

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
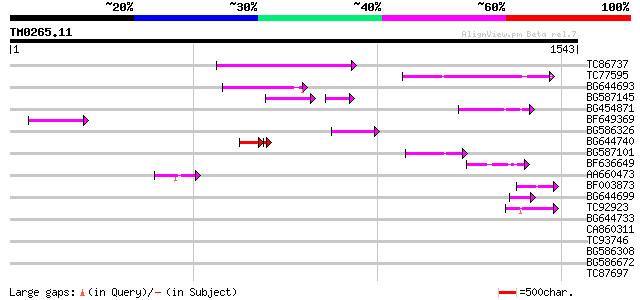
Query= TM0265.11
(1543 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 256 5e-68
TC77595 weakly similar to PIR|T18350|T18350 probable pol polypro... 181 3e-45
BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin... 172 1e-42
BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - ... 100 5e-28
BG454871 weakly similar to GP|10140673|g putative gag-pol polypr... 122 1e-27
BF649369 120 3e-27
BG586326 similar to PIR|G84493|G8 probable retroelement pol poly... 92 2e-18
BG644740 similar to PIR|A84460|A84 probable retroelement pol pol... 63 9e-14
BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana... 73 1e-12
BF636649 similar to GP|21628724|emb OSJNBa0033H08.7 {Oryza sativ... 70 5e-12
AA660473 65 3e-10
BF003873 similar to GP|14715222|em putative polyprotein {Cicer a... 55 2e-07
BG644699 similar to PIR|T07863|T078 probable polyprotein - pinea... 54 5e-07
TC92923 similar to GP|6466937|gb|AAF13073.1| putative retroeleme... 51 4e-06
BG644733 weakly similar to GP|15289942|db putative polyprotein {... 41 0.004
CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product ... 40 0.005
TC93746 weakly similar to GP|22830935|dbj|BAC15800. hypothetical... 38 0.035
BG586308 weakly similar to PIR|F84528|F8 probable retroelement p... 33 0.85
BG586672 33 0.85
TC87697 similar to PIR|T47802|T47802 hypothetical protein F24G16... 30 5.5
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 256 bits (654), Expect = 5e-68
Identities = 148/395 (37%), Positives = 223/395 (55%), Gaps = 16/395 (4%)
Frame = +1
Query: 564 VLADFHEVFT--KKIQLPPVRSRV-HQITLNPEHG----PINVRP-YRYPHHQKEEIERQ 615
VL +F ++F K Q+P R + H I L P+ P+ P Y + +++
Sbjct: 343 VLEEFPDLFNPEKAYQVPASRGLLDHAIPLIPDKDGNDPPLPWGPLYGMSRQELLVLKKT 522
Query: 616 VTELLEAGIIRPSMSSFSSPVILVKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDE 675
+ +LL+ G I+ S S+ +PV+ V+K R CVDYRALN T D+YP+P++ E L
Sbjct: 523 LEDLLDKGFIKASGSAAGAPVLFVRKPGGGIRFCVDYRALNAITKKDRYPLPLISETLRR 702
Query: 676 LNGAVIFSKIDLKSGYHQIRVHEEDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMND 735
+ GA F+K+D+ + +H++R+ +ED KT FRT G +E++V PFGL APATFQ +N
Sbjct: 703 VAGARWFTKLDVVAAFHKMRIKDEDQEKTAFRTRYGLFEWIVCPFGLTGAPATFQRYINK 882
Query: 736 IFRPYLRKFVLVFFDDILIYSK-SLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDY 794
+L FV + DD+LIY+ S +H ++ VL L + KC+F + Y
Sbjct: 883 TLHEFLDDFVTAYIDDVLIYTTGSKKDHEAQVRRVLRRLADAGLSLDPKKCEFSVTTVKY 1062
