

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0269a.3
(490 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
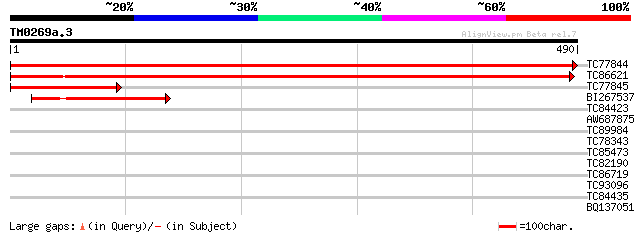
TC77844 similar to PIR|T49049|T49049 hypothetical protein T5P19.... 847 0.0
TC86621 similar to PIR|F84664|F84664 hypothetical protein At2g26... 587 e-168
TC77845 weakly similar to PIR|T49049|T49049 hypothetical protein... 162 4e-40
BI267537 similar to PIR|F84664|F846 hypothetical protein At2g267... 130 9e-31
TC84423 similar to PIR|F84730|F84730 probable myosin heavy chain... 38 0.008
AW687875 similar to PIR|T05990|T05 hypothetical protein F17M5.15... 33 0.19
TC89984 similar to GP|16604601|gb|AAL24093.1 unknown protein {Ar... 33 0.25
TC78343 weakly similar to GP|15724195|gb|AAL06489.1 AT5g49210/K2... 33 0.32
TC85473 homologue to SP|Q9XGM1|VATD_ARATH Vacuolar ATP synthase ... 33 0.32
TC82190 weakly similar to GP|13937297|gb|AAK50128.1 unknown prot... 32 0.42
TC86719 similar to GP|599956|emb|CAA83434.1| chloroplast inner m... 30 1.6
TC93096 29 4.7
TC84435 similar to GP|20259307|gb|AAM14389.1 putative nodulin pr... 28 6.1
BQ137051 homologue to GP|20259307|gb putative nodulin protein {A... 28 6.1
>TC77844 similar to PIR|T49049|T49049 hypothetical protein T5P19.130 -
Arabidopsis thaliana, partial (70%)
Length = 1969
Score = 847 bits (2187), Expect = 0.0
Identities = 433/490 (88%), Positives = 463/490 (94%)
Frame = +2
Query: 1 MTKISPEIEIKMPMEAVPPVSADVSFISDRFPKYKLGADSQILEETVEDNQGPSLKDVIE 60
MTKISPEIE KMPMEAVPPVSADVSFIS+ FPKYKLGAD Q+ +ET EDNQGPSLK+VIE
Sbjct: 281 MTKISPEIETKMPMEAVPPVSADVSFISNSFPKYKLGADHQVFQETAEDNQGPSLKEVIE 460
Query: 61 QEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKIREVASLEGHVLLKKLRDALEYL 120
QEA+NLS+Q+ RISVRDLA KFDKNL+AAAKLSNEAK+REV SLEGHVLLKKLRDALEYL
Sbjct: 461 QEASNLSEQNNRISVRDLASKFDKNLSAAAKLSNEAKLREVPSLEGHVLLKKLRDALEYL 640
Query: 121 RVRFAGRNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQASEDAKKLVNQ 180
+ RF GRNKEDV AISMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQASEDAKKLVNQ
Sbjct: 641 KGRFTGRNKEDVANAISMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQASEDAKKLVNQ 820
Query: 181 EKSFACAEIESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEVQEARRIKLLHQPSKV 240
EKSFACAEIESAR+VVLRIGEALEEQE+ ++ASKPQDVDGLVEEVQEARRI+LLHQPSKV
Sbjct: 821 EKSFACAEIESARAVVLRIGEALEEQEKVTEASKPQDVDGLVEEVQEARRIRLLHQPSKV 1000
Query: 241 MAMEYELRALRDQIREKSIFSIKLQKELTMSKRDEENKSHPYMLHGSEALGSYLKVQPRS 300
MAMEYELRALRDQI+EKS+FSI+LQKELTMSK D+ENKSH Y L GSEALGSY++V+P +
Sbjct: 1001MAMEYELRALRDQIQEKSVFSIQLQKELTMSKWDKENKSHSYKLDGSEALGSYMQVKPCT 1180
Query: 301 GEVPQVSKCSFQWYRLSSEGSWREVISGADKSIYAPDPFDVGRILQVDIVSNGKKLTLTT 360
EVPQVSKCSFQWYRLSSEGSWREVISGA+KSIYAPDP DVGR+LQVDIVS+GKKLTLTT
Sbjct: 1181SEVPQVSKCSFQWYRLSSEGSWREVISGANKSIYAPDPLDVGRMLQVDIVSDGKKLTLTT 1360
