

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0258a.2
(167 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
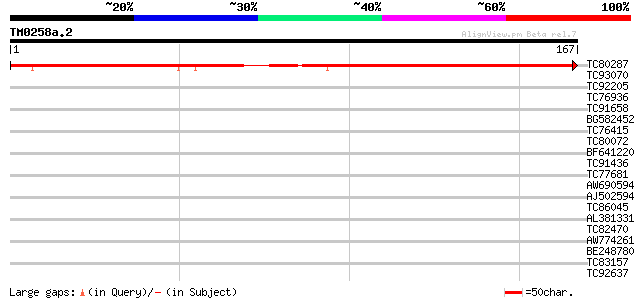
Score E
Sequences producing significant alignments: (bits) Value
TC80287 weakly similar to GP|12667516|gb|AAG61157.1 DBC2 {Homo s... 179 6e-46
TC93070 weakly similar to GP|17065230|gb|AAL32769.1 Unknown prot... 34 0.024
TC92205 similar to GP|12723779|gb|AAK04950.1 UNKNOWN PROTEIN {La... 33 0.070
TC76936 similar to GP|20160782|dbj|BAB89723. putative nascent po... 33 0.070
TC91658 similar to GP|14329812|emb|CAC40753. putative nucleosome... 33 0.070
BG582452 similar to GP|21593180|gb| putative DNA-binding protein... 32 0.091
TC76415 similar to PIR|T05479|T05479 hypothetical protein T8O5.1... 31 0.27
TC80072 similar to GP|14423520|gb|AAK62442.1 Unknown protein {Ar... 30 0.35
BF641220 weakly similar to PIR|G86203|G86 probable N-arginine di... 30 0.35
TC91436 weakly similar to GP|21593499|gb|AAM65466.1 putative nod... 30 0.45
TC77681 similar to GP|22597156|gb|AAN03465.1 nucleolar histone d... 30 0.45
AW690594 similar to GP|23498163|emb hypothetical protein {Plasmo... 30 0.45
AJ502594 weakly similar to PIR|T05755|T057 hypothetical protein ... 30 0.59
TC86045 similar to PIR|T46225|T46225 alpha NAC-like protein - Ar... 29 0.77
AL381331 weakly similar to GP|14209525|dbj contains ESTs C73715(... 29 1.0
TC82470 homologue to GP|23497460|gb|AAN37003.1 hypothetical prot... 29 1.0
AW774261 weakly similar to EGAD|146423|156 vitellogenin {Anolis ... 28 1.3
BE248780 similar to GP|5929884|gb|A nucleolin-related protein NR... 28 1.3
TC83157 similar to GP|21595368|gb|AAM66095.1 unknown {Arabidopsi... 28 1.3
TC92637 similar to PIR|T39903|T39903 serine-rich protein - fissi... 28 1.3
>TC80287 weakly similar to GP|12667516|gb|AAG61157.1 DBC2 {Homo sapiens},
partial (4%)
Length = 779
Score = 179 bits (453), Expect = 6e-46
Identities = 100/184 (54%), Positives = 125/184 (67%), Gaps = 17/184 (9%)
Frame = -3
Query: 1 MENQP-----MQFLLLEQVQFLKVQDDSLLQWELVDVVDAEEEQEQEPDQNQQ------- 48
MENQ MQFL+LEQVQF+KV DDSLLQWELVDVVDAEE+ E+E +
Sbjct: 735 MENQTSSSSSMQFLILEQVQFVKVNDDSLLQWELVDVVDAEEQSEEEERVDDDGDSFISW 556
Query: 49 -DAAPV-EEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAYED---DEEQVVDDEED 103
D++ + + I+++ R L DP ++ +H++ GY+ E D++ DDE +
Sbjct: 555 SDSSRIGDPIELLTHRRIHLDDP-------VENHHHHHV-GYSEEQVNHDDDDDDDDESE 400
Query: 104 DDDRCDLDDELVPWEVGNKLGRQRMKKLGKRACSKMKNSKRSPYLFVRPGCVGGKHGLGL 163
