 467 ------YRRLIGKLIY-LTTTRPDISFAVQQLSQFVSKPQQVHYQAAIRVLQYLKTAPAK 625
Query: 1424 GIMFSREQGVVPLVVGYVDSDYAGDLDDRRSTTGY 1458
G+ +S + + + DSD+A R+S TGY
Sbjct: 626 GLFYSATSNL--KLSSFADSDWATCPTTRKSVTGY 724
>BG587170 similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 718
Score = 62.0 bits (149), Expect = 2e-09
Identities = 38/124 (30%), Positives = 58/124 (46%), Gaps = 2/124 (1%)
Frame = -3
Query: 418 LWHMRLGHLSERGMMELYKRNLLKGVRSCTIGLCKYCVLGKQCRVRFKTGQHKTKGILDY 477
LWH RLGH R + + + + C+ C+LGK C+ F + D
Sbjct: 584 LWHARLGHPHGRALNLMLPGVVFENKN------CEACILGKHCKNVFPRTSTVYENCFDL 423
Query: 478 VHSDVWGPTKEPSVG--GFRYFVTFTDDFSRKVWVYFLKYKSEVFAKFKLWKAEVENQTG 535
+++D+W PS+ +YFVTF D+ S+ W+ + K V FK ++A V N
Sbjct: 422 IYTDLW---TAPSLSRDNHKYFVTFIDEKSKYTWLTLIPSKDRVIDAFKNFQAYVTNHYH 252
Query: 536 RKIK 539
KIK
Sbjct: 251 AKIK 240
>CB893680 weakly similar to GP|1167523|db ORF(AA 1-1338) {Nicotiana tabacum},
partial (7%)
Length = 780
Score = 52.0 bits (123), Expect = 2e-06
Identities = 31/59 (52%), Positives = 38/59 (63%)
Frame = -3
Query: 1300 AKKILGMEIHREKGAKKLWLSQNSYVEGVLSRFDMSKANPVSTPLANHFKLSLEQCPKT 1358
AKKI+GM+I +K L LSQ Y+ VL F+M A VST LA+HF LS EQ P+T
Sbjct: 184 AKKIIGMQIMIDKQKGVL*LSQVEYITRVLQIFNMGNAILVSTTLASHFCLSHEQSPQT 8
>AJ499215 weakly similar to GP|6642775|gb| gag-pol polyprotein {Vitis
vinifera}, partial (18%)
Length = 567
Score = 49.3 bits (116), Expect = 1e-05
Identities = 37/142 (26%), Positives = 63/142 (44%), Gaps = 12/142 (8%)
Frame = +3
Query: 252 GNSSSAANVVQSDGSCSEEDLLCVSYVKCTDA---WVLDSGCSCHMTPHREWFNSFKSCD 308
G+S S N + S+E LL +S W++DSGC+ HMT R+ F
Sbjct: 3 GDSPSHKNKHKPHQ*GSKEQLLAMSTFATKQPSKYWLIDSGCTHHMTHDRDLFKELNKST 182
Query: 309 FGYVYLGDDKPCIIKGM*QVKIALDDGGVRTLSQVRYVPEVTKNLISLGTLHENGYS--- 365
V + + ++G+ V + G + +S V Y P++ ++L+S+ L GY
Sbjct: 183 ISKVRMLNGAHIEVEGIGTV-LVKSHSGYKQISNVLYAPKLNQSLLSVPQLLTKGYKVLF 359
Query: 366 ------FKSEENRDILRVSKGA 381
K + N+++LR A
Sbjct: 360 EHEKCVIKDQNNKEVLRAQMKA 425
>AW773859
Length = 538
Score = 44.3 bits (103), Expect = 4e-04
Identities = 20/74 (27%), Positives = 35/74 (47%)
Frame = -3
Query: 418 LWHMRLGHLSERGMMELYKRNLLKGVRSCTIGLCKYCVLGKQCRVRFKTGQHKTKGILDY 477
LWH RLGHLS R ++ L+ + ++ C C + ++ F+ ++ +
Sbjct: 230 LWHFRLGHLSNRKLLSLHSNFPFITIDQNSV--CDICHYSRHKKLPFQLSTNRASKCYEL 57
Query: 478 VHSDVWGPTKEPSV 491
H D+WGP S+
Sbjct: 56 FHFDIWGPFSTQSI 15
>TC93065
Length = 783
Score = 42.4 bits (98), Expect = 0.001
