

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0253.9
(677 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
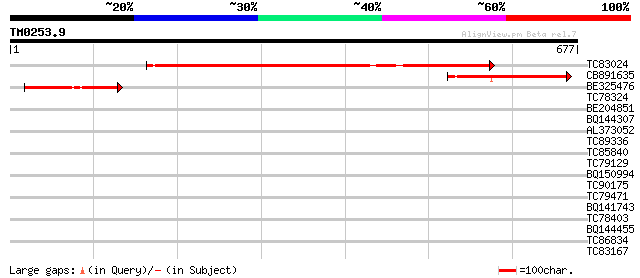
Score E
Sequences producing significant alignments: (bits) Value
TC83024 similar to PIR|T23875|T23875 hypothetical protein R03E1.... 416 e-116
CB891635 weakly similar to GP|5081545|gb|A CENPCB protein {Zea m... 142 3e-34
BE325476 114 9e-26
TC78324 similar to GP|8567798|gb|AAF76370.1| unknown protein {Ar... 33 0.27
BE204851 weakly similar to GP|14149112|db bg55 {Bruguiera gymnor... 32 0.61
BQ144307 weakly similar to GP|6523547|emb| hydroxyproline-rich g... 32 0.80
AL373052 similar to GP|21593014|gb| putative aldolase {Arabidops... 32 1.0
TC89336 weakly similar to GP|21554135|gb|AAM63215.1 unknown {Ara... 31 1.8
TC85840 similar to SP|P35016|ENPL_CATRO Endoplasmin homolog prec... 31 1.8
TC79129 similar to PIR|A96725|A96725 hypothetical protein F20P5.... 30 2.3
BQ150994 similar to GP|22122173|dbj preproneuropeptide B {Homo s... 30 2.3
TC90175 weakly similar to GP|6002488|gb|AAF00009.1| hypothetical... 30 3.0
TC79471 similar to GP|21536629|gb|AAM60961.1 seed maturation-lik... 29 6.7
BQ141743 homologue to GP|21112788|gb| CTP synthetase {Xanthomona... 29 6.7
TC78403 similar to PIR|E96603|E96603 unknown protein F14G9.26 [i... 29 6.7
BQ144455 homologue to PIR|B64826|B648 probable membrane protein ... 29 6.7
TC86834 similar to PIR|T46158|T46158 hypothetical protein T4D2.5... 28 8.8
TC83167 similar to GP|10177502|dbj|BAB10896. peroxidase ATP26a h... 28 8.8
>TC83024 similar to PIR|T23875|T23875 hypothetical protein R03E1.1 -
Caenorhabditis elegans, partial (1%)
Length = 1233
Score = 416 bits (1069), Expect = e-116
Identities = 225/418 (53%), Positives = 288/418 (68%), Gaps = 2/418 (0%)
Frame = +1
Query: 164 VSVEQNQDDVSTRPRQRRPGLRGNDQRSIKYRHRYSTEISDKNDHAMASQEAFGSDSLDP 223
VS + +QD S +PR+RRPGL G ++R ++YRHR STE ND ++SQEAF SD LDP
Sbjct: 10 VSSQLSQD--SNKPRERRPGLPGFNRRPVRYRHRVSTEALVNNDDVLSSQEAFESDGLDP 183
Query: 224 DTEKIENGEACLTSLENEVTDSSATEGNKICEILDGLLRCNSEDLEGDGAMDLLQERLQI 283
+ + G+ L SL+NEVTDS A E NK+ +IL GLL C+SE+LEG+GAM+LLQERLQ+
Sbjct: 184 VGDNTDKGKFSLASLDNEVTDSPAIEENKMNDILKGLLDCDSEELEGEGAMNLLQERLQV 363
Query: 284 KPVVLEKVSVPDFPDNQPIDLKSLRGSLSKPR--KALSNIDNLLNRRKNKTPLRQDAGSP 341
K +V EK+SVPDF D QPIDLKSL+G+LSKP KA S++DN L +TPLR+ G
Sbjct: 364 KSIVFEKLSVPDFLDIQPIDLKSLQGTLSKPSKGKAFSDVDNWLKGMNIQTPLRRSVGYA 543
Query: 342 AEQLASPTPPRSPFAPLSSLLNHISRLKPSVDPFSAHDIDHLSTTKYSPVHKMNQELNVV 401
+QLASPTPP+SPFA LSSL HISR K S DPFS H+ID + T YSP+H +QE+++V
Sbjct: 544 EKQLASPTPPKSPFASLSSLQKHISRSKLSTDPFSTHEIDLVPTRSYSPIHMADQEVDIV 723
Query: 402 GSAKPSNELNDHITKDAVAIGETNAVPDTLRNCASASENSKEDNSGKSSNKLDAPLIEDI 461
GS+K S+EL T+D +A GE N +P+T SENSKE NS S++++AP+IEDI
Sbjct: 724 GSSKLSDELTAPTTEDVIAAGEKNTIPET-------SENSKEHNSRNPSDEVNAPIIEDI 882
Query: 462 LAVSESCLIEDAVMNSTSTSQMPMEDNSMEPEFDANVDRNEPHADMDVDNGGSGMGETVM 521
+++ N T T Q M DNS EP F+ANVD NEP DMDVD G SGMG+ VM
Sbjct: 883 --------VDNPDRNCTITPQKSMVDNSTEPGFNANVDSNEPAVDMDVDIGRSGMGKRVM 1038
Query: 522 DDTVAKPNIEVNEQCQFEGKILTENTDAFTASMPTDDTDIDVVNPLADQSSPGVIQAN 579
