

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0263.7
(787 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
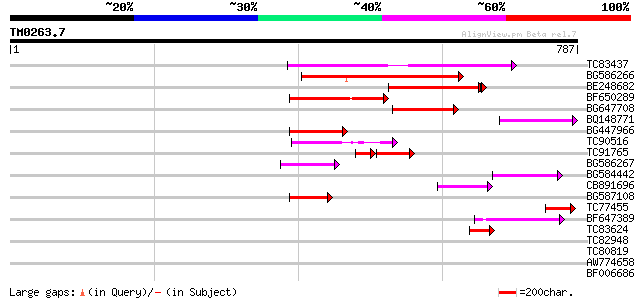
Score E
Sequences producing significant alignments: (bits) Value
TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imp... 194 8e-50
BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-li... 164 9e-41
BE248682 similar to GP|18568269|gb putative gag-pol polyprotein ... 130 2e-30
BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgari... 111 1e-24
BG647708 weakly similar to GP|13786450|gb| putative reverse tran... 71 2e-12
BQ148771 68 2e-11
BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [impo... 66 5e-11
TC90516 60 2e-09
TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative ret... 45 1e-08
BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At... 53 5e-07
BG584442 50 3e-06
CB891696 50 3e-06
BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - ... 50 4e-06
TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarot... 49 6e-06
BF647389 49 6e-06
TC83624 homologue to PIR|G84581|G84581 copia-like retroelement p... 44 2e-04
TC82948 40 0.003
TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non... 38 0.013
AW774658 similar to GP|2808681|emb| Hcr9-4B {Lycopersicon hirsut... 25 2.0
BF006686 31 2.1
>TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imported] -
Arabidopsis thaliana, partial (6%)
Length = 951
Score = 194 bits (494), Expect = 8e-50
Identities = 123/320 (38%), Positives = 183/320 (56%), Gaps = 2/320 (0%)
Frame = +2
Query: 386 RGGNASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFI 445
+G N++FI LIPKV+NPQ L ++RPISLVG +YKI+ KLL+ RLR V+ VI D Q+AF+
Sbjct: 44 KGINSTFIALIPKVDNPQRLNDFRPISLVGSLYKILGKLLANRLRVVIGSVISDAQSAFV 223
Query: 446 GGRFMLDSVVVANEVVEEAKRRKKECFVFKVDFEKAYDSVDWNFLFYMM-SRIGFSHKWI 504
R +L+ V + + R ++ F K ++ ++ + + F W
Sbjct: 224 KNRQILEMVFL*QMRLWMRLRN*RKIFCCLRWILKRLITLSIGLIWILF*VGMSFLVLWR 403
Query: 505 HWIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKL 564
WIK C+ +A++SVLVNGSPT N LM+ VQ +L
Sbjct: 404 KWIKECVSTATTSVLVNGSPT---------------------------NVLMKSLVQTQL 502
Query: 565 FTGYRVGG-DEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLA 623
FT Y G + V +S LQFA+DTL + NI +++ L F AMSGLKVNF KS L
Sbjct: 503 FTRYSFGVVNPVVVSHLQFANDTLLLETKNWANIRALRAALVIF*AMSGLKVNFHKSGLV 682
Query: 624 GISMVSRETQIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKS 683
+++ A++L+ KV VPF YLG+P+ N R+ + WEP++ +++ +L+ +
Sbjct: 683 CVNIAPSWLSEAASVLSWKVGKVPFLYLGMPIEGNSRRLSFWEPIVNRIKARLTGWNSRF 862
Query: 684 LSFGGRICLIKSVLSSLPLF 703
LSFGGR+ L+KSVL+SL ++
Sbjct: 863 LSFGGRLVLLKSVLTSLSVY 922
>BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 789