DD DLDDELVPW++GNKLGRQRM+KLGKR SKM NSKRSP LFVRPGCV GKHG+GL
Sbjct: 399 DDGDDDLDDELVPWDIGNKLGRQRMRKLGKRVSSKMNNSKRSPILFVRPGCVRGKHGMGL 220
Query: 164 KHSF 167
KH F
Sbjct: 219 KHCF 208
>TC93070 weakly similar to GP|17065230|gb|AAL32769.1 Unknown protein
{Arabidopsis thaliana}, partial (19%)
Length = 679
Score = 34.3 bits (77), Expect = 0.024
Identities = 14/24 (58%), Positives = 19/24 (78%)
Frame = +1
Query: 90 YEDDEEQVVDDEEDDDDRCDLDDE 113
+EDDE++ D++EDDDD D DDE
Sbjct: 250 FEDDEDEDEDEDEDDDDDDDDDDE 321
Score = 31.2 bits (69), Expect = 0.20
Identities = 19/52 (36%), Positives = 28/52 (53%), Gaps = 2/52 (3%)
Frame = +1
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE--LVPWEVGNKLGRQRMKKLGKRACSKMK 140
ED++E DD++DDDD D DD+ P ++ K R + K + R K K
Sbjct: 268 EDEDEDEDDDDDDDDDDEDDDDDGFAEPTDLNAKDKRLKSKTVPDRQQEKEK 423
>TC92205 similar to GP|12723779|gb|AAK04950.1 UNKNOWN PROTEIN {Lactococcus
lactis subsp. lactis}, partial (2%)
Length = 640
Score = 32.7 bits (73), Expect = 0.070
Identities = 23/74 (31%), Positives = 32/74 (43%), Gaps = 2/74 (2%)
Frame = +3
Query: 34 DAEEEQEQEPDQNQQD--AAPVEEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAYE 91
D +E+ + +P Q Q D A IH++ P P+A V + Q Y+
Sbjct: 378 DDDEDDDWDPSQKQADDGVAQTAVSDSIHEVSEANPQPNAKRV----KVQKELTFQDLYQ 545
Query: 92 DDEEQVVDDEEDDD 105
E DDEEDDD
Sbjct: 546 SQELFDEDDEEDDD 587
>TC76936 similar to GP|20160782|dbj|BAB89723. putative nascent polypeptide
associated complex alpha chain {Oryza sativa (japonica
cultivar-group)}, partial (86%)
Length = 914
Score = 32.7 bits (73), Expect = 0.070
Identities = 17/70 (24%), Positives = 37/70 (52%)
Frame = +3
Query: 70 DAVAVVEIDRQQYHYLHGYAYEDDEEQVVDDEEDDDDRCDLDDELVPWEVGNKLGRQRMK 129
+ + +++Q+ H+ EDD+++ +D+EDDDD D + E + + + R +
Sbjct: 93 EELLAAHLEQQKIHHDEPVV-EDDDDEDDEDDEDDDDEDDDNAEGFEGDASGRSKQTRSE 269
Query: 130 KLGKRACSKM 139
K ++A K+
Sbjct: 270 KKSRKAMLKL 299
>TC91658 similar to GP|14329812|emb|CAC40753. putative nucleosome assembly
protein 1 {Atropa belladonna}, partial (46%)
Length = 583
Score = 32.7 bits (73), Expect = 0.070
Identities = 19/54 (35%), Positives = 29/54 (53%)
Frame = +2
Query: 91 EDDEEQVVDDEEDDDDRCDLDDELVPWEVGNKLGRQRMKKLGKRACSKMKNSKR 144
+ DEE DD++DDDD D DDE E G+ + K+ K K ++++R
Sbjct: 101 DGDEEDDDDDDDDDDDEEDEDDE----EEDEGKGKSKSKRGSKAKPGKDQSTER 250
Score = 28.5 bits (62), Expect = 1.3
Identities = 14/23 (60%), Positives = 17/23 (73%)
Frame = +2
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
EDDE+ D+E+DDDD D DDE
Sbjct: 89 EDDEDG--DEEDDDDDDDDDDDE 151
