

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0258b.1
(207 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
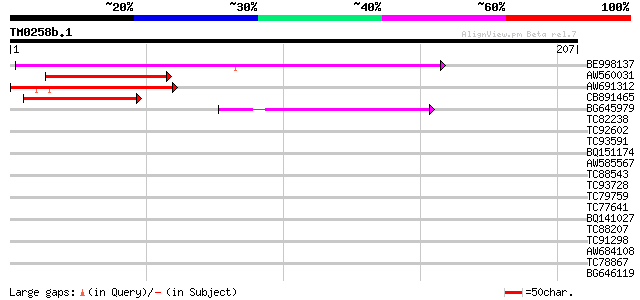
BE998137 similar to GP|10177429|db zinc finger protein SHI-like ... 151 1e-37
AW560031 similar to GP|882341|gb|AA LRP1 {Arabidopsis thaliana},... 81 3e-16
AW691312 similar to GP|14586363|emb lateral root primordia (LRP1... 79 2e-15
CB891465 similar to PIR|G84600|G846 hypothetical protein At2g214... 77 5e-15
BG645979 similar to GP|15528818|dbj putative LRP(lateral root pr... 55 2e-08
TC82238 weakly similar to GP|8570046|dbj|BAA96751.1 Similar to A... 31 0.38
TC92602 similar to SP|P57997|IF2C_PHAVU Translation initiation f... 31 0.38
TC93591 similar to PIR|T05209|T05209 hypothetical protein F24J7.... 30 0.65
BQ151174 similar to GP|11036868|gb| PxORF73 peptide {Plutella xy... 30 0.65
AW585567 30 0.85
TC88543 similar to GP|20466782|gb|AAM20708.1 unknown protein {Ar... 29 1.1
TC93728 similar to GP|1621467|gb|AAB17194.1| laccase {Liriodendr... 29 1.1
TC79759 homologue to GP|15215590|gb|AAK91340.1 AT5g62070/mtg10_9... 29 1.5
TC77641 similar to GP|21595125|gb|AAM66074.1 endosperm-specific ... 29 1.5
BQ141027 weakly similar to GP|11320891|gb| pinin {Homo sapiens},... 29 1.5
TC88207 28 1.9
TC91298 similar to GP|3928086|gb|AAC79612.1| unknown protein {Ar... 28 1.9
AW684108 28 2.5
TC78867 28 2.5
BG646119 weakly similar to PIR|T46196|T46 cytochrome P450-like p... 28 3.2
>BE998137 similar to GP|10177429|db zinc finger protein SHI-like {Arabidopsis
thaliana}, partial (36%)
Length = 592
Score = 151 bits (382), Expect = 1e-37
Identities = 75/159 (47%), Positives = 95/159 (59%), Gaps = 2/159 (1%)
Frame = +1
Query: 3 MMLGQEVKGSRCQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLMEH 62
M +G CQ+CGNQAKK C + RCRTCC ++GFQCQTHV+STW+P RRR R +
Sbjct: 4 MRSSSSAEGISCQDCGNQAKKDCPHMRCRTCCKSRGFQCQTHVKSTWVPASRRRERQQQL 183
Query: 63 QPPPTTNNPHHLHEDIPQS--HNQNPFTSLEELKFPEAMSSMAVFSSVRVRSMDDSVNEM 120
P + D + +N T LEE FP +SS A F VRV S+DD+ +
Sbjct: 184 SSSPLQRDISKRPRDGSNALVSTRNFHTGLEEANFPAVVSSPAEFRCVRVSSIDDADDRY 363
Query: 121 AYQTSVNIGGHRFSGILYDQGPEQQSLNASPLDQHQNLN 159
AYQT+VNIGGH F GILYD GPE + N++ + N N
Sbjct: 364 AYQTAVNIGGHLFKGILYDFGPESSTNNSNNNSNYNNSN 480
>AW560031 similar to GP|882341|gb|AA LRP1 {Arabidopsis thaliana}, partial
(24%)
Length = 756
Score = 80.9 bits (198), Expect = 3e-16
Identities = 31/46 (67%), Positives = 37/46 (80%)
Frame = +1
Query: 14 CQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRL 59
CQ+CGNQAKK C+ RCRTCC ++GF C THV+STW+P RRR RL
Sbjct: 493 CQDCGNQAKKDCSNRRCRTCCKSRGFDCPTHVKSTWVPAARRRERL 630