Score = 164 bits (416), Expect = 9e-41
Identities = 85/229 (37%), Positives = 138/229 (60%), Gaps = 3/229 (1%)
Frame = -3
Query: 405 LGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGRFMLDSVVVANEVVEEA 464
+ YR I+ YKI+AK+LS R++ +L +I Q+AF+ GR + D+V++ ++++
Sbjct: 778 VSEYRTIAPCNTQYKIIAKILSKRMQPLLRSIISPSQSAFVPGRAISDNVLITHKILHYL 599
Query: 465 KR---RKKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHWIKGCIQSASSSVLVN 521
++ +K K D KAYD + WNFL +++R+GF WI WI C+ + S S L+N
Sbjct: 598 RQSGAKKHVSMAVKTDMTKAYDRIAWNFLREVLTRLGFHGIWISWIMECVSTVSYSFLIN 419
Query: 522 GSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFTGYRVGGDEVEISLLQ 581
G P RGLRQGDPL+P+LF++ E L+GL +QA+++ G +V + I+ L
Sbjct: 418 GGPQGRVLPSRGLRQGDPLSPYLFILCTEVLSGLCQQALRKGTLPGVKVARNCPPINHLL 239
Query: 582 FADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGISMVSR 630
FADDT+FFG+++ + ++ SI+ + A SG +N KS + S S+
Sbjct: 238 FADDTMFFGKSNASSCAILLSIMDKYRAASGRCIN*TKSAITFSSKTSQ 92
>BE248682 similar to GP|18568269|gb putative gag-pol polyprotein {Zea mays},
partial (1%)
Length = 441
Score = 130 bits (328), Expect(2) = 2e-30
Identities = 65/132 (49%), Positives = 89/132 (67%)
Frame = +3
Query: 526 EEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFTGYRVGGDEVEISLLQFADD 585
EE ++ RGL+QGDPLAPFLFL+VAEG++GLM+ AV + LF G+ V +S LQ+ADD
Sbjct: 6 EEISVQRGLKQGDPLAPFLFLLVAEGISGLMKNAVNRNLFQGFDVKRGGTRVSHLQYADD 185
Query: 586 TLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGISMVSRETQIFAAMLNCKVMG 645
TL G + N+ +K++L+ FE SGLKVNF KS L GI++ + LNC+
Sbjct: 186 TLCIGMPTVDNLWTLKALLQGFEMASGLKVNFHKSSLIGINVPRDFMEAACRFLNCREES 365
Query: 646 VPFTYLGIPVGA 657
+PF YLG+P G+
Sbjct: 366 IPFIYLGLPGGS 401
Score = 20.4 bits (41), Expect(2) = 2e-30
Identities = 7/8 (87%), Positives = 8/8 (99%)
Frame = +1
Query: 654 PVGANPRK 661
PVGANP+K
Sbjct: 391 PVGANPKK 414
>BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgaris}, partial
(69%)
Length = 616
Score = 111 bits (277), Expect = 1e-24
Identities = 49/139 (35%), Positives = 92/139 (65%), Gaps = 2/139 (1%)
Frame = +3
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N +++TL+PK N + N+RPI+ +YKI++K+L+ R++ VL+ V+ + Q+AF+ GR
Sbjct: 195 NCTYVTLLPKEVNVTSVKNFRPIACCSVIYKIISKILTSRMQGVLNSVVSENQSAFVKGR 374
Query: 449 FMLDSVVVANEVVEEAKRR--KKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHW 506
+ D++++++E+V+ R+ C V K+D KAYDS +W F+ ++M +GF +K+++W
Sbjct: 375 VIFDNIILSHELVKSYSRKGISPRCMV-KIDLXKAYDSXEWPFIKHLMLELGFPYKFVNW 551
Query: 507 IKGCIQSASSSVLVNGSPT 525
+ + +AS + NG T
Sbjct: 552 VMAXLTTASYTFNXNGDLT 608
>BG647708 weakly similar to GP|13786450|gb| putative reverse transcriptase
{Oryza sativa}, partial (9%)
Length = 708