Query: 795 LGHII-SGAGVSVDPEKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTEL 853
+G I+ +G GVS DP K+ I +W P +VKG R FLG YY+ FI Y ++ +PLT L
Sbjct: 1063VGFILTAGKGVSCDPLKLAAIRDWLPPGSVKGARSFLGFCNYYKDFIPGYSEITEPLTRL 1242
Query: 854 TKKD-NFTWGEEAKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQ--- 909
T+KD F WG E + AF KLK + PVL + D + VE D +G +G VL Q+
Sbjct: 1243TRKDFPFRWGAEQEAAFTKLKRLFAEEPVLRMFDPEAVTTVETDCSGFALGGVLTQEDGT 1422
Query: 910 --RQPIAYFSKALSNGNLTKSVYEKELMALVLCIQ 942
P+A+ S+ LS +++KEL+A+ C++
Sbjct: 1423GAAHPVAFHSQRLSPAEYNYPIHDKELLAVWACLR 1527
>TC77595 weakly similar to PIR|T18350|T18350 probable pol polyprotein - rice
blast fungus gypsy retroelement (fragment), partial (14%)
Length = 1708
Score = 181 bits (458), Expect = 3e-45
Identities = 133/431 (30%), Positives = 202/431 (46%), Gaps = 18/431 (4%)
Frame = +2
Query: 1070 SPTSPSIPWLLQEYHGSPTGGHSGFLRTYRRLAESLYWVGMQRTVRDFVRACDTCQRQKY 1129
SP + L+QE H S GH G T ++ +W G +TVR FVR CD C
Sbjct: 152 SPLNELRTKLVQESHDSTAAGHPGRNGTLEIVSRKFFWPGQSQTVRRFVRNCDVCGGIHI 331
Query: 1130 SAMSPGGLLQPFPVPNAVWEDLSLDFIIGLP--KSKGYEAILVVVDRLSKYSHFILLKHP 1187
+ G L+P PVPN + DLS+DFI LP + +G + + V+VDRLSK + L+
Sbjct: 332 WRQAKRGFLKPLPVPNRLHSDLSMDFITSLPPTRGRGSQYLWVIVDRLSKS---VTLEEM 502
Query: 1188 YT--AKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFWSELFKLQGTKLKMSSSYHPET 1245
T A++ A+ F+ R HG+P ++ SDR +V FW E +L G +S+SYHP+T
Sbjct: 503 DTMEAEACAQRFLSCHYRFHGMPQSIVSDRGSNWVGRFWREFCRLTGVTQLLSTSYHPQT 682
Query: 1246 DGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNSTFHCSIGQTPFEVVYGRKAPP 1305
DG TE N+ +++ LR + W +P + + + SIG TPF V +G P
Sbjct: 683 DGGTERWNQEIQAVLRAYVCWSQDNWGDLLPTVQLALRNRHNSSIGATPFFVEHGYHVDP 862
Query: 1306 IIRFLTNETKV----VAVALELSERDEALKQLRTHLQRAQEQMASYANKKRRDLS-FDIG 1360
I V A L + + ++ + AQ++ + ANK+R + +G
Sbjct: 863 IPTVEDTGGVVSEGEAAAQLLVKRMKDVTGFIQAEIVAAQQRSEASANKRRCPADRYQVG 1042
Query: 1361 EWVFLKLRPHRQHSVVKRI----NQKLAARFYGPFQIEDKIGTVAYKLKLPVESRIHPVF 1416
+ V+L + ++ K++ ++ RF P +E L V ++P F
Sbjct: 1043 DKVWLNVSNYKSPRPSKKLDWLHHKYEVTRFVTPHVVE-----------LNVPGTVYPKF 1189
Query: 1417 HVSLLKRAI-----GNYQVQGQLPTDLGIEEATDIYPEAILGTRIIRQGDSEVHQSLIKW 1471
HV LL+RA G V Q P + + + E IL R + G Q+L+KW
Sbjct: 1190 HVDLLRRAASDPLPGQEVVDPQPPPIVDDDGEVEWEVEEILAARWHQVGRGRRRQALVKW 1369
Query: 1472 KNRSIDDVTWE 1482
K D TWE
Sbjct: 1370 K--GFVDATWE 1396
>BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin cluster;
Ty3-Gypsy type {Oryza sativa}, partial (15%)