Identities = 43/194 (22%), Positives = 82/194 (42%), Gaps = 13/194 (6%)
Frame = +2
Query: 101 KQRLYSLKMQEGGDLQAHVYAFNNILADLTRLGVTVDDEDKTIILLCSLPGSYDHLVTTL 160
++ +LKM+E ++ + ++ + LG + D+ +L LP ++ +++L
Sbjct: 218 RREFEALKMKETETVREFSDRISKVVTQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSL 397
Query: 161 TYGKD--SITLDSISSTLLQHAQRRRSVEEGGGSSGEGLFVKGGQDRGR-----GKGKAV 213
K+ IT+ + + L QRR E + EG F+ + + + GK K
Sbjct: 398 EENKNFSEITVAELVNALQASEQRRSLRME---ENVEGAFLANNKGKNQSFKSFGKKKFP 568
Query: 214 DSGKKKRSKSKD-----RKTAECYSCKQIGHWKRDCPNRSGKSGNSSSAANVVQSDGSCS 268
K+ D R +C +C Q+GH ++ C N++ + + Q D
Sbjct: 569 PCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQEQEARVVEHHQED---- 736
Query: 269 EEDLLCVS-YVKCT 281
EE L S Y+ C+
Sbjct: 737 EEPLFKASCYLACS 778
>AW256617
Length = 718
Score = 39.7 bits (91), Expect = 0.009
Identities = 24/72 (33%), Positives = 39/72 (53%)
Frame = -2
Query: 347 PEVTKNLISLGTLHENGYSFKSEENRDILRVSKGAMTVMRAKRTAGNIYKLLGSTVMGDV 406
P + K S+ TL NG++ + ++DILR+ GA LLG T++ D+
Sbjct: 330 P*IKK*FDSMHTL*ANGFANRINRDKDILRLRNGA---------------LLGKTIV*DI 196
Query: 407 ASVETDDDATKL 418
ASV+ D+DA ++
Sbjct: 195 ASVDPDNDAAEI 160
>TC87868 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana, partial (91%)
Length = 860
Score = 38.5 bits (88), Expect = 0.021
Identities = 22/61 (36%), Positives = 27/61 (44%)
Frame = +3
Query: 189 GGGSSGEGLFVKGGQDRGRGKGKAVDSGKKKRSKSKDRKTAECYSCKQIGHWKRDCPNRS 248
GGG G G GG+ RG G G D +CY C + GH+ R+C NR
Sbjct: 264 GGGGGGGG----GGRGRGGGGGGGSD--------------LKCYECGEPGHFARECRNRG 389
Query: 249 G 249
G
Sbjct: 390 G 392
>BG645355 similar to PIR|G96590|G965 hypothetical protein T24C10.5 [imported]
- Arabidopsis thaliana, partial (5%)
Length = 627
Score = 37.0 bits (84), Expect = 0.061
Identities = 18/48 (37%), Positives = 22/48 (45%), Gaps = 8/48 (16%)
Frame = -1
Query: 228 TAECYSCKQIGHWKRDCPNRSG---KSGNSSSAANVV-----QSDGSC 267
+ CY C Q GHW +CPN S GNS++ Q DG C
Sbjct: 411 SGNCYKCNQPGHWANNCPNMSAAPQSHGNSNTGQGRYGIAQNQPDGRC 268
Score = 34.7 bits (78), Expect = 0.30
Identities = 14/34 (41%), Positives = 20/34 (58%), Gaps = 3/34 (8%)
Frame = -1
Query: 228 TAECYSCKQIGHWKRDCPNRSGK---SGNSSSAA 258
+ +CY C+Q GHW +CP+ S SG S A+
Sbjct: 510 SGKCYKCQQPGHWASNCPSMSAANRVSGGSGGAS 409
>AJ388976 similar to PIR|E84638|E84 probable RSZp22 splicing factor
[imported] - Arabidopsis thaliana, partial (62%)
Length = 508
Score = 37.0 bits (84), Expect = 0.061