Query: 361 NPIQTASGLGSHVEALLRKSNTDFHVVISQMNGKDHSSRSTHSFNVGRMRIKLCRGWITK 420
NPIQT GLGS VEALLRKSNTDF+VVISQMNGKDHSS STHSFNVGRMRIKLCRGWITK
Sbjct: 1361NPIQTVPGLGSQVEALLRKSNTDFNVVISQMNGKDHSSHSTHSFNVGRMRIKLCRGWITK 1540
Query: 421 AREIYSPSMQLCGVRGDASNAAKALFWQARKGLSFVLTFESEKERNVAIMIARKYALDCN 480
AREIYSP+MQLCGVR D SNAAK LFWQARKGLSFVLTFESEK+RNVAIMIARK+ALDCN
Sbjct: 1541AREIYSPTMQLCGVRSDVSNAAKTLFWQARKGLSFVLTFESEKDRNVAIMIARKHALDCN 1720
Query: 481 VVLAGPDDLV 490
VVLAGPDDLV
Sbjct: 1721VVLAGPDDLV 1750
>TC86621 similar to PIR|F84664|F84664 hypothetical protein At2g26770
[imported] - Arabidopsis thaliana, partial (97%)
Length = 1890
Score = 587 bits (1513), Expect = e-168
Identities = 302/489 (61%), Positives = 391/489 (79%), Gaps = 1/489 (0%)
Frame = +1
Query: 1 MTKISPEIEIKMPMEAVPPVSADVSFISDRFPKYKLGADSQILEETVEDNQGPSLKDVIE 60
MT+++ + M EAVP VS+DV F + RFP Y++GA++QI+E +D + S+K+VI
Sbjct: 130 MTRVTRDFGDTMQKEAVPAVSSDVVFATSRFPNYRIGANNQIMEAK-DDPKVLSMKEVIA 306
Query: 61 QEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKIREVASLEGHVLLKKLRDALEYL 120
+E L DQH R+SVRDLA KF+K LAAAAKLS EA++RE ASLE HVLLKKLRD+LE L
Sbjct: 307 RETAMLLDQHNRLSVRDLASKFEKGLAAAAKLSEEARLREAASLEKHVLLKKLRDSLESL 486
Query: 121 RVRFAGRNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQASEDAKKLVNQ 180
+ R AGRNK+DVE AI+MVEALAV+LTQ EGEL+QEK EVKKL NFLKQASEDAKKLV++
Sbjct: 487 KGRVAGRNKDDVEDAIAMVEALAVQLTQREGELLQEKAEVKKLANFLKQASEDAKKLVDE 666
Query: 181 EKSFACAEIESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEVQEARRIKLLHQPSKV 240
E++FA AEI++ARS V R+ E+L+E E+ SQAS QDV+ L++EVQEARRIK+LHQPSKV
Sbjct: 667 ERAFARAEIDNARSAVQRVEESLQEHERMSQASGKQDVEQLMKEVQEARRIKMLHQPSKV 846
Query: 241 MAMEYELRALRDQIREKSIFSIKLQKELTMSKRDEENKSHPYMLHGSEALGSYLKVQPRS 300
M ME+EL+ALR Q+ EK+ ++LQKE+T +K+ EEN H Y L G+E LGSYL++QP S
Sbjct: 847 MDMEHELQALRAQLAEKTRHYLRLQKEITRTKKGEENVPHLYELEGNETLGSYLQIQPCS 1026
Query: 301 GEVPQVSKCSFQWYRLSSEGSWREVISGADKSIYAPDPFDVGRILQVDIVSNGKKLTL-T 359
P +S CS QW R+SS+G+ +E+ISGA KS+YAP+PFDVGRIL VD++S + + L T
Sbjct: 1027DNAPDLSNCSIQWCRVSSDGAKKELISGAIKSVYAPEPFDVGRILHVDVISENQHIILST 1206
Query: 360 TNPIQTASGLGSHVEALLRKSNTDFHVVISQMNGKDHSSRSTHSFNVGRMRIKLCRGWIT 419
T PI A+GLG++VEAL+RK +T+F+VV++QM+G H + S H +VG+MRIKLC+G T
Sbjct: 1207TGPIDPAAGLGTYVEALVRKHDTEFNVVVTQMSGLHHPTESIHVLHVGKMRIKLCKGKTT 1386
Query: 420 KAREIYSPSMQLCGVRGDASNAAKALFWQARKGLSFVLTFESEKERNVAIMIARKYALDC 479
A+E YS SMQLCGVRG + AA+ALFWQ ++GLSFVL FESE+ERN AIM+AR++A DC
Sbjct: 1387IAKEYYSSSMQLCGVRGGGNAAAQALFWQPKQGLSFVLAFESERERNAAIMLARRFAFDC 1566
Query: 480 NVVLAGPDD 488
N++LAGPDD
Sbjct: 1567NIMLAGPDD 1593
>TC77845 weakly similar to PIR|T49049|T49049 hypothetical protein T5P19.130
- Arabidopsis thaliana, partial (16%)
Length = 793
Score = 162 bits (409), Expect = 4e-40
Identities = 81/96 (84%), Positives = 89/96 (92%)