Length = 716
Score = 172 bits (435), Expect = 1e-42
Identities = 102/243 (41%), Positives = 134/243 (54%), Gaps = 10/243 (4%)
Frame = +2
Query: 578 LPPVRSRVHQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLEAGIIRPSMSSFSSPVI 637
+PP I L P PI + YR + + ++ Q+ +LLE G I+PS+ V+
Sbjct: 17 VPPEWKIDFGIDLLPNMNPI*IPSYRINPLKLKVLKLQLKDLLEKGFIQPSIYP*GVVVL 196
Query: 638 LVKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIFSKIDLKSGYHQIRVH 697
+KKKD RM +DY LN I KYP+P++DEL D L G+ F KIDL+ G HQ RV
Sbjct: 197 FLKKKDGFLRMSIDYPQLNNVNIKIKYPLPLIDELFDNLQGSKWFFKIDLRLG*HQHRVI 376
Query: 698 EEDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLVFFDDILIYSK 757
ED+PKT FR GHYE LVM FG N P F +MN +F+ YL V+VF +DILIYSK
Sbjct: 377 GEDVPKTAFRIRYGHYEILVMSFG*TNPPMAFMELMNRVFQDYLDSLVIVFSNDILIYSK 556
Query: 758 SLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYL----------GHIISGAGVSVD 807
+ EH +HL+L L VL G QI Y+ H+ISG G+ VD
Sbjct: 557 NENEHENHLRLALKVLKD-----------IGLCQISYV*ILVEVGFFSLHVISGEGLKVD 703
Query: 808 PEK 810
++
Sbjct: 704 SKR 712
>BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (13%)
Length = 763
Score = 100 bits (250), Expect(2) = 5e-28
Identities = 50/136 (36%), Positives = 79/136 (57%)
Frame = +2
Query: 696 VHEEDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLVFFDDILIY 755
+H +D+ KT F T G Y Y VMPFGL NA +T+Q ++N +F L + V+ DD+L+
Sbjct: 2 MHPDDLEKTAFITDRGTYCYKVMPFGLKNAGSTYQRLVNRMFADKLGNTMEVYIDDMLVK 181
Query: 756 SKSLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGVSVDPEKVQCIV 815
S +H +HLK L N +KC FG ++LG+I++ G+ V+P+++ I+
Sbjct: 182 SLRATDHLNHLKE*FKTLDEYIMKLNPAKCTFGVTSGEFLGYIVTQQGIEVNPKQITAIL 361
Query: 816 EWPEPKNVKGVRGFLG 831
+ P PKN + V+ G
Sbjct: 362 DLPSPKNSREVQRLTG 409
Score = 43.5 bits (101), Expect(2) = 5e-28
Identities = 22/84 (26%), Positives = 45/84 (53%), Gaps = 4/84 (4%)
Frame = +3
Query: 859 FTWGEEAKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVLMQ----QRQPIA 914
F W E+ ++AF +LK +T+ PVL+ P+ + + + + +VL++ +++PI
Sbjct: 495 FVWDEKCEEAFEQLKQYLTTPPVLSKPEAGDTLSLYIAISSTAVSSVLIREDRGEQKPIF 674
Query: 915 YFSKALSNGNLTKSVYEKELMALV 938
Y SK +++ EK A++
Sbjct: 675 YTSKRMTDPETRYPTLEKMAFAVI 746
>BG454871 weakly similar to GP|10140673|g putative gag-pol polyprotein {Oryza
sativa (japonica cultivar-group)}, partial (7%)
Length = 674
Score = 122 bits (306), Expect = 1e-27
Identities = 72/207 (34%), Positives = 111/207 (52%)
Frame = +2
Query: 1221 SHFWSELFKLQGTKLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEF 1280
S+FW +LFKL GT L MSS+YHP +DGQ+E +N+ E YLRC P WS PWAE+
Sbjct: 32 SNFWKQLFKLHGTILTMSSAYHP*SDGQSEALNKGXEMYLRCLMFTDPLKWSKAFPWAEY 211
Query: 1281 WYNSTFHCSIGQTPFEVVYGRKAPPIIRFLTNETKVVAVALELSERDEALKQLRTHLQRA 1340
WYN++++ S TPF+ +YGR +IR + + +L++R+E L QL++ R
Sbjct: 212 WYNTSYNISAAMTPFKALYGRDLSMLIRSKGSSKDTADLQSQLAQREELLSQLQSISTRL 391
Query: 1341 QEQMASYANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTV 1400
+ + K R+ + WV L + ++ + + + + +
Sbjct: 392 NKL*SI---KLIRNAAILSSSWVSTSL*NCNLINSLR*HCGNIKSSVHP--TLVHY*QSA 556
Query: 1401 AYKLKLPVESRIHPVFHVSLLKRAIGN 1427
AYKL LP +++ P+FHVS LK GN
Sbjct: 557 AYKLSLPSTAKVPPIFHVSQLKPFHGN 637
>BF649369
Length = 631
Score = 120 bits (302), Expect = 3e-27
Identities = 70/163 (42%), Positives = 95/163 (57%)
Frame = +2
Query: 51 *IQCE*ITSRREESEAASVRRRRPSSLDHESGDLF*CARNPR*TSGEARTLEYGRINDPL 110
*I E +T +ESE +VRR S +DH DL CA R* GE EYGR DPL
Sbjct: 86 *IITERVTFGWKESEITTVRRG*SSCVDHTC*DLLRCAEYSR*HEGEIIPFEYGRSYDPL 265
Query: 111 VQSVDGNRG*TFLGKTQASVDRTLWRTTTRKPIRRTVDAEAARKCGGICGGFRVAVFSGR 170
VQ +GNRG + G+ + VDR+L R+ T + IRR ++ + K +C FRVA+ +G
Sbjct: 266 VQLTNGNRGRSIQGEIEEGVDRSLRRSKTGESIRRIINTSSNWKR*RVCRSFRVAIVAGW 445
Query: 171 KVTRRSVFGVFHEWVETTDPATGANVESQYPYGDDENRQGC*G 213
KVT ++FG+ HEW+E++ T + ES DDENR+ C G
Sbjct: 446 KVTGGAIFGLLHEWIESSHQKTSSYAESYDADADDENRKRCGG 574
>BG586326 similar to PIR|G84493|G8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 736
Score = 91.7 bits (226), Expect = 2e-18
Identities = 49/130 (37%), Positives = 74/130 (56%)
Frame = +2
Query: 877 TSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNGNLTKSVYEKELMA 936
TS+P+L LP+ ++ V DA+ G+G VL Q + IAY S+ L ++ E+ A
Sbjct: 8 TSAPILVLPELI-TYVVYTDASITGLGCVLTQHEKVIAYASRQLRKHEGNYPTHDLEMAA 184
Query: 937 LVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLLGYQFEVKYKPGQE 996
+V ++ WR YL G ++TDHKSLK+ Q + Q+ W+ + Y ++ Y PG+
Sbjct: 185 VVFALKIWRSYLYGAKVQIHTDHKSLKYIFTQPELNLRQRRWMEFVADYDLDITYYPGKA 364
Query: 997 NKAADALSRR 1006
N ADALSRR
Sbjct: 365 NLVADALSRR 394
>BG644740 similar to PIR|A84460|A84 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 754
Score = 62.8 bits (151), Expect(2) = 9e-14
Identities = 28/65 (43%), Positives = 42/65 (64%)
Frame = -1
Query: 626 RPSMSSFSSPVILVKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIFSKI 685
+PS+S + ++ V+KKD +RMC+DYR NK T +KYP+P +D L D++ F I
Sbjct: 262 QPSISP*GAALLFVRKKDGYFRMCIDYRQFNKVTTKNKYPLPRIDNLFDKIQEDCYF*NI 83
