

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
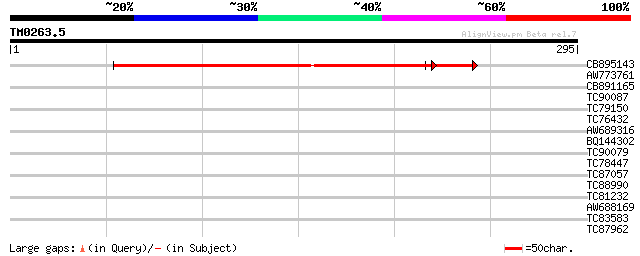
Query= TM0263.5
(295 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CB895143 218 3e-62
AW773761 similar to GP|21618133|gb| bcnt-like protein {Arabidops... 32 0.29
CB891165 homologue to PIR|H82068|H820 RNA polymerase-associated ... 32 0.38
TC90087 similar to PIR|T05841|T05841 spliceosome-associated prot... 31 0.50
TC79150 similar to GP|18461304|dbj|BAB84499. putative receptor-l... 31 0.65
TC76432 weakly similar to GP|15810523|gb|AAL07149.1 unknown prot... 30 0.85
AW689316 similar to GP|20259452|g unknown protein {Arabidopsis t... 30 1.1
BQ144302 similar to GP|6815109|dbj| hypothetical protein {Oryza ... 25 1.8
TC90079 similar to GP|5454204|gb|AAD43619.1| T3P18.18 {Arabidops... 29 2.5
TC78447 similar to GP|7768151|emb|CAB90633.1 protein phpsphatase... 28 3.2
TC87057 similar to PIR|T48135|T48135 nucleotide sugar epimerase-... 28 4.2
TC88990 similar to PIR|T11751|T11751 transcription repressor ROM... 27 9.4
TC81232 similar to GP|14423428|gb|AAK62396.1 Unknown protein {Ar... 27 9.4
AW688169 weakly similar to GP|21745054|gb| putative disease resi... 27 9.4
TC83583 similar to PIR|T06379|T06379 SAR DNA-binding protein 2 -... 27 9.4
TC87962 similar to GP|14596039|gb|AAK68747.1 Unknown protein {Ar... 27 9.4
>CB895143
Length = 770
Score = 218 bits (554), Expect(2) = 3e-62
Identities = 103/168 (61%), Positives = 133/168 (78%)
Frame = +2
Query: 55 WNTEQNLVLISGWIKYGTSSVVGRNQKGETYWSQIAEYCHEHCSFDPPREGGACRNCFNY 114
WN +QNL+LISGWIKYGT+SVVGRNQK ++Y +I +Y +EHCSF+PP + CRN +NY
Sbjct: 209 WNIDQNLILISGWIKYGTNSVVGRNQKCDSY*GKIVDYFNEHCSFNPPCDAAPCRNHYNY 388
Query: 115 MSNQLNKWIGAYDGAKRVQGSGWSEDDILAKAQDIYTNGKNSHPFTLMKEWLALCNEPHY 174
M+ LNKWI AYDGAKR+Q SG S+ D+LAKA ++Y++GK+ H F L+ W + ++P Y
Sbjct: 389 MNKILNKWIDAYDGAKRMQQSG*SKIDVLAKAHELYSSGKSEH-FILISGWFVVRDQPRY 565
Query: 175 CSQVGGNLGSGSSGSKRSHESDACGSNSIGSIPRPMGREAAKKKNKKK 222
SQVGGN GSGSSGSKR+H+SDA SNS+GS PMGR+AAKK+ K+K
Sbjct: 566 DSQVGGNTGSGSSGSKRAHKSDASDSNSVGSSSCPMGRDAAKKR*KEK 709
Score = 38.5 bits (88), Expect(2) = 3e-62
Identities = 16/27 (59%), Positives = 21/27 (77%)
Frame = +3
Query: 217 KKNKKKSRDAALEVVNNEWSEYKQLRE 243
KK +KKS+ A LE VN EW+E+KQ +E
Sbjct: 690 KKGEKKSKGATLESVNEEWNEFKQCKE 770
>AW773761 similar to GP|21618133|gb| bcnt-like protein {Arabidopsis
thaliana}, partial (35%)
Length = 622
Score = 32.0 bits (71), Expect = 0.29