>BG582452 similar to GP|21593180|gb| putative DNA-binding protein
{Arabidopsis thaliana}, partial (29%)
Length = 762
Score = 32.3 bits (72), Expect = 0.091
Identities = 24/59 (40%), Positives = 30/59 (50%)
Frame = +2
Query: 99 DDEEDDDDRCDLDDELVPWEVGNKLGRQRMKKLGKRACSKMKNSKRSPYLFVRPGCVGG 157
DDEE+DDD D E EVGN+ R R + K+ + +KR P F P C GG
Sbjct: 533 DDEEEDDDNRDEPREGAV-EVGNRRPRGRPPWIQKQTKTTYLCNKR*PKCFEEP-CHGG 703
>TC76415 similar to PIR|T05479|T05479 hypothetical protein T8O5.180 -
Arabidopsis thaliana, partial (30%)
Length = 1114
Score = 30.8 bits (68), Expect = 0.27
Identities = 25/72 (34%), Positives = 35/72 (47%), Gaps = 15/72 (20%)
Frame = +1
Query: 86 HGYAYEDDEEQVV----DDEEDDDDRCDLDDELVPWE-VGNKLGRQRM----------KK 130
+GY YE D+E V D EEDDDD + + P E + +L R ++ +
Sbjct: 580 YGY-YEGDDEVVYHGGEDGEEDDDDDDEYSTRVPPHEIISRRLARSQISSFSVFEGVGRT 756
Query: 131 LGKRACSKMKNS 142
L R SKM+NS
Sbjct: 757 LKGRDLSKMRNS 792
>TC80072 similar to GP|14423520|gb|AAK62442.1 Unknown protein {Arabidopsis
thaliana}, partial (17%)
Length = 746
Score = 30.4 bits (67), Expect = 0.35
Identities = 22/79 (27%), Positives = 40/79 (49%), Gaps = 2/79 (2%)
Frame = +1
Query: 54 EEIKIIHQIRPVLP-DPDAVAVVEIDRQQYHYLHGYAYEDDE-EQVVDDEEDDDDRCDLD 111
+EI I H+ R ++P D +A + + + + ++DE ++ D+EEDDDD D
Sbjct: 367 DEIDIFHKQRDIVPLDINADSAESDEDDEMPIFNDKDIDNDETDEDGDEEEDDDDEDDEY 546
Query: 112 DELVPWEVGNKLGRQRMKK 130
D +G + +Q+ K
Sbjct: 547 DGKEKGFIGQLIRQQKYLK 603
>BF641220 weakly similar to PIR|G86203|G86 probable N-arginine dibasic
convertase [imported] - Arabidopsis thaliana, partial
(5%)
Length = 634
Score = 30.4 bits (67), Expect = 0.35
Identities = 13/23 (56%), Positives = 17/23 (73%)
Frame = +2
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
E+D+E DDEEDDD+ D +DE
Sbjct: 374 EEDDEDEEDDEEDDDEGEDDEDE 442
Score = 26.9 bits (58), Expect = 3.8
Identities = 14/23 (60%), Positives = 18/23 (77%)
Frame = +2
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
EDDEE+ DDE+++DD D DDE
Sbjct: 362 EDDEEE--DDEDEEDDEED-DDE 421
>TC91436 weakly similar to GP|21593499|gb|AAM65466.1 putative nodulin
protein N21 {Arabidopsis thaliana}, partial (16%)
Length = 909
Score = 30.0 bits (66), Expect = 0.45
Identities = 15/34 (44%), Positives = 19/34 (55%)
Frame = +3
Query: 82 YHYLHGYAYEDDEEQVVDDEEDDDDRCDLDDELV 115
Y L G + E D +V D +DDDD D DDE +
Sbjct: 480 YSVLWGKSKEVDNNKVEDATDDDDDDDDDDDEAI 581
>TC77681 similar to GP|22597156|gb|AAN03465.1 nucleolar histone deacetylase
HD2-P39 {Glycine max}, partial (47%)
Length = 1233