>AW691312 similar to GP|14586363|emb lateral root primordia (LRP1)
{Arabidopsis thaliana}, partial (14%)
Length = 652
Score = 78.6 bits (192), Expect = 2e-15
Identities = 35/66 (53%), Positives = 46/66 (69%), Gaps = 5/66 (7%)
Frame = +1
Query: 1 MSMMLGQE--VKGSR---CQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRR 55
++MM +E V GS CQ+CGN+AKK C + RCRTCC +G+ C TH++STWIP RR
Sbjct: 370 IAMMESEENGVYGSEYRVCQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRR 549
Query: 56 RHRLME 61
R R +E
Sbjct: 550 REREVE 567
>CB891465 similar to PIR|G84600|G846 hypothetical protein At2g21400
[imported] - Arabidopsis thaliana, partial (20%)
Length = 607
Score = 77.0 bits (188), Expect = 5e-15
Identities = 29/43 (67%), Positives = 37/43 (85%)
Frame = -1
Query: 6 GQEVKGSRCQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRST 48
G+ +KG +CQ+CGNQAKK CAYSRCR+CC NK F CQTH+R++
Sbjct: 427 GEVLKGLKCQDCGNQAKKDCAYSRCRSCCKNKDFNCQTHIRTS 299
Score = 29.6 bits (65), Expect = 0.85
Identities = 15/53 (28%), Positives = 25/53 (46%), Gaps = 3/53 (5%)
Frame = -3
Query: 6 GQEVKGSRCQECGNQAKKGCAYSRCRTCCNNKGF---QCQTHVRSTWIPVDRR 55
G+ VKG++ +K+G + + + F + RSTWIP DR+
Sbjct: 428 GRSVKGTKMSRLWEPSKEGLCILKVQKLLQKQRF*LPNTYQNFRSTWIPADRK 270
>BG645979 similar to GP|15528818|dbj putative LRP(lateral root primordia) 1
{Oryza sativa (japonica cultivar-group)}, partial (22%)
Length = 713
Score = 54.7 bits (130), Expect = 2e-08
Identities = 31/79 (39%), Positives = 39/79 (49%)
Frame = -3
Query: 77 DIPQSHNQNPFTSLEELKFPEAMSSMAVFSSVRVRSMDDSVNEMAYQTSVNIGGHRFSGI 136
D SH F P + + AVF VRV S+DD +E AYQ V IGGH F G
Sbjct: 690 DTTSSHQDAGFKD----SMPGQVRAPAVFKCVRVTSVDDGKDEYAYQAVVKIGGHVFKGF 523
Query: 137 LYDQGPEQQSLNASPLDQH 155
LYD G E + + + + H
Sbjct: 522 LYDHGVENREVYPNLSELH 466
>TC82238 weakly similar to GP|8570046|dbj|BAA96751.1 Similar to Arabidopsis
thaliana chromosome4 BAC clone T16H5; lectin like
protein (AL024486), partial (24%)
Length = 819
Score = 30.8 bits (68), Expect = 0.38
Identities = 15/51 (29%), Positives = 23/51 (44%), Gaps = 1/51 (1%)
Frame = +3
Query: 63 QPPPTTNNPHHLHEDI-PQSHNQNPFTSLEELKFPEAMSSMAVFSSVRVRS 112
QPP + H H + P H QN T+ EEL+ P ++ + +S
Sbjct: 33 QPPQSQPQQTHQHSQLQPHQHRQNEPTNKEELRLPSRRENLLLSPQYHTKS 185
>TC92602 similar to SP|P57997|IF2C_PHAVU Translation initiation factor IF-2
chloroplast precursor (PvIF2cp). [Kidney bean French
bean], partial (17%)
Length = 912
Score = 30.8 bits (68), Expect = 0.38
Identities = 17/42 (40%), Positives = 23/42 (54%), Gaps = 5/42 (11%)
Frame = -1
Query: 151 PLDQHQNLNLTTIHSHDG-----ATMAPPSSSATAAHELFFP 187
PL H NL LT +H+ D +++P S + HELFFP
Sbjct: 447 PLPNHHNL-LTQVHTQDRHLPL*QSLSPVSKNQEYQHELFFP 325
