

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
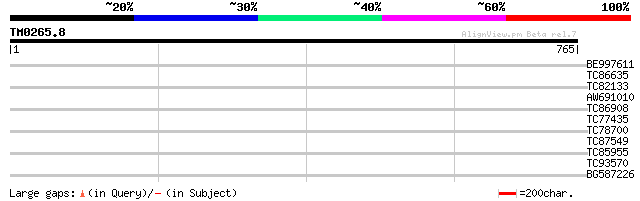
Query= TM0265.8
(765 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BE997611 39 0.007
TC86635 weakly similar to GP|9758600|dbj|BAB09233.1 gene_id:MDF2... 32 1.2
TC82133 similar to PIR|G84597|G84597 probable XAP-5 protein [Hom... 32 1.2
AW691010 similar to GP|8656003|gb| Contains similarity to formin... 31 2.0
TC86908 similar to PIR|G96708|G96708 hypothetical protein T26J14... 30 2.6
TC77435 similar to GP|7211427|gb|AAF40306.1| RNA helicase {Vigna... 30 4.5
TC78700 similar to GP|21554835|gb|AAM63703.1 unknown {Arabidopsi... 29 5.9
TC87549 similar to PIR|T01734|T01734 hypothetical protein A_IG00... 29 5.9
TC85955 similar to GP|3860319|emb|CAA10127.1 nucleolar protein {... 29 7.7
TC93570 similar to GP|21553436|gb|AAM62529.1 unknown {Arabidopsi... 29 7.7
BG587226 similar to GP|10176955|d phosphatidylinositol 4-kinase ... 29 7.7
>BE997611
Length = 547
Score = 38.9 bits (89), Expect = 0.007
Identities = 33/130 (25%), Positives = 57/130 (43%)
Frame = -2
Query: 436 NVNDEFVGDIEPLGENKKKQKEKEEEKRKVELEKKFTKVPFPPFPTNIAKRRLEKQFSKF 495
N+ DE + +E + E+K + ++ EEK PP +F+
Sbjct: 516 NMEDEHINAVE-VAEDKPQTEDA*EEKE------------VPP------------KFNAL 412
Query: 496 ISMFKKLRVELPFSEVLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKLPP 555
+ K +V +P +E +E++P KF++E+L LSEE E + L+ E +
Sbjct: 411 HHILKGYKVNMPITETVEQIPCCIKFLQELLKTNANLSEE-EFISLSSEFHHTYEVPAVV 235
Query: 556 KRKDPGSFTL 565
+ G FTL
Sbjct: 234 RFDGEGCFTL 205
>TC86635 weakly similar to GP|9758600|dbj|BAB09233.1
gene_id:MDF20.10~pir||T08929~similar to unknown protein
{Arabidopsis thaliana}, partial (32%)
Length = 1963
Score = 31.6 bits (70), Expect = 1.2
Identities = 20/76 (26%), Positives = 35/76 (45%)
Frame = +2
Query: 426 EPKQKIVENNNVNDEFVGDIEPLGENKKKQKEKEEEKRKVELEKKFTKVPFPPFPTNIAK 485
E + ++ + D+ G G K+ + + EEK KV+ KK K+P P PT+
Sbjct: 149 EADEDTIDKSKEEDKAEGSKGEKGSKKRARGKVNEEKVKVK--KKELKLPEPKTPTSDRP 322
Query: 486 RRLEKQFSKFISMFKK 501
R K + +++ K
Sbjct: 323 VRERKSVERLVALIDK 370
>TC82133 similar to PIR|G84597|G84597 probable XAP-5 protein [Homo sapiens]
[imported] - Arabidopsis thaliana, partial (60%)
Length = 926
Score = 31.6 bits (70), Expect = 1.2
Identities = 16/48 (33%), Positives = 30/48 (62%), Gaps = 1/48 (2%)
Frame = +3
Query: 424 LSEPKQKIVENNNVNDEFVGDIEPLGENKKKQKEKEEEKRKV-ELEKK 470
LSE K++ E + + D +VG + ++ +K++E E+RK+ EL+ K
Sbjct: 135 LSEEKKRAREMSGMGDGYVGTAQDAVRIRRLEKQREAERRKIQELKSK 278
>AW691010 similar to GP|8656003|gb| Contains similarity to formin binding