Score = 30.0 bits (66), Expect = 0.45
Identities = 30/115 (26%), Positives = 52/115 (45%), Gaps = 3/115 (2%)
Frame = +2
Query: 35 AEEEQEQEPDQ-NQQDAAPVEEIKIIHQIRPVLP-DPDAVAVVEIDRQQYHYLHGYAYED 92
AEE + EP + + ++AAP + ++ V P D D+ E D + D
Sbjct: 464 AEELKVSEPKKADAKNAAPAKNAATAKHVKVVDPKDEDSDDDDESDDE-------IGSSD 622
Query: 93 DEEQVVD-DEEDDDDRCDLDDELVPWEVGNKLGRQRMKKLGKRACSKMKNSKRSP 146
DE + D D ED+DD + D+E P + ++ ++ + K S K+ +P
Sbjct: 623 DEMENADSDSEDEDDSDEDDEEETPVKKVDQGKKRPNESASKTPISSKKSKNATP 787
>AW690594 similar to GP|23498163|emb hypothetical protein {Plasmodium
falciparum 3D7}, partial (10%)
Length = 633
Score = 30.0 bits (66), Expect = 0.45
Identities = 19/76 (25%), Positives = 34/76 (44%)
Frame = +3
Query: 37 EEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAYEDDEEQ 96
EE+E E D+ ++D EE E + ++ H E++EE+
Sbjct: 81 EEEEDEHDEEEEDEHDEEE-------------------EEEEEEEEEDEHDDEEEEEEEE 203
Query: 97 VVDDEEDDDDRCDLDD 112
++EEDDD+ + D+
Sbjct: 204 EEEEEEDDDEEGEEDE 251
>AJ502594 weakly similar to PIR|T05755|T057 hypothetical protein M4I22.120 -
Arabidopsis thaliana, partial (41%)
Length = 353
Score = 29.6 bits (65), Expect = 0.59
Identities = 10/27 (37%), Positives = 20/27 (74%)
Frame = +3
Query: 92 DDEEQVVDDEEDDDDRCDLDDELVPWE 118
+DE+ DD++DD + D ++++VPW+
Sbjct: 273 NDEDNDQDDDDDDTEDDDEENQVVPWK 353
>TC86045 similar to PIR|T46225|T46225 alpha NAC-like protein - Arabidopsis
thaliana, partial (69%)
Length = 1166
Score = 29.3 bits (64), Expect = 0.77
Identities = 17/50 (34%), Positives = 30/50 (60%), Gaps = 1/50 (2%)
Frame = +1
Query: 91 EDDEEQVVDDEEDDDDRCDLD-DELVPWEVGNKLGRQRMKKLGKRACSKM 139
+DD++ DD+EDDDD D D ++ + G+K + R +K ++A K+
Sbjct: 229 KDDDKDDEDDDEDDDDEDDDDKEDALAGADGSK--QSRSEKKSRKAMLKL 372
Score = 26.6 bits (57), Expect = 5.0
Identities = 11/23 (47%), Positives = 16/23 (68%)
Frame = +1
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
ED ++ DDE+DD+D D DD+
Sbjct: 220 EDVKDDDKDDEDDDEDDDDEDDD 288
>AL381331 weakly similar to GP|14209525|dbj contains ESTs C73715(E20247)
C99497(E20247)~similar to Arabidopsis thaliana
chromosome 2 At2g20670~, partial (14%)
Length = 475
Score = 28.9 bits (63), Expect = 1.0
Identities = 13/32 (40%), Positives = 20/32 (61%)
Frame = +2
Query: 92 DDEEQVVDDEEDDDDRCDLDDELVPWEVGNKL 123
+DEE +DDEED+D C ++ WE ++L
Sbjct: 107 NDEE--IDDEEDEDSVCTIEKNKAFWEEQDQL 196
>TC82470 homologue to GP|23497460|gb|AAN37003.1 hypothetical protein
{Plasmodium falciparum 3D7}, partial (8%)
Length = 1064
Score = 28.9 bits (63), Expect = 1.0