>TC93591 similar to PIR|T05209|T05209 hypothetical protein F24J7.50 -
Arabidopsis thaliana, partial (12%)
Length = 688
Score = 30.0 bits (66), Expect = 0.65
Identities = 15/36 (41%), Positives = 19/36 (52%), Gaps = 3/36 (8%)
Frame = +1
Query: 47 STWIPVDRRRHRLMEHQP---PPTTNNPHHLHEDIP 79
S WIP R +HQP P +T P HLH+ +P
Sbjct: 67 SKWIPHLLNNRRGEDHQPFPPPISTAQPPHLHQPLP 174
>BQ151174 similar to GP|11036868|gb| PxORF73 peptide {Plutella xylostella
granulovirus}, partial (42%)
Length = 909
Score = 30.0 bits (66), Expect = 0.65
Identities = 11/26 (42%), Positives = 13/26 (49%)
Frame = +2
Query: 61 EHQPPPTTNNPHHLHEDIPQSHNQNP 86
+H P T +PHH H P H NP
Sbjct: 191 QHDKPQKTTHPHHPHTPPPTPHPPNP 268
>AW585567
Length = 491
Score = 29.6 bits (65), Expect = 0.85
Identities = 23/90 (25%), Positives = 38/90 (41%), Gaps = 7/90 (7%)
Frame = +3
Query: 45 VRSTWIPV----DRRRHRLMEHQPPPTTNNPHHLHEDIPQSHNQNPFTSLEELKFPEAMS 100
+R W P+ + + R H PPP + + + + + T + LKFPE+ S
Sbjct: 18 LRYKWQPIKGVFNAQVKRERNHAPPPKKTPDNSVSWQLQIKNRRITGTITKTLKFPESNS 197
Query: 101 SMAVF---SSVRVRSMDDSVNEMAYQTSVN 127
S +F S R S N++ +S N
Sbjct: 198 SKFIFFFCDSQRFFCKSSSSNQILRNSSAN 287
>TC88543 similar to GP|20466782|gb|AAM20708.1 unknown protein {Arabidopsis
thaliana}, partial (46%)
Length = 2101
Score = 29.3 bits (64), Expect = 1.1
Identities = 12/28 (42%), Positives = 18/28 (63%)
Frame = +3
Query: 11 GSRCQECGNQAKKGCAYSRCRTCCNNKG 38
G++C CG ++K CAY C+TC +G
Sbjct: 981 GAQC--CG*VSRKNCAY*TCKTCSCTQG 1058
>TC93728 similar to GP|1621467|gb|AAB17194.1| laccase {Liriodendron
tulipifera}, partial (21%)
Length = 514
Score = 29.3 bits (64), Expect = 1.1
Identities = 18/63 (28%), Positives = 23/63 (35%)
Frame = -1
Query: 25 CAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLMEHQPPPTTNNPHHLHEDIPQSHNQ 84
CA C TCC+N C + P T +N H HE + + Q
Sbjct: 154 CANMFCMTCCSNMSCMC-----------------*LWQYPQST*HNQHCSHEGLTF*YKQ 26
Query: 85 NPF 87
NPF
Sbjct: 25 NPF 17
>TC79759 homologue to GP|15215590|gb|AAK91340.1 AT5g62070/mtg10_90
{Arabidopsis thaliana}, partial (22%)
Length = 861
Score = 28.9 bits (63), Expect = 1.5
Identities = 16/38 (42%), Positives = 20/38 (52%)
Frame = +1
Query: 7 QEVKGSRCQECGNQAKKGCAYSRCRTCCNNKGFQCQTH 44
Q+ RC CGN + +G + RC TCC G QTH
Sbjct: 400 QQTCNRRC--CGNSSSRGGSSGRC-TCCGGGG---QTH 495
>TC77641 similar to GP|21595125|gb|AAM66074.1 endosperm-specific
protein-like protein {Arabidopsis thaliana}, partial
(82%)
Length = 1488
Score = 28.9 bits (63), Expect = 1.5
Identities = 13/42 (30%), Positives = 22/42 (51%), Gaps = 5/42 (11%)
Frame = +1
Query: 60 MEHQPPPTTNNPHHLHEDIPQSHNQNPFTS-----LEELKFP 96
+ H PPP +P H H ++ H++NP +++L FP
Sbjct: 34 LSHPPPP---HPLHRHHNLRPQHHRNPLLQP*LLPIQQLPFP 150