Identities = 22/64 (34%), Positives = 26/64 (40%)
Frame = +2
Query: 189 GGGSSGEGLFVKGGQDRGRGKGKAVDSGKKKRSKSKDRKTAECYSCKQIGHWKRDCPNRS 248
GGG G G GG RGR G D +CY C + GH+ R C +
Sbjct: 260 GGGGGGRGGGGGGGGGRGRSGGGGSD--------------LKCYXCGEPGHFARXCNSSP 397
Query: 249 GKSG 252
G SG
Sbjct: 398 GGSG 409
>TC81207 similar to GP|21322752|dbj|BAB78536. cold shock protein-1 {Triticum
aestivum}, partial (39%)
Length = 630
Score = 36.6 bits (83), Expect = 0.079
Identities = 25/87 (28%), Positives = 33/87 (37%), Gaps = 17/87 (19%)
Frame = +3
Query: 184 RSVEEGGGSSGEGLFVKGGQDRGRGKGK---------AVDSGKKKRSKSKDR-------- 226
R GGG G G +GG+ R G G A D + R+ DR
Sbjct: 276 RQDNHGGGGGGRGF--RGGERRNGGGGCYTCGDTGHIARDCDRSDRNDRNDRSGGGGGGD 449
Query: 227 KTAECYSCKQIGHWKRDCPNRSGKSGN 253
+ CY+C H+ RDC G + N
Sbjct: 450 RDRACYTCGSFEHFARDCMRGGGNNNN 530
Score = 33.9 bits (76), Expect = 0.51
Identities = 21/87 (24%), Positives = 31/87 (35%)
Frame = +3
Query: 170 DSISSTLLQHAQRRRSVEEGGGSSGEGLFVKGGQDRGRGKGKAVDSGKKKRSKSKDRKTA 229
D + T+ + + G GE L V+ G G G+ G+++
Sbjct: 186 DRVEFTIADGDNGKTKAVDVTGPKGEPLQVRQDNHGGGGGGRGFRGGERRNGGGG----- 350
Query: 230 ECYSCKQIGHWKRDCPNRSGKSGNSSS 256
CY+C GH RDC N S
Sbjct: 351 -CYTCGDTGHIARDCDRSDRNDRNDRS 428
>TC84935 similar to PIR|G96631|G96631 probable RNA-binding protein F8A5.17
[imported] - Arabidopsis thaliana, partial (41%)
Length = 552
Score = 35.8 bits (81), Expect = 0.14
Identities = 18/53 (33%), Positives = 24/53 (44%), Gaps = 1/53 (1%)
Frame = +2
Query: 201 GGQDRGRGKGKAVDSGKKKRSKSKDRKTAE-CYSCKQIGHWKRDCPNRSGKSG 252
GG D + SG + + DR + C+ C + GHW RDCP G G
Sbjct: 380 GGDDADQRYRGGFSSGGRGSYGAGDRVGQDDCFKCGRPGHWARDCPLAGGDGG 538
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.341 0.149 0.500
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 52,931,827
Number of Sequences: 36976
Number of extensions: 822491
Number of successful extensions: 7107
Number of sequences better than 10.0: 60
Number of HSP's better than 10.0 without gapping: 3851
Number of HSP's successfully gapped in prelim test: 273
Number of HSP's that attempted gapping in prelim test: 2833
Number of HSP's gapped (non-prelim): 4636
length of query: 1582
length of database: 9,014,727
effective HSP length: 109
effective length of query: 1473
effective length of database: 4,984,343
effective search space: 7341937239
effective search space used: 7341937239
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0279a.6