Identities = 10/24 (41%), Positives = 17/24 (70%)
Frame = +2
Query: 90 YEDDEEQVVDDEEDDDDRCDLDDE 113
Y+DD++ DD++DDDD D+ +
Sbjct: 32 YDDDDDDDDDDDDDDDDDDDMSTD 103
Score = 26.2 bits (56), Expect = 6.5
Identities = 10/24 (41%), Positives = 15/24 (61%)
Frame = +2
Query: 88 YAYEDDEEQVVDDEEDDDDRCDLD 111
Y +DD++ DD++DDDD D
Sbjct: 32 YDDDDDDDDDDDDDDDDDDDMSTD 103
>AW774261 weakly similar to EGAD|146423|156 vitellogenin {Anolis pulchellus},
partial (40%)
Length = 495
Score = 28.5 bits (62), Expect = 1.3
Identities = 19/81 (23%), Positives = 38/81 (46%)
Frame = +1
Query: 33 VDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAYED 92
+D EEE + E + + + VEE + + +I + + + A E E+
Sbjct: 289 IDLEEENDPEEEMEEIEYEEVEEEEEVEEIEEEVEENEEDAEEE--------------EE 426
Query: 93 DEEQVVDDEEDDDDRCDLDDE 113
+ E+ ++ E+DD +LDD+
Sbjct: 427 E*EKEEEEVEEDDTMQNLDDD 489
>BE248780 similar to GP|5929884|gb|A nucleolin-related protein NRP {Rattus
norvegicus}, partial (4%)
Length = 561
Score = 28.5 bits (62), Expect = 1.3
Identities = 13/35 (37%), Positives = 19/35 (54%)
Frame = +2
Query: 79 RQQYHYLHGYAYEDDEEQVVDDEEDDDDRCDLDDE 113
+Q H G +E ++E D +DDDD D D+E
Sbjct: 290 KQDDHIKEGGTHEIEDEDDDSDGDDDDDDDDYDEE 394
>TC83157 similar to GP|21595368|gb|AAM66095.1 unknown {Arabidopsis
thaliana}, partial (57%)
Length = 1108
Score = 28.5 bits (62), Expect = 1.3
Identities = 13/35 (37%), Positives = 19/35 (54%)
Frame = +3
Query: 79 RQQYHYLHGYAYEDDEEQVVDDEEDDDDRCDLDDE 113
+Q H G +E ++E D +DDDD D D+E
Sbjct: 297 KQDDHIKEGGTHEIEDEDDDSDGDDDDDDDDYDEE 401
>TC92637 similar to PIR|T39903|T39903 serine-rich protein - fission yeast
(Schizosaccharomyces pombe), partial (7%)
Length = 671
Score = 28.5 bits (62), Expect = 1.3
Identities = 17/47 (36%), Positives = 28/47 (59%)
Frame = +1
Query: 91 EDDEEQVVDDEEDDDDRCDLDDELVPWEVGNKLGRQRMKKLGKRACS 137
+DD E+V +DEE+D++ D DD ++ G K G R ++ K + S
Sbjct: 445 DDDAEEVDEDEEEDEEDYDDDD----YDGGGK-GSSRKRQYRKVSAS 570
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,165,862
Number of Sequences: 36976
Number of extensions: 68901
Number of successful extensions: 966
Number of sequences better than 10.0: 89
Number of HSP's better than 10.0 without gapping: 544
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 730
length of query: 167
length of database: 9,014,727
effective HSP length: 89
effective length of query: 78
effective length of database: 5,723,863
effective search space: 446461314
effective search space used: 446461314
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0258a.2