Identities = 23/97 (23%), Positives = 44/97 (44%), Gaps = 3/97 (3%)
Frame = -1
Query: 194 ESDACGSNSIGSIPRPMGREAAKKKNKKKSRDAALEVVNNEWSEYKQLR---EQELERLE 250
+SD+ + P P +A ++ KKK++ + L+ +W E+K+ E EL+ +
Sbjct: 604 DSDSKEAIERAKAPAPSAVDAVLEQIKKKAKLSVLDKTKKDWGEFKEENKGLENELDTYK 425
Query: 251 KISSVQQEANRLEKMKMFLQLSFEEHLDDRRKELLKQ 287
K S+ L+K+ + + E +R L Q
Sbjct: 424 KSSN-----QYLDKVSFLQRTDYREFERERDARLALQ 329
>CB891165 homologue to PIR|H82068|H820 RNA polymerase-associated protein HepA
VC2506 [imported] - Vibrio cholerae, partial (1%)
Length = 692
Score = 31.6 bits (70), Expect = 0.38
Identities = 18/65 (27%), Positives = 37/65 (56%)
Frame = -2
Query: 218 KNKKKSRDAALEVVNNEWSEYKQLREQELERLEKISSVQQEANRLEKMKMFLQLSFEEHL 277
++ KK + + ++ + E K+L E+ + EKI + ++ N +E K+ L+ S E H
Sbjct: 451 RDGKKQKKSMMKEKSQFEDEAKRLVEEGEKVEEKIRILVEQKNSIELEKIKLKESMERHE 272
Query: 278 DDRRK 282
D+++K
Sbjct: 271 DEKKK 257
>TC90087 similar to PIR|T05841|T05841 spliceosome-associated protein homolog
F17L22.120 - Arabidopsis thaliana, partial (12%)
Length = 733
Score = 31.2 bits (69), Expect = 0.50
Identities = 29/105 (27%), Positives = 49/105 (46%), Gaps = 6/105 (5%)
Frame = +2
Query: 180 GNLGSGSSGSKRSHESDACGSNSIGSIPRPMGREAAKKKNK-KKSRDAALEVVNNEWSEY 238
G+LGS ++ K+S ESD R R KK NK K ++A + + E
Sbjct: 140 GDLGSKTATVKKSRESD-----------RRRRRRKQKKINKASKEQNANASEDDTDAKEN 286
Query: 239 KQLREQELERLEKISSVQQEANRLEKM-----KMFLQLSFEEHLD 278
+ ++Q++ +I V ++A E + K+F + SF E +D
Sbjct: 287 TEQQQQQVVEQVEIEYVPEKAELYEGLDEEFKKIFEKFSFSEVID 421
>TC79150 similar to GP|18461304|dbj|BAB84499. putative receptor-like protein
kinase {Oryza sativa (japonica cultivar-group)}, partial
(7%)
Length = 1077
Score = 30.8 bits (68), Expect = 0.65
Identities = 21/76 (27%), Positives = 29/76 (37%), Gaps = 11/76 (14%)
Frame = +1
Query: 58 EQNLVLISGWIKYGTSSVVGRNQKGETYWSQIAEYC-----------HEHCSFDPPREGG 106
++N + S WI V N K +T+ S +A YC + CS+DP G
Sbjct: 796 QRNFIDYSKWIMMVNPRVYSFNDKTKTWGSFMASYC*TM*GSWYLWRNSMCSYDPV-NGR 972
Query: 107 ACRNCFNYMSNQLNKW 122
C Y N W
Sbjct: 973 TCYCLKGYKLKNRNDW 1020
>TC76432 weakly similar to GP|15810523|gb|AAL07149.1 unknown protein
{Arabidopsis thaliana}, partial (43%)
Length = 1594
Score = 30.4 bits (67), Expect = 0.85
Identities = 15/41 (36%), Positives = 20/41 (48%)
Frame = +1
Query: 96 HCSFDPPREGGACRNCFNYMSNQLNKWIGAYDGAKRVQGSG 136
HC F PR+ A +NC+NY N+ DG + G G
Sbjct: 1327 HCPFFTPRQVSAGKNCWNYSCNK--------DGGNKEGGFG 1425