Query: 686 DLKSG 690
DL+ G
Sbjct: 82 DLRLG 68
Score = 33.5 bits (75), Expect(2) = 9e-14
Identities = 14/23 (60%), Positives = 17/23 (73%)
Frame = -3
Query: 690 GYHQIRVHEEDIPKTTFRTHNGH 712
GYHQ RV+E +IPKTT T G+
Sbjct: 71 GYHQFRVNEVNIPKTTLLTWRGY 3
>BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana}, partial
(10%)
Length = 624
Score = 72.8 bits (177), Expect = 1e-12
Identities = 49/170 (28%), Positives = 82/170 (47%), Gaps = 1/170 (0%)
Frame = +2
Query: 1076 IPWLLQEYHGSPTGGHSGFLRTYRRLAES-LYWVGMQRTVRDFVRACDTCQRQKYSAMSP 1134
IP +L HGS GH +T ++ ++ +W M + F+ CD CQRQ +
Sbjct: 95 IPGILFHCHGSNYAGHFAVSKTVSKIQQAGFWWPTMFKDAHSFISKCDPCQRQG-NIS*R 271
Query: 1135 GGLLQPFPVPNAVWEDLSLDFIIGLPKSKGYEAILVVVDRLSKYSHFILLKHPYTAKSIA 1194
+ Q F + V++ +DF+ P S + ILV VD +SK+ I A +
Sbjct: 272 NEMPQNFILEVEVFDVWGIDFMGPFPSSYNNKYILVAVDYVSKWVEAIA-SPTNDATVVV 448
Query: 1195 EVFVREIVRLHGIPNTVKSDRDPIFVSHFWSELFKLQGTKLKMSSSYHPE 1244
++F I G+P V SD F++ + +L K G + K++++YHP+
Sbjct: 449 KMFKSVIFPRFGVPRVVISDGGSHFINKVFEKLLKKNGVRHKVATAYHPQ 598
>BF636649 similar to GP|21628724|emb OSJNBa0033H08.7 {Oryza sativa}, partial
(4%)
Length = 653
Score = 70.5 bits (171), Expect = 5e-12
Identities = 56/179 (31%), Positives = 89/179 (49%), Gaps = 9/179 (5%)
Frame = +1
Query: 1244 ETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNSTFHCSIGQTPFEVVYGRKA 1303
++D Q ++N LE++L F S+ +F++ WAE YN+ FH + G TPF+VVY
Sbjct: 4 DSDEQAGLLNHTLETHLLYFTSEQQGV*NFFLTWAECLYNTNFHRTAGCTPFKVVY---- 171
Query: 1304 PPIIRFLTNETKVVAVALELSERDEALKQLRT-----HLQRAQEQMASYANK----KRRD 1354
+ + VA +L R+E L + T RA E ++ A + RR
Sbjct: 172 ------VVAHLQKFVVARDLIYRNEGLHKSST*TSFGRGTRAYEALSRPAYETC*HPRRP 333
Query: 1355 LSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKLPVESRIH 1413
LS V+ + R + ++ K A YGP+Q+ +IG+VA+KL LP + +IH
Sbjct: 334 LSI-----VYTRDRTYEW-----QVLPKYVA*CYGPYQVIKQIGSVAFKL*LPEQHQIH 480
>AA660473
Length = 655
Score = 64.7 bits (156), Expect = 3e-10
Identities = 46/134 (34%), Positives = 69/134 (51%), Gaps = 8/134 (5%)
Frame = -2
Query: 394 TMKLAGQVANVELLILIDSGASHNFISPKITSALGLVITPITTRSIKLGDGHKVV----- 448
T++L G + V ++IL+D GA+HNF S +G I ++ ++G + +
Sbjct: 543 TLQLVGCLKGVPIVILVDIGANHNFDFASSVSDVGCKGRVILSQESEVG*WSENINVG*A 364
Query: 449 --TRGICRGIKAKLGSIEITIDALVLELGGLDLVLGVSWLSTLGK-VIMDWKLLTMQFVY 505
+RG RG +DA LELG D+ LGV+WL LG V+ DW T+ F +
Sbjct: 363 Y*SRGTVRGFHC*------GVDAYELELGEFDMFLGVAWLEKLGN*VVFDWDERTICFEW 202
Query: 506 GEQVVQLQGMRGKD 519
+VV+LQG KD
Sbjct: 201 KGEVVKLQGQILKD 160