Frame = +3
Query: 1 MTKISPEIEIKMPMEAVPPVSADVSFISDRFPKYKLGADSQILEETVEDNQGPSLKDVIE 60
MTKISPEIE KMPMEAVPPVSADVSFIS+ FPKYKLGAD Q+ +ET EDNQGPSLK+VIE
Sbjct: 504 MTKISPEIETKMPMEAVPPVSADVSFISNSFPKYKLGADHQVFQETAEDNQGPSLKEVIE 683
Query: 61 QEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEA 96
QEA+NLS+Q+ RISVRDLA KFDKNL+AAAKLSNEA
Sbjct: 684 QEASNLSEQNNRISVRDLASKFDKNLSAAAKLSNEA 791
>BI267537 similar to PIR|F84664|F846 hypothetical protein At2g26770
[imported] - Arabidopsis thaliana, partial (17%)
Length = 499
Score = 130 bits (328), Expect = 9e-31
Identities = 71/120 (59%), Positives = 91/120 (75%)
Frame = +1
Query: 20 VSADVSFISDRFPKYKLGADSQILEETVEDNQGPSLKDVIEQEATNLSDQHKRISVRDLA 79
VSADV F S RFP Y++G++++I++ D + S+K+V+ +E L +Q KR+SVRDLA
Sbjct: 148 VSADVIFASLRFPTYQIGSNNKIMD----DPKVLSMKEVVARETAQLLEQQKRLSVRDLA 315
Query: 80 CKFDKNLAAAAKLSNEAKIREVASLEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMV 139
KF+K LAAAAKLS EAK+RE ASLE HVLLKKLRD LE L+ R GRNK+DV AIS+V
Sbjct: 316 SKFEKGLAAAAKLSEEAKLREAASLEKHVLLKKLRDGLESLKGRVTGRNKDDVHXAISLV 495
>TC84423 similar to PIR|F84730|F84730 probable myosin heavy chain [imported]
- Arabidopsis thaliana, partial (10%)
Length = 787
Score = 38.1 bits (87), Expect = 0.008
Identities = 62/248 (25%), Positives = 107/248 (43%), Gaps = 24/248 (9%)
Frame = +2
Query: 26 FISDRFPKYKLGADSQILEETVED--NQGPSLKDVIE--QEATNLSDQ--HKRISVRDLA 79
F+S + KL A+ + LEETVED K++ +E LSD+ K S+ A
Sbjct: 2 FVSVQEELTKLNAEKKGLEETVEDLTVNAKHFKELCSDLEEKLKLSDESFSKTDSLLSQA 181
Query: 80 CKFDKNLAAAAK----LSNEA-KIREVAS-----LEGHV-----LLKKLRDALEYLRVRF 124
+ L K L NE+ + AS LEGH+ ++ + L L RF
Sbjct: 182 LSNNSELEQKVKSLEDLHNESGAVAATASQRSLELEGHIEATNAAAEEAKSQLRELETRF 361
Query: 125 AGRNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQASEDAKKLVN---QE 181
+++VE + + +K E ++ + ++ L LK+A E+ K L+N QE
Sbjct: 362 IAAEQKNVELE-QQLNLVQLKANDAERDVTEFSEKISHLDAKLKEAEEE-KNLLNSLLQE 535
Query: 182 KSFACAEIESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEVQEARRIKLLHQPSKVM 241
+++ES + + LEE+ + + + D + +R ++ L Q S
Sbjct: 536 HMDKLSQLESDLNQSTQKNSQLEEELKIVKEKCSEHEDRATMNNERSRELEDLIQSSHSK 715
Query: 242 AMEYELRA 249
+ + E RA
Sbjct: 716 SEKCEKRA 739
>AW687875 similar to PIR|T05990|T05 hypothetical protein F17M5.150 -
Arabidopsis thaliana, partial (16%)
Length = 578
Score = 33.5 bits (75), Expect = 0.19
Identities = 34/134 (25%), Positives = 60/134 (44%), Gaps = 11/134 (8%)
Frame = +2
Query: 129 KEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQASEDAKKL---VNQEKS-- 183
K D+EKA S V L V T EL QEK + + AS + +N+ KS
Sbjct: 2 KLDIEKATSEVNNLKVAATSLRSELEQEKSSLASIGQREGMASITVASIEVELNKTKSDI 181
Query: 184 -FACAEIESARSVVLRIGEALEEQEQDSQ-----ASKPQDVDGLVEEVQEARRIKLLHQP 237
F + + + ++L + + L+E +++ A + +V V+E E +