Score = 70.9 bits (172), Expect = 2e-12
Identities = 34/92 (36%), Positives = 59/92 (63%)
Frame = +1
Query: 532 RGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFTGYRVGGDEVEISLLQFADDTLFFGE 591
+GLRQGDPL+P+LF++ A L+GL+++ ++ G +V + +I+ L FADD+L F
Sbjct: 10 KGLRQGDPLSPYLFILCANVLSGLLKREGNKQNLHGIQVARSDPKITHLLFADDSLLFAR 189
Query: 592 ASTQNILVVKSILRWFEAMSGLKVNFFKSKLA 623
A+ + +L +++ SG VNF KS+++
Sbjct: 190 ANLTEAATIMQVLHSYQSASGQLVNFEKSEVS 285
>BQ148771
Length = 680
Score = 67.8 bits (164), Expect = 2e-11
Identities = 39/109 (35%), Positives = 57/109 (51%), Gaps = 1/109 (0%)
Frame = -3
Query: 680 KHKSLSFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGEE-GRGVAWV 738
K LS R+ L KSV+ ++PL+ PK I++ K+QRKF+WG E R V
Sbjct: 561 KANHLSLARRVTLAKSVIEAVPLYPMMTTIIPKACIEEIQKLQRKFVWGDTEVSRRYHAV 382
Query: 739 KWEEICKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
WE + K K GLG++ L+ NKA ++K W + L +V+ K
Sbjct: 381 GWETMSKPKTIYGLGLRRLDVMNKACIMKLGWSIYSGSNSLCTEVMRGK 235
>BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [imported] -
Arabidopsis thaliana, partial (7%)
Length = 687
Score = 66.2 bits (160), Expect = 5e-11
Identities = 34/81 (41%), Positives = 52/81 (63%)
Frame = +2
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N +FI LIPK +NP +YRPISL V KI+ K+++ R+++ L VID Q+AF+ GR
Sbjct: 428 NKTFIVLIPKGKNPNTPKDYRPISLCNVVMKIITKVIANRVKQTLPDVIDVEQSAFVQGR 607
Query: 449 FMLDSVVVANEVVEEAKRRKK 469
+ D+ ++A V + R+K
Sbjct: 608 LITDNALIAWSVSIG*RXRRK 670
>TC90516
Length = 983
Score = 60.5 bits (145), Expect = 2e-09
Identities = 46/147 (31%), Positives = 72/147 (48%)
Frame = -2
Query: 392 FITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGRFML 451
F+TLI KVE P LG++ +S +G + K++AK+L++RL ++ K+I Q+ F+ GR +
Sbjct: 340 FLTLISKVEYPILLGDFSLMSFLGSL*KLMAKVLALRLAHIMEKIIFVNQSTFVRGRQHV 161
Query: 452 DSVVVANEVVEEAKRRKKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHWIKGCI 511
D VV NE++ V D VD + Y
Sbjct: 160 DGVVAINEII-----------VLMGD-----GVVD*SVCVY------------------- 86
Query: 512 QSASSSVLVNGSPTEEFNMGRGLRQGD 538
++LVN SPT E ++ +GL+QGD
Sbjct: 85 --K*PAILVNCSPT*EIDIQKGLKQGD 11
>TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative retroelement
{Oryza sativa (japonica cultivar-group)}, partial (1%)
Length = 625
Score = 45.1 bits (105), Expect(2) = 1e-08
Identities = 21/53 (39%), Positives = 33/53 (61%)
Frame = +2
Query: 510 CIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQ 562
C++S VLVN + RGL+QGD L+P++F+I EGL+ L+ A ++
Sbjct: 104 CVESNDYYVLVNNDAVDPIIPSRGLQQGDHLSPYIFIICVEGLSFLIPHAKER 262
Score = 32.7 bits (73), Expect(2) = 1e-08
Identities = 11/28 (39%), Positives = 22/28 (78%)
Frame = +1
Query: 480 KAYDSVDWNFLFYMMSRIGFSHKWIHWI 507
K Y+ VD ++L +M ++GF+++WI+W+