>BF003873 similar to GP|14715222|em putative polyprotein {Cicer arietinum},
partial (82%)
Length = 559
Score = 55.5 bits (132), Expect = 2e-07
Identities = 33/119 (27%), Positives = 61/119 (50%), Gaps = 5/119 (4%)
Frame = +2
Query: 1380 NQKLAARFYGPFQIEDKIGTVAYKLKLPVE-SRIHPVFHVSLLKRAI--GNYQVQGQLPT 1436
++KL RF GP+QI +++GTVAY++ LP +H VFHVS L++ + ++ +Q
Sbjct: 23 SKKLTVRFIGPYQISERVGTVAYRVGLPPHLLNLHDVFHVSQLRKYVPDPSHVIQSD--- 193
Query: 1437 DLGIEE--ATDIYPEAILGTRIIRQGDSEVHQSLIKWKNRSIDDVTWEDNEVLAGQFPE 1493
D+ + + + P I ++ E+ + W + + +TWE + +PE
Sbjct: 194 DVQVRDNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWDRANGESLTWELESKMVESYPE 370
>BG644699 similar to PIR|T07863|T078 probable polyprotein - pineapple
retrotransposon dea1 (fragment), partial (5%)
Length = 231
Score = 53.9 bits (128), Expect = 5e-07
Identities = 32/74 (43%), Positives = 44/74 (59%), Gaps = 3/74 (4%)
Frame = +2
Query: 1361 EWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKLPVE-SRIHPVFHVS 1419
E V LK+ P + KL+ R+ GPF++ +IG VAY+L LP S +HPVFHVS
Sbjct: 2 EQVLLKVLPTERGDCRFGKRGKLSLRYIGPFEVIKRIGEVAYELALPPGLSGVHPVFHVS 181
Query: 1420 LLKR--AIGNYQVQ 1431
+ KR GNY ++
Sbjct: 182 MFKRYHGDGNYIIR 223
>TC92923 similar to GP|6466937|gb|AAF13073.1| putative retroelement pol
polyprotein {Arabidopsis thaliana}, partial (1%)
Length = 625
Score = 50.8 bits (120), Expect = 4e-06
Identities = 43/155 (27%), Positives = 74/155 (47%), Gaps = 12/155 (7%)
Frame = +2
Query: 1350 KKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAARF---------YGPFQIEDKIGTV 1400
KK D++ VFLK RP+ K+ + KL YG + D V
Sbjct: 20 KKFIDVTIICSWRVFLKFRPYLTPLYGKKNDPKLLVLLEIQAILTPLYG--KKNDPKLLV 193
Query: 1401 AYKLKLPVESRIHPVFHV---SLLKRAIGNYQVQGQLPTDLGIEEATDIYPEAILGTRII 1457
+ L + + + F++ S K ++G+Y V+ QLP + +E + P+ +L TR I
Sbjct: 194 LFNL*IAAHTNFYDPFNIPCFSA*KSSVGSYPVELQLPEGMEVEINDEADPQFMLATRQI 373
Query: 1458 RQGDSEVHQSLIKWKNRSIDDVTWEDNEVLAGQFP 1492
R+GD + IKWK ++++ + +D+ + QFP
Sbjct: 374 REGDITT**A-IKWKAKTLEHASLKDDFTIRSQFP 475
>BG644733 weakly similar to GP|15289942|db putative polyprotein {Oryza sativa
(japonica cultivar-group)}, partial (1%)
Length = 174
Score = 40.8 bits (94), Expect = 0.004
Identities = 22/43 (51%), Positives = 29/43 (67%)
Frame = -3
Query: 1163 KGYEAILVVVDRLSKYSHFILLKHPYTAKSIAEVFVREIVRLH 1205
K YE+I VVVDRL+K + FI K Y+AK A + + EIV +H
Sbjct: 130 KSYESI*VVVDRLTKSTLFIPFKTSYSAK*YARILLDEIVCIH 2
>CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product {Drosophila
