

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0278.5.1
(30 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
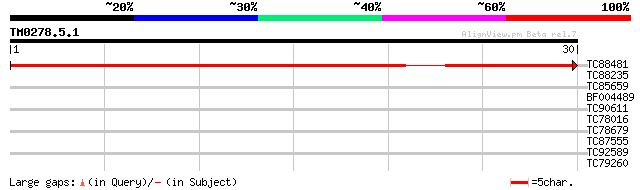
TC88481 similar to PIR|T00806|T02445 probable U4/U6 small nuclea... 44 1e-05
TC88235 similar to GP|12656803|gb|AAK00964.1 unknown protein {Or... 27 2.4
TC85659 SP|O24076|GBLP_MEDSA Guanine nucleotide-binding protein ... 26 4.0
BF004489 similar to GP|20161344|d contains ESTs AU086067(S4642) ... 25 6.9
TC90611 similar to GP|10177223|dbj|BAB10298. contains similarity... 25 6.9
TC78016 similar to GP|21537191|gb|AAM61532.1 PRL1 protein {Arabi... 25 6.9
TC78679 similar to GP|20466538|gb|AAM20586.1 putative WD-repeat ... 25 6.9
TC87555 weakly similar to GP|22775656|dbj|BAC15510. contains EST... 25 6.9
TC92589 similar to GP|11141605|gb|AAG32022.1 LEUNIG {Arabidopsis... 25 6.9
TC79260 weakly similar to GP|9759350|dbj|BAB10005.1 WD-repeat pr... 25 9.0
>TC88481 similar to PIR|T00806|T02445 probable U4/U6 small nuclear
ribonucleoprotein [imported] - Arabidopsis thaliana,
partial (60%)
Length = 1836
Score = 44.3 bits (103), Expect = 1e-05
Identities = 22/30 (73%), Positives = 22/30 (73%)
Frame = +1
Query: 1 GGYIVTVSHDRTIKLWSSNMTNDNEHTMDV 30
GGYIVTVSHDRTIKLWS T H MDV
Sbjct: 1744 GGYIVTVSHDRTIKLWSXXTT--XXHAMDV 1827
>TC88235 similar to GP|12656803|gb|AAK00964.1 unknown protein {Oryza sativa},
partial (91%)
Length = 1080
Score = 26.6 bits (57), Expect = 2.4
Identities = 9/18 (50%), Positives = 13/18 (72%)
Frame = +2
Query: 1 GGYIVTVSHDRTIKLWSS 18
G Y++T S D T +LWS+
Sbjct: 1016 GAYLITASSDSTARLWST 1069
>TC85659 SP|O24076|GBLP_MEDSA Guanine nucleotide-binding protein beta
subunit-like protein. [Alfalfa] {Medicago sativa},
complete
Length = 1261
Score = 25.8 bits (55), Expect = 4.0
Identities = 10/15 (66%), Positives = 13/15 (86%)
Frame = +2
Query: 4 IVTVSHDRTIKLWSS 18
IV+ S DRTIKLW++
Sbjct: 419 IVSASRDRTIKLWNT 463
Score = 25.0 bits (53), Expect = 6.9
Identities = 11/19 (57%), Positives = 15/19 (78%)
Frame = +2
Query: 4 IVTVSHDRTIKLWSSNMTN 22
IV+ S DRT+K+W N+TN
Sbjct: 557 IVSASWDRTVKVW--NLTN 607
>BF004489 similar to GP|20161344|d contains ESTs AU086067(S4642)
D41819(S4642)~similar to Arabidopsis thaliana chromosome
1 At1g49040~, partial (16%)
Length = 605
Score = 25.0 bits (53), Expect = 6.9
Identities = 8/16 (50%), Positives = 12/16 (75%)
Frame = +1
Query: 1 GGYIVTVSHDRTIKLW 16
G ++T SHD T+K+W
Sbjct: 178 GERVLTASHDGTVKMW 225
>TC90611 similar to GP|10177223|dbj|BAB10298. contains similarity to
GTP-binding regulatory protein and WD-repeat
protein~gene_id:MPF21.14, partial (40%)
Length = 855
Score = 25.0 bits (53), Expect = 6.9
Identities = 9/16 (56%), Positives = 13/16 (81%)
Frame = +1
Query: 1 GGYIVTVSHDRTIKLW 16
G ++ +VS DRTIK+W
Sbjct: 517 GKFLYSVSWDRTIKIW 564
>TC78016 similar to GP|21537191|gb|AAM61532.1 PRL1 protein {Arabidopsis
thaliana}, partial (81%)
Length = 1759
Score = 25.0 bits (53), Expect = 6.9
Identities = 9/13 (69%), Positives = 11/13 (84%)
Frame = +3
Query: 4 IVTVSHDRTIKLW 16
+VT SHD TIK+W
Sbjct: 1011 VVTGSHDSTIKMW 1049
>TC78679 similar to GP|20466538|gb|AAM20586.1 putative WD-repeat protein
{Arabidopsis thaliana}, partial (55%)
Length = 1198
Score = 25.0 bits (53), Expect = 6.9
Identities = 12/28 (42%), Positives = 18/28 (63%), Gaps = 5/28 (17%)
Frame = +1
Query: 1 GGYIVTVSHDRTIKLW-----SSNMTND 23
G Y ++ D+TIKLW SSN+T++
Sbjct: 487 GRYFISNGKDQTIKLWDIRKMSSNVTSN 570
>TC87555 weakly similar to GP|22775656|dbj|BAC15510. contains EST
C27990(C53630)~similar to G-protein beta family {Oryza
sativa (japonica cultivar-group), partial (33%)
Length = 685
Score = 25.0 bits (53), Expect = 6.9
Identities = 9/23 (39%), Positives = 12/23 (52%)
Frame = +1
Query: 4 IVTVSHDRTIKLWSSNMTNDNEH 26
+ T S DRT++LW N H
Sbjct: 583 LATASKDRTLRLWKINTEEATNH 651
>TC92589 similar to GP|11141605|gb|AAG32022.1 LEUNIG {Arabidopsis thaliana},
partial (5%)
Length = 415
Score = 25.0 bits (53), Expect = 6.9
Identities = 8/15 (53%), Positives = 11/15 (73%)
Frame = +1
Query: 2 GYIVTVSHDRTIKLW 16
G + + SHDR +KLW
Sbjct: 103 GLVASASHDRFVKLW 147
>TC79260 weakly similar to GP|9759350|dbj|BAB10005.1 WD-repeat protein-like
{Arabidopsis thaliana}, partial (50%)
Length = 1227
Score = 24.6 bits (52), Expect = 9.0
Identities = 11/23 (47%), Positives = 15/23 (64%)
Frame = +2
Query: 1 GGYIVTVSHDRTIKLWSSNMTND 23
G Y+ T S+DRT +W + TND
Sbjct: 581 GKYLATASNDRTAIIWEVD-TND 646
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.313 0.128 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,033,420
Number of Sequences: 36976
Number of extensions: 4781
Number of successful extensions: 65
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 58
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 65
length of query: 30
length of database: 9,014,727
effective HSP length: 6
effective length of query: 24
effective length of database: 8,792,871
effective search space: 211028904
effective search space used: 211028904
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0278.5.1