>AW689316 similar to GP|20259452|g unknown protein {Arabidopsis thaliana},
partial (13%)
Length = 648
Score = 30.0 bits (66), Expect = 1.1
Identities = 15/34 (44%), Positives = 22/34 (64%), Gaps = 1/34 (2%)
Frame = -1
Query: 104 EGGACRNCFNYMSNQLNKWI-GAYDGAKRVQGSG 136
EG ACR C S+ +N+WI G+ D ++R + SG
Sbjct: 240 EGEACRLCIPPSSSFINRWITGSKDESRRRERSG 139
>BQ144302 similar to GP|6815109|dbj| hypothetical protein {Oryza sativa
(japonica cultivar-group)}, partial (3%)
Length = 1047
Score = 25.0 bits (53), Expect(2) = 1.8
Identities = 15/46 (32%), Positives = 22/46 (47%)
Frame = -1
Query: 180 GNLGSGSSGSKRSHESDACGSNSIGSIPRPMGREAAKKKNKKKSRD 225
GN+G G + G+N I R G E K+++KKK R+
Sbjct: 489 GNMGEKGGGG------E*WGNNKIKGKTRANGVEKNKRRDKKKKRE 370
Score = 22.7 bits (47), Expect(2) = 1.8
Identities = 9/25 (36%), Positives = 12/25 (48%)
Frame = -2
Query: 117 NQLNKWIGAYDGAKRVQGSGWSEDD 141
N KW G G K +G W E++
Sbjct: 608 NGWGKWKGERYGKKEKKGRKWGEEE 534
>TC90079 similar to GP|5454204|gb|AAD43619.1| T3P18.18 {Arabidopsis
thaliana}, partial (38%)
Length = 677
Score = 28.9 bits (63), Expect = 2.5
Identities = 13/47 (27%), Positives = 21/47 (44%)
Frame = -2
Query: 6 SQTPPFNVGVTDFPEFSTNMTLGDMTAVNEVTPNEEDSTPRSRKTQS 52
+++PP + FP F + T VN PN ++ + R T S
Sbjct: 445 TESPPLDSTTQSFPPFLSTFTTSVSNRVNS*IPNSSANSSKKRSTSS 305
>TC78447 similar to GP|7768151|emb|CAB90633.1 protein phpsphatase 2C (PP2C)
{Fagus sylvatica}, partial (37%)
Length = 833
Score = 28.5 bits (62), Expect = 3.2
Identities = 25/112 (22%), Positives = 43/112 (38%), Gaps = 1/112 (0%)
Frame = +1
Query: 69 KYGTSSVVGRNQKGETYWSQIAEYCHE-HCSFDPPREGGACRNCFNYMSNQLNKWIGAYD 127
KYG +SV GR ++ E S +C E F +G C + +L++ +
Sbjct: 388 KYGITSVCGRRREMEDAVSVHPSFCREKQDHFFGVYDGHGCSHVATMCKERLHEIVEEEV 567
Query: 128 GAKRVQGSGWSEDDILAKAQDIYTNGKNSHPFTLMKEWLALCNEPHYCSQVG 179
++V E + +++ + K F+ E PH C VG
Sbjct: 568 EKEKVDWKSTMEKSFIRMDEEVLNSSKTKQSFSCKCE----LQTPH-CDAVG 708
>TC87057 similar to PIR|T48135|T48135 nucleotide sugar epimerase-like
protein - Arabidopsis thaliana, partial (80%)
Length = 1655
Score = 28.1 bits (61), Expect = 4.2
Identities = 14/38 (36%), Positives = 22/38 (57%)
Frame = +2
Query: 99 FDPPREGGACRNCFNYMSNQLNKWIGAYDGAKRVQGSG 136
F+ P +GG+ F Y+ + + +GA D AK+ GSG
Sbjct: 938 FESP-DGGSVARDFTYIDDVVKGCLGALDTAKKSTGSG 1048
>TC88990 similar to PIR|T11751|T11751 transcription repressor ROM1 - kidney
bean, partial (91%)
Length = 1528
Score = 26.9 bits (58), Expect = 9.4