Sbjct: 13 KVYNRVD*DYLKEIMIKMGFNNRWIYWM 96
>BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (16%)
Length = 794
Score = 52.8 bits (125), Expect = 5e-07
Identities = 30/82 (36%), Positives = 46/82 (55%)
Frame = +1
Query: 377 NSIKMGGGLRGGNASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKV 436
NS + G R L+PK + L +RPISL YKIV+K+LS RL+ VL +
Sbjct: 547 NSSEQGNLKRESTKQTSGLVPKKLEAKRLVEFRPISLCNVAYKIVSKVLSKRLKSVLPWI 726
Query: 437 IDDRQTAFIGGRFMLDSVVVAN 458
I + Q AF + + D++++A+
Sbjct: 727 ITETQAAFGRRQLISDNILIAH 792
>BG584442
Length = 775
Score = 50.4 bits (119), Expect = 3e-06
Identities = 31/99 (31%), Positives = 49/99 (49%), Gaps = 2/99 (2%)
Frame = +1
Query: 671 KMQRKLSL*KHKSLSFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWG-- 728
K +K++ *++K LS +IK L S+ + S F +D+ KI F W
Sbjct: 370 KFDKKINF*RNKCLSKVM*EVMIKYALQSISSYVMSIFLLLNSQVDEIEKIMNTFSWVHV 549
Query: 729 GEEGRGVAWVKWEEICKKKEEGGLGVKDLESFNKALLVK 767
GE +G+ W+ E++ K GG+G D +FN +L K
Sbjct: 550 GENRKGMHWMS*EKLFVHKNYGGMGFTDFTTFNIPMLGK 666
>CB891696
Length = 638
Score = 50.1 bits (118), Expect = 3e-06
Identities = 30/76 (39%), Positives = 43/76 (56%)
Frame = +1
Query: 595 QNILVVKSILRWFEAMSGLKVNFFKSKLAGISMVSRETQIFAAMLNCKVMGVPFTYLGIP 654
+NIL +K+I+ +FE S L VNF KS L ++++ + CKV V F YLGI
Sbjct: 7 ENILTMKTIVSYFELASSLWVNFLKSGLINLNVIGHF*GW*NIYIKCKVH*VIFKYLGIL 186
Query: 655 VGANPRKAATWEPVIK 670
VG NP + E ++K
Sbjct: 187 VGENPCRVNM*ELLLK 234
>BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - Arabidopsis
thaliana, partial (17%)
Length = 677
Score = 49.7 bits (117), Expect = 4e-06
Identities = 25/59 (42%), Positives = 36/59 (60%)
Frame = +3
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGG 447
N + I LIPK + P + RPISL YKI++K+L RL+ L +I + Q+AF+ G
Sbjct: 501 NTTNICLIPKKKRPTRMTELRPISLCNVGYKIISKVLCQRLKVCLPSLISETQSAFVHG 677
>TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarotenoid
dioxygenase1 {Pisum sativum}, partial (43%)
Length = 1865
Score = 49.3 bits (116), Expect = 6e-06
Identities = 21/42 (50%), Positives = 30/42 (71%)
Frame = -3
Query: 744 CKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLE 785
C + +GGLGV+D+ N +LL KW WRLL+++ LW +VLE
Sbjct: 969 CLPRCKGGLGVRDIRLVNVSLLAKWWWRLLQDQSSLWKEVLE 844
>BF647389
Length = 404
Score = 49.3 bits (116), Expect = 6e-06
Identities = 39/126 (30%), Positives = 58/126 (45%), Gaps = 1/126 (0%)
Frame = +2
Query: 646 VPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSLSFGGRICLIKSVLSSLPLFFF 705
+P YLG+P+ + K +KK+ ++ * K LS+ G + LIK VL + ++
Sbjct: 32 LPSRYLGVPLSS---KKLYVIQRVKKIICRIEN*SSKLLSYAGSLQLIKIVLFGVQPYWS 202
Query: 706 SFFKAPKCIIDQCNKIQRKFLWGGEEGRGV-AWVKWEEICKKKEEGGLGVKDLESFNKAL 764
F P +I R FL G+ G A + E IC K GG V DL+ N+