protein 11 from Mus musculus gb|AF135439 and contains,
partial (18%)
Length = 655
Score = 30.8 bits (68), Expect = 2.0
Identities = 29/123 (23%), Positives = 54/123 (43%), Gaps = 11/123 (8%)
Frame = +3
Query: 437 VNDEFVGDIEPLGENKKKQKEKEEEKRKVELEKKFTK-----VPFPPFPTNIAKRRLEKQ 491
+ND+ G ++ LGE K+ E +++K E E+K K F + +
Sbjct: 75 INDKRYGALKSLGERKQAFNEYLSQRKKQEAEEKRMKHKKAREDFRKMLEESTELTSSIR 254
Query: 492 FSKFISMFKKLRVELPFSEVLEKMPQYAKFIKEILSKKR-RLSEEN-----EIVELTEEC 545
+SK I++F+ ++ F++E+L+K+R ++ EE E + E C
Sbjct: 255 YSKAIAIFENDDRFKAVERERDRKDMIESFLEELLNKERAKVLEERKRNTVEYRKFLESC 434
Query: 546 SVI 548
I
Sbjct: 435 DFI 443
>TC86908 similar to PIR|G96708|G96708 hypothetical protein T26J14.4
[imported] - Arabidopsis thaliana, partial (65%)
Length = 1587
Score = 30.4 bits (67), Expect = 2.6
Identities = 16/56 (28%), Positives = 31/56 (54%), Gaps = 3/56 (5%)
Frame = +1
Query: 450 ENKKKQKEKEEEKRKVELEKKFTKVPFPP---FPTNIAKRRLEKQFSKFISMFKKL 502
+ KKK+K+K+++K+ L + + PP F N+ ++ + SKF +M K+
Sbjct: 88 KKKKKKKKKKKKKKTSRLVPFISYISLPPSLFFLLNVFQKVITNSTSKFSTMHNKI 255
>TC77435 similar to GP|7211427|gb|AAF40306.1| RNA helicase {Vigna radiata},
partial (67%)
Length = 1828
Score = 29.6 bits (65), Expect = 4.5
Identities = 23/91 (25%), Positives = 41/91 (44%), Gaps = 12/91 (13%)
Frame = +1
Query: 395 RKTPGMFPSDTVINPKENCSAITLRSGATLSEPKQKIVENNNVNDEFVGDIEPLGENKKK 454
R P F S T+ + +++ S + S K+K+++N+ V++ + +KK+
Sbjct: 82 RFIPSRFHSSTMPSLPITNDTVSMESPKSKSSKKKKLLQNDVVSEVEAVSAKKKESSKKR 261
Query: 455 QKEK------------EEEKRKVELEKKFTK 473
+K EE RKV+ EKK K
Sbjct: 262 KKSSDDDEETKSDISSEEGSRKVKKEKKKKK 354
>TC78700 similar to GP|21554835|gb|AAM63703.1 unknown {Arabidopsis
thaliana}, partial (5%)
Length = 687
Score = 29.3 bits (64), Expect = 5.9
Identities = 21/53 (39%), Positives = 29/53 (54%)
Frame = +2
Query: 419 RSGATLSEPKQKIVENNNVNDEFVGDIEPLGENKKKQKEKEEEKRKVELEKKF 471
+SG+ E K+K V N++ E VGD KKK+K K +E K EKK+
Sbjct: 428 KSGSGEEEEKEKEV---NLDGEGVGD-------KKKKKRKRDEVEKEWEEKKY 556
>TC87549 similar to PIR|T01734|T01734 hypothetical protein A_IG002N01.30 -
Arabidopsis thaliana (fragment), partial (38%)
Length = 1766
Score = 29.3 bits (64), Expect = 5.9
Identities = 50/216 (23%), Positives = 87/216 (40%), Gaps = 9/216 (4%)
Frame = +2
Query: 485 KRRLEKQFSKFISMFKKLRVELPFSEVLEKMPQYAKFIKEILSKKRRLSEENEIVELTEE 544
KRR E F I KKL+ L ++L P ++E+ +R L ++ +
Sbjct: 458 KRRKELPFDNVIQKDKKLKFVLKVRKILVSKPTRVMSLQELGKYRRELG-----LDKKRK 622
Query: 545 CSVILQRKLPPKRKDPGSFTLPVNFGASKQVRALCDLGSSVNLMPLSM---FERLNVGEL 601
VIL R+ PG F + V G + + L L + E + V +L