melanogaster}, partial (20%)
Length = 192
Score = 40.4 bits (93), Expect = 0.005
Identities = 23/58 (39%), Positives = 34/58 (57%), Gaps = 5/58 (8%)
Frame = +1
Query: 901 GIGAVLMQQRQ-----PIAYFSKALSNGNLTKSVYEKELMALVLCIQHWRHYLLGKAF 953
GIGA L Q+ + PIAY S+ L+ +V E+E +A + I+++RHYL G F
Sbjct: 13 GIGAGLSQKDEENHEHPIAYASRLLTAAERNYTVVERECLAAIWAIRNFRHYLHGPKF 186
>TC93746 weakly similar to GP|22830935|dbj|BAC15800. hypothetical
protein~similar to gag-pol polyprotein {Oryza sativa
(japonica cultivar-group)}, partial (4%)
Length = 1019
Score = 37.7 bits (86), Expect = 0.035
Identities = 18/43 (41%), Positives = 30/43 (68%), Gaps = 1/43 (2%)
Frame = +2
Query: 1142 PVPNAVWEDLSLDFIIGLPKSKGY-EAILVVVDRLSKYSHFIL 1183
PVP WED+++DF +GL ++ ++ +VV D+ S+ +HFIL
Sbjct: 473 PVPKPPWEDVTIDFSLGLL*TQQLKDSKMVVGDKFSRMAHFIL 601
>BG586308 weakly similar to PIR|F84528|F8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 686
Score = 33.1 bits (74), Expect = 0.85
Identities = 14/52 (26%), Positives = 29/52 (54%)
Frame = -2
Query: 1205 HGIPNTVKSDRDPIFVSHFWSELFKLQGTKLKMSSSYHPETDGQTEVINRCL 1256
HG+P + +D F+S+ + E + +L +S +P+++GQ E N+ +
Sbjct: 685 HGLPYEIVTDNGSHFISNKFREFCERWRIRLNTASPRYPQSNGQAEASNKII 530
>BG586672
Length = 667
Score = 33.1 bits (74), Expect = 0.85
Identities = 14/28 (50%), Positives = 23/28 (82%), Gaps = 1/28 (3%)
Frame = +2
Query: 440 KLGDGHKVVTRGICRGIK-AKLGSIEIT 466
KLGDGH+V+++G+C+G + K GS+ +T
Sbjct: 23 KLGDGHEVLSQGVCKGREGTKEGSVFLT 106
>TC87697 similar to PIR|T47802|T47802 hypothetical protein F24G16.50 -
Arabidopsis thaliana, partial (7%)
Length = 1514
Score = 30.4 bits (67), Expect = 5.5
Identities = 19/47 (40%), Positives = 23/47 (48%)
Frame = +3
Query: 1443 ATDIYPEAILGTRIIRQGDSEVHQSLIKWKNRSIDDVTWEDNEVLAG 1489
A ++PEAIL + I G E SLIK N S+ EV AG
Sbjct: 495 APTVHPEAILSSTDITSGKIESVPSLIKGGNESLAATKASATEVFAG 635
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.140 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 47,795,275
Number of Sequences: 36976
Number of extensions: 690578
Number of successful extensions: 3480
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 3321
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3419
length of query: 1543
length of database: 9,014,727
effective HSP length: 109
effective length of query: 1434
effective length of database: 4,984,343
effective search space: 7147547862
effective search space used: 7147547862
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0265.11