Sbjct: 182 AFVQMKEKEGKEMILELPKKLQEASEEANKANLLAREACEVFRRVKEEAEQAKAGASTMH 361
Query: 238 SKVMAMEYELRALR 251
S+++A + E+ A R
Sbjct: 362 SRLLAAQKEIEAAR 403
>TC89984 similar to GP|16604601|gb|AAL24093.1 unknown protein {Arabidopsis
thaliana}, partial (32%)
Length = 1303
Score = 33.1 bits (74), Expect = 0.25
Identities = 30/108 (27%), Positives = 52/108 (47%), Gaps = 16/108 (14%)
Frame = +3
Query: 84 KNLAAAAKLSNEAK-------IREVASLEGHVLLKKLRDALEYLRV------RFAGRNKE 130
K+L KL +E+ +E + VLL+ L A E L + ++K
Sbjct: 543 KSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKA 722
Query: 131 DVEKAISMVEALAVKLTQNEGEL---IQEKFEVKKLLNFLKQASEDAK 175
D++ + V++L T+ + EL I+EK+E +KLL ++ SE A+
Sbjct: 723 DIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAE 866
>TC78343 weakly similar to GP|15724195|gb|AAL06489.1 AT5g49210/K21P3_8
{Arabidopsis thaliana}, partial (78%)
Length = 903
Score = 32.7 bits (73), Expect = 0.32
Identities = 39/137 (28%), Positives = 62/137 (44%)
Frame = +3
Query: 111 KKLRDALEYLRVRFAGRNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQA 170
K+ L LR +A + KE ++ I VEA+ ++ Q K E +K L+ A
Sbjct: 207 KEAEAQLAQLRREYAKQVKEVRKEYIVEVEAMRLEK--------QRKDEARK--EALRVA 356
Query: 171 SEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEVQEARR 230
+E+ KKL QEK E A+ L+E+ + Q+ + Q E ++A +
Sbjct: 357 NEERKKLKAQEKELRAQERNIAQQQFRE--TLLKERAEKLQSWRMQ---AKKHEEKKAEK 521
Query: 231 IKLLHQPSKVMAMEYEL 247
LLH+ S + E EL
Sbjct: 522 KDLLHKRSSLWVDEAEL 572
>TC85473 homologue to SP|Q9XGM1|VATD_ARATH Vacuolar ATP synthase subunit D
(EC 3.6.3.14) (V-ATPase D subunit) (Vacuolar proton pump
D subunit)., partial (80%)
Length = 1267
Score = 32.7 bits (73), Expect = 0.32
Identities = 40/176 (22%), Positives = 75/176 (41%), Gaps = 5/176 (2%)
Frame = +3
Query: 85 NLAAAAKLSNEAKIREVASLEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAV 144
N+ + K R V + GH LLKK DA L V+F ++ ++K +S E++
Sbjct: 123 NVVPTVTMLGVVKARLVGATRGHALLKKKSDA---LTVQF----RQILKKIVSTKESMGD 281
Query: 145 KLTQNEGELIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALE 204
+ + L + K+ A ++ K +V + A + S V +
Sbjct: 282 IMKTSSFALTEAKY----------VAGDNIKHVVLENVKEASLRVRSRTENVAGVKLPKF 431
Query: 205 EQEQDSQASKPQDVDGLV---EEVQEAR--RIKLLHQPSKVMAMEYELRALRDQIR 255
+ D +A+K D+ GL ++VQ+ R IK + ++ +++ L D I+
Sbjct: 432 DYSADGEATK-NDLTGLARGGQQVQQCRVAYIKAIEVLVELASLQTSFLTLDDAIK 596
>TC82190 weakly similar to GP|13937297|gb|AAK50128.1 unknown protein {Oryza
sativa}, partial (22%)
Length = 1001
Score = 32.3 bits (72), Expect = 0.42
Identities = 16/47 (34%), Positives = 24/47 (51%)
Frame = +2
Query: 304 PQVSKCSFQWYRLSSEGSWREVISGADKSIYAPDPFDVGRILQVDIV 350
P S C FQW R +G+ R+ I GA Y DV +++ V+ +
Sbjct: 8 PSTSLCMFQWVRHLQDGT-RQYIEGATNPEYVVTADDVDKLIAVECI 145
>TC86719 similar to GP|599956|emb|CAA83434.1| chloroplast inner membrane