Sbjct: 623 LIVIL-------RRFPGVFEI-VEDGCYSLKFKMTSEAEKLYLEELRVRNEMEDVVVTKL 778
Query: 602 KPTMMMLQLADRSIVTPWGVVEDVLVRVGEF------EFPVDFVIIDMDEDSKIPLILGR 655
+ ++M+ L R ++ G + + L EF +P F ++ + + L
Sbjct: 779 R-KLLMMPLEKRILLEKIGHLANDLGLPREFRDTICHRYPEFFKVVQTERGPALELTHWD 955
Query: 656 PFLATSQAKINVGKGTISLRVADEKIGFNIFDLKPK 691
P LA S A+++ + I V ++ + I D PK
Sbjct: 956 PHLAVSAAELSAEENRIR-EVEEQNL---IIDRAPK 1051
>TC85955 similar to GP|3860319|emb|CAA10127.1 nucleolar protein {Cicer
arietinum}, partial (98%)
Length = 1981
Score = 28.9 bits (63), Expect = 7.7
Identities = 17/44 (38%), Positives = 24/44 (53%), Gaps = 1/44 (2%)
Frame = +2
Query: 655 RPFLATSQAKINVGK-GTISLRVADEKIGFNIFDLKPKPVEKND 697
R L T+ K+ GK SL VA+ KIG +I + PV+ N+
Sbjct: 311 RTVLETNLPKVKEGKKAKFSLGVAESKIGSHIHEATKIPVQSNE 442
>TC93570 similar to GP|21553436|gb|AAM62529.1 unknown {Arabidopsis
thaliana}, partial (57%)
Length = 759
Score = 28.9 bits (63), Expect = 7.7
Identities = 35/137 (25%), Positives = 57/137 (41%), Gaps = 8/137 (5%)
Frame = +1
Query: 450 ENKKKQKEKEEEKRKVELEKKFTKVPFPPFPTNIAKRRLEKQFSKFISMFKKLRVELPFS 509
+ KKK+ +++ FT FPT+I+ + + F K L ++ P S
Sbjct: 1 KKKKKKSSNQQKNMSAITTLTFTNTTNITFPTSISTKNPIFHTTHFT---KPLHLQNPLS 171
Query: 510 EVLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKLPPKRKDPGSFTLP--- 566
L K I I+ K + E+E+ + E+ + +KLP K K + +LP
Sbjct: 172 ISLRKP------ISTIVLCKGSVESESEVPFVPEDEWL---QKLPEKTKPLYTHSLPCIE 324
Query: 567 -----VNFGASKQVRAL 578
+ F SK RAL
Sbjct: 325 AWLKSLGFNQSKDDRAL 375
>BG587226 similar to GP|10176955|d phosphatidylinositol 4-kinase {Arabidopsis
thaliana}, partial (10%)
Length = 473
Score = 28.9 bits (63), Expect = 7.7
Identities = 18/47 (38%), Positives = 22/47 (46%)
Frame = +3
Query: 439 DEFVGDIEPLGENKKKQKEKEEEKRKVELEKKFTKVPFPPFPTNIAK 485
D + D E LG K+QKEK + EK TK P P P+ K
Sbjct: 201 DRSIEDSELLGS--KRQKEKHPGSPMPQSEKSSTKPPLPINPSQFRK 335
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.337 0.147 0.466
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,541,054
Number of Sequences: 36976
Number of extensions: 304092
Number of successful extensions: 2172
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 1220
Number of HSP's successfully gapped in prelim test: 100
Number of HSP's that attempted gapping in prelim test: 917
Number of HSP's gapped (non-prelim): 1374
length of query: 765
length of database: 9,014,727
effective HSP length: 104
effective length of query: 661
effective length of database: 5,169,223
effective search space: 3416856403
effective search space used: 3416856403
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0265.8