Identities = 19/50 (38%), Positives = 21/50 (42%)
Frame = +1
Query: 179 GGNLGSGSSGSKRSHESDACGSNSIGSIPRPMGREAAKKKNKKKSRDAAL 228
GG GSGS SH D S S GS + +NKK S D L
Sbjct: 622 GGKAGSGSGNDGFSHSGD---SGSEGSSNASDENQQESARNKKGSFDLML 762
>TC81232 similar to GP|14423428|gb|AAK62396.1 Unknown protein {Arabidopsis
thaliana}, partial (64%)
Length = 957
Score = 26.9 bits (58), Expect = 9.4
Identities = 11/34 (32%), Positives = 18/34 (52%)
Frame = +1
Query: 4 SPSQTPPFNVGVTDFPEFSTNMTLGDMTAVNEVT 37
+PSQ PP +F + ST T+ T+ + +T
Sbjct: 241 NPSQPPPLTTNAVEFLQSSTGATVTAFTSTSNLT 342
>AW688169 weakly similar to GP|21745054|gb| putative disease resistance gene
analog NBS-LRR {Malus prunifolia}, partial (16%)
Length = 540
Score = 26.9 bits (58), Expect = 9.4
Identities = 12/41 (29%), Positives = 24/41 (58%)
Frame = +2
Query: 223 SRDAALEVVNNEWSEYKQLREQELERLEKISSVQQEANRLE 263
+R+ + V +E K++ + + LEK+ V +EAN+L+
Sbjct: 44 ARERMIHSVKSERENGKEIEKDVMNWLEKVDGVIKEANQLQ 166
>TC83583 similar to PIR|T06379|T06379 SAR DNA-binding protein 2 - garden
pea, partial (33%)
Length = 852
Score = 26.9 bits (58), Expect = 9.4
Identities = 12/33 (36%), Positives = 18/33 (54%)
Frame = +2
Query: 213 EAAKKKNKKKSRDAALEVVNNEWSEYKQLREQE 245
E KK+ KKK +D+ + EW Y + R +E
Sbjct: 488 EVVKKEKKKKKKDST*KGRGAEWRPYFEWRRKE 586
>TC87962 similar to GP|14596039|gb|AAK68747.1 Unknown protein {Arabidopsis
thaliana}, partial (90%)
Length = 1268
Score = 26.9 bits (58), Expect = 9.4
Identities = 20/82 (24%), Positives = 35/82 (42%), Gaps = 5/82 (6%)
Frame = +3
Query: 212 REAAKKKNKKKSRDAALEVVNNEWSEYKQLREQELERLE-----KISSVQQEANRLEKMK 266
RE K + + + +V W E+ + + +E ER ++ ++++ K K
Sbjct: 459 RELQKALDLGRGANPKGYMVEEIWQEFAKAKYKEWERSSSQRSWELQNLKEACKSALKEK 638
Query: 267 MFLQLSFEEHLDDRRKELLKQL 288
FL E +DD LKQL
Sbjct: 639 HFLGSEMEGFVDDATTSHLKQL 704
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.312 0.128 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,941,094
Number of Sequences: 36976
Number of extensions: 153873
Number of successful extensions: 672
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 667
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 671
length of query: 295
length of database: 9,014,727
effective HSP length: 95
effective length of query: 200
effective length of database: 5,502,007
effective search space: 1100401400
effective search space used: 1100401400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0263.5