Sbjct: 203 QVFVLP*KVIKLIQTTCRIFL*TGKSGTSKRALIAREHICLPKTAGGWNVIDLKVXNQTA 382
Query: 765 LVKWRW 770
+ K W
Sbjct: 383 ICKLXW 400
>TC83624 homologue to PIR|G84581|G84581 copia-like retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(1%)
Length = 831
Score = 43.9 bits (102), Expect = 2e-04
Identities = 17/35 (48%), Positives = 27/35 (76%)
Frame = +1
Query: 639 LNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQ 673
L C V VPF +LG+P+GANP++++T +PV+ +Q
Sbjct: 322 LLCNVNEVPFCFLGLPIGANPKRSSTRKPVLDSLQ 426
>TC82948
Length = 705
Score = 40.4 bits (93), Expect = 0.003
Identities = 16/43 (37%), Positives = 27/43 (62%)
Frame = +3
Query: 735 VAWVKWEEICKKKEEGGLGVKDLESFNKALLVKWRWRLLRERE 777
V V WE++C+ +EG LG++ L N+AL +K W ++ +E
Sbjct: 258 VVKVSWEKVCRPIKEGSLGIRSLSKLNEALNLKLCWDMMISKE 386
>TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non-LTR
retroelement reverse transcriptase {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 1262
Score = 38.1 bits (87), Expect = 0.013
Identities = 19/37 (51%), Positives = 27/37 (72%)
Frame = +1
Query: 751 GLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
GLGV +FN +LL KW WRLL ++E LW +VL+++
Sbjct: 31 GLGVG---AFNLSLLGKWCWRLLVDKEGLWHRVLKAR 132
>AW774658 similar to GP|2808681|emb| Hcr9-4B {Lycopersicon hirsutum}, partial
(4%)
Length = 665
Score = 25.0 bits (53), Expect(2) = 2.0
Identities = 14/27 (51%), Positives = 18/27 (65%)
Frame = -3
Query: 590 GEASTQNILVVKSILRWFEAMSGLKVN 616
GE + ++ILV+ FE MSGLKVN
Sbjct: 333 GERALRSILVI------FENMSGLKVN 271
Score = 24.3 bits (51), Expect(2) = 2.0
Identities = 12/19 (63%), Positives = 12/19 (63%)
Frame = -1
Query: 578 SLLQFADDTLFFGEASTQN 596
S LQFADDTL G S N
Sbjct: 383 SHLQFADDTLLLGVKSWAN 327
>BF006686
Length = 325
Score = 30.8 bits (68), Expect = 2.1
Identities = 13/29 (44%), Positives = 20/29 (68%)
Frame = +3
Query: 665 WEPVIKKMQRKLSL*KHKSLSFGGRICLI 693
WEP+++ + + L +K LSFGGRI L+
Sbjct: 237 WEPLLEHVNKMLKSWGNKLLSFGGRIVLL 323
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.341 0.150 0.502
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,058,922
Number of Sequences: 36976
Number of extensions: 374454
Number of successful extensions: 2951
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 1665
Number of HSP's successfully gapped in prelim test: 132
Number of HSP's that attempted gapping in prelim test: 1214
Number of HSP's gapped (non-prelim): 1891
length of query: 787
length of database: 9,014,727
effective HSP length: 104
effective length of query: 683
effective length of database: 5,169,223
effective search space: 3530579309
effective search space used: 3530579309
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0263.7