protein {Pisum sativum}, partial (63%)
Length = 2950
Score = 30.4 bits (67), Expect = 1.6
Identities = 42/220 (19%), Positives = 79/220 (35%), Gaps = 19/220 (8%)
Frame = +3
Query: 4 ISPEIEIKMPMEAVPPVSADVSFI--------------SDRFPKYKLGADSQILEETVED 49
+ P+ + M +PP VS + S + PK L + LEE +E
Sbjct: 1512 VDPKKKRNMKKSDIPPAKISVSEVEVQIEKLKQQILKSSSQPPKLNLDKTIKKLEEKLEK 1691
Query: 50 NQGPSLKDV-----IEQEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKIREVASL 104
L + + Q + L D+ + S D D L + R +++
Sbjct: 1692 EVDQELSEAAKALGLTQSLSKLRDEFSKASSDDQP--LDPLLKGKIEKLQADFNRRLSAA 1865
Query: 105 EGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKLL 164
LKK D L +++ + RNKE I+ A + + + + + +K+
Sbjct: 1866 PNANKLKKKHDELTKVKLLLSDRNKEVASSKINKEAATLTQELKKKFDDVMNNPRIKENY 2045
Query: 165 NFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALE 204
L+ + A + + IE V L++ A++
Sbjct: 2046 EALQSEIQRASSASDLDDELKKKIIEFNNEVDLQVANAVK 2165
>TC93096
Length = 686
Score = 28.9 bits (63), Expect = 4.7
Identities = 26/93 (27%), Positives = 39/93 (40%), Gaps = 7/93 (7%)
Frame = +1
Query: 30 RFPKYKLGADSQILEETVEDNQGPSLKDVIEQEATNLSDQ-------HKRISVRDLACKF 82
R +YK Q + V D + SLK +E +N Q H++ V +
Sbjct: 292 RHARYKKKTSLQDQQNAVTDAE--SLKVSQHREISNAKGQASSGNALHQKKDVNTTTDQG 465
Query: 83 DKNLAAAAKLSNEAKIREVASLEGHVLLKKLRD 115
N + K N + EV ++E + LLK LRD
Sbjct: 466 SLNRDISIKFKNNDDVLEVINVEKNALLKFLRD 564
>TC84435 similar to GP|20259307|gb|AAM14389.1 putative nodulin protein
{Arabidopsis thaliana}, partial (41%)
Length = 618
Score = 28.5 bits (62), Expect = 6.1
Identities = 14/46 (30%), Positives = 23/46 (49%), Gaps = 8/46 (17%)
Frame = +1
Query: 382 TDFHVVISQMNGKDH--------SSRSTHSFNVGRMRIKLCRGWIT 419
+ FH++IS DH SS +H + ++++ C GWIT
Sbjct: 184 SSFHLLISWKRRTDHRLI*TSSASSFVSHLLGLQQIKVSTCLGWIT 321
>BQ137051 homologue to GP|20259307|gb putative nodulin protein {Arabidopsis
thaliana}, partial (28%)
Length = 696
Score = 28.5 bits (62), Expect = 6.1
Identities = 14/46 (30%), Positives = 23/46 (49%), Gaps = 8/46 (17%)
Frame = +3
Query: 382 TDFHVVISQMNGKDH--------SSRSTHSFNVGRMRIKLCRGWIT 419
+ FH++IS DH SS +H + ++++ C GWIT
Sbjct: 246 SSFHLLISWKRRTDHRLI*TSSASSFVSHLLGLQQIKVSTCLGWIT 383
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.131 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,786,242
Number of Sequences: 36976
Number of extensions: 138232
Number of successful extensions: 659
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 653
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 656
length of query: 490
length of database: 9,014,727
effective HSP length: 100
effective length of query: 390
effective length of database: 5,317,127
effective search space: 2073679530
effective search space used: 2073679530
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0269a.3