>BQ141027 weakly similar to GP|11320891|gb| pinin {Homo sapiens}, partial
(4%)
Length = 633
Score = 28.9 bits (63), Expect = 1.5
Identities = 15/57 (26%), Positives = 27/57 (47%)
Frame = +1
Query: 59 LMEHQPPPTTNNPHHLHEDIPQSHNQNPFTSLEELKFPEAMSSMAVFSSVRVRSMDD 115
L+ H PP ++++ SH+ F EE +F E + ++ F S ++ DD
Sbjct: 304 LLSHGPPSSSSSAMSTSSSSQASHSSPQFKEFEEHEFNEVL-ALGYFESQKISYHDD 471
>TC88207
Length = 617
Score = 28.5 bits (62), Expect = 1.9
Identities = 15/45 (33%), Positives = 21/45 (46%), Gaps = 2/45 (4%)
Frame = +2
Query: 7 QEVKGSRCQECGNQAKKGCAYSR--CRTCCNNKGFQCQTHVRSTW 49
Q+ G+ C C K C++S CR C + C+T RS W
Sbjct: 71 QKTVGNACFWCPENFGKCCSWSS*DCRRLCYHGC*LCKTQTRSYW 205
>TC91298 similar to GP|3928086|gb|AAC79612.1| unknown protein {Arabidopsis
thaliana}, partial (66%)
Length = 777
Score = 28.5 bits (62), Expect = 1.9
Identities = 15/46 (32%), Positives = 24/46 (51%), Gaps = 3/46 (6%)
Frame = +1
Query: 43 THVRSTWIPVDRRRHRLMEHQPPPTTNNPHHLHEDIPQ---SHNQN 85
+H +T IP+ R +R +P P++ P+ H+ PQ H QN
Sbjct: 52 SHHSNTRIPISRTLNRRKSEKPKPSSIQPNPNHQWPPQHRHHHRQN 189
>AW684108
Length = 447
Score = 28.1 bits (61), Expect = 2.5
Identities = 11/23 (47%), Positives = 15/23 (64%)
Frame = -1
Query: 18 GNQAKKGCAYSRCRTCCNNKGFQ 40
G + KKG +Y + CNNKGF+
Sbjct: 165 GEREKKGVSYFSFGSFCNNKGFE 97
>TC78867
Length = 800
Score = 28.1 bits (61), Expect = 2.5
Identities = 16/45 (35%), Positives = 24/45 (52%), Gaps = 2/45 (4%)
Frame = -1
Query: 29 RCRTCCNNKGFQCQTHVRSTWIPV--DRRRHRLMEHQPPPTTNNP 71
+C TCC C+ RS P+ +RR HRL+ ++ P+T P
Sbjct: 674 KCWTCC------CRR*RRSRTSPIHHERRPHRLLRNRHLPSTPTP 558
>BG646119 weakly similar to PIR|T46196|T46 cytochrome P450-like protein -
Arabidopsis thaliana, partial (37%)
Length = 712
Score = 27.7 bits (60), Expect = 3.2
Identities = 14/49 (28%), Positives = 23/49 (46%), Gaps = 12/49 (24%)
Frame = +2
Query: 48 TWIPVDRRRHRLMEHQPPPT------------TNNPHHLHEDIPQSHNQ 84
++IP R + +++ QPPP+ +NNPH P HN+
Sbjct: 125 SFIPHHRLPYLILQKQPPPSRFLHSPSLPISNSNNPHQPPRRTPNHHNR 271
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.129 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,255,592
Number of Sequences: 36976
Number of extensions: 155440
Number of successful extensions: 1388
Number of sequences better than 10.0: 71
Number of HSP's better than 10.0 without gapping: 1341
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1377
length of query: 207
length of database: 9,014,727
effective HSP length: 92
effective length of query: 115
effective length of database: 5,612,935
effective search space: 645487525
effective search space used: 645487525
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0258b.1