DDT + N+E NE F+ L N FT+S+PTDD +++ PLADQS+P QAN
Sbjct: 1039DDTEGRQNVEPNEPFHFDEHTLERNMQGFTSSIPTDDANLNTELPLADQSNPVTYQAN 1212
>CB891635 weakly similar to GP|5081545|gb|A CENPCB protein {Zea mays},
partial (12%)
Length = 684
Score = 142 bits (359), Expect = 3e-34
Identities = 85/154 (55%), Positives = 106/154 (68%), Gaps = 5/154 (3%)
Frame = -3
Query: 523 DTVAKPNIEVNEQCQFEGKILTENTDAFTASMPTDDTDIDVVNPLADQSSPG---VIQAN 579
D V + N+E + Q E L EN FTAS+PTDD ++++V PLADQS+ QAN
Sbjct: 682 DIVGRQNVEP-QPYQSEDN-LPENMHEFTASLPTDDANLNLVIPLADQSNASNQDEHQAN 509
Query: 580 SIDKRTTCTNEGSEQCLQEERDGSRAPVE-QKRVKSRSQKDTKSKRLAK-RQSLAAAGTS 637
S+DKR+ +N+G E LQE GS APV QKRVK +QK +K K+ + R SLA AGTS
Sbjct: 508 SMDKRSGRSNDGPELSLQENTVGSVAPVNGQKRVKVCAQKVSKGKKREQHRMSLADAGTS 329
Query: 638 WTSGIRRSTRIRTRPLEYWKGERLVYGRIHQSKS 671
W SG+RRS R RTRPLEYWKGER+VYGR+H+S S
Sbjct: 328 WESGVRRSKRFRTRPLEYWKGERMVYGRVHESLS 227
>BE325476
Length = 394
Score = 114 bits (286), Expect = 9e-26
Identities = 68/120 (56%), Positives = 87/120 (71%), Gaps = 3/120 (2%)
Frame = +2
Query: 18 DPLSNYRGLSLFPSTSSALPLSVA-DDLDAIHNHLRSMALRSPAKLVEQTKAILEENPGL 76
DP++NY GLSLF ST S P S DLDAI+N+LRSM L SP +L EQ ++ILE N G
Sbjct: 41 DPIANYSGLSLFRSTFSLQPSSNPFHDLDAINNNLRSMDLGSPTRLAEQGQSILENNLG- 217
Query: 77 FNSQDDERNDVVDVDVDVAEDGQDFPRKRRPGLGLKRA--RFSLKPSTSVSLESLLPNLD 134
FN+++ + DV + DV E+G++FPRKRRPGLGL RA RFSLKP+ S+E L+P+LD
Sbjct: 218 FNTENLTQ-DVENDDVFAVEEGEEFPRKRRPGLGLNRARPRFSLKPTKKPSVEDLIPSLD 394
Score = 33.5 bits (75), Expect = 0.27
Identities = 27/94 (28%), Positives = 47/94 (49%), Gaps = 11/94 (11%)
Frame = +2
Query: 108 GLGLKRARFSLKPSTSV--SLESLLPNLDIENLKDPVEFFKAHERM---------ENARR 156
GL L R+ FSL+PS++ L+++ NL +L P + + + EN +
Sbjct: 62 GLSLFRSTFSLQPSSNPFHDLDAINNNLRSMDLGSPTRLAEQGQSILENNLGFNTENLTQ 241
Query: 157 EIQKQMGVSVEQNQDDVSTRPRQRRPGLRGNDQR 190
+++ +VE+ ++ PR+RRPGL N R
Sbjct: 242 DVENDDVFAVEEGEEF----PRKRRPGLGLNRAR 331
>TC78324 similar to GP|8567798|gb|AAF76370.1| unknown protein {Arabidopsis
thaliana}, complete
Length = 1108
Score = 33.5 bits (75), Expect = 0.27
Identities = 41/211 (19%), Positives = 82/211 (38%), Gaps = 17/211 (8%)
Frame = +1
Query: 160 KQMGVSVEQNQDDVS-TRPRQRRPGLRGNDQRSIKYRHRYSTEISDKNDHAMASQEAFGS 218
K++ V + + +D + TRP + G++ R +K + Y + M + F
Sbjct: 274 KKLDVELNRYKDQIKKTRPGPMQDGIKARAMRVLKQKRMY------EGQRDMLYNQTFNL 435
Query: 219 DSLDPDTEKIENGEACLTSLENEVTDSSATEGNKICEILDGLLRCNSEDLEGDGAMDLLQ 278
D + E I++ + +++L++ D + +D L D MDL+
Sbjct: 436 DQVQFAAEGIKDAQQTMSALKSANKDLKGMMKTVKIQDIDNLQ---------DEMMDLMD 588
Query: 279 ERLQIKPVVLEKVSVPDFPDNQPI--DLKSLRGSLSKPRKAL---------SNIDNLLNR 327
+I+ + SVPD D + + +L +L + + + S+ D+ LN
Sbjct: 589 VSNEIQETLGRSYSVPDDIDEEDLMGELDALEADMENESEGVPSYLQPDKESDFDSELNL 768
Query: 328 RKNKT-----PLRQDAGSPAEQLASPTPPRS 353
T P + ++L P PR+
Sbjct: 769 PSAPTGQTAVPHGRANAQTEDELGLPAVPRA 861
>BE204851 weakly similar to GP|14149112|db bg55 {Bruguiera gymnorrhiza},
partial (19%)
Length = 559
Score = 32.3 bits (72), Expect = 0.61
Identities = 19/56 (33%), Positives = 29/56 (50%)
Frame = +3
Query: 411 NDHITKDAVAIGETNAVPDTLRNCASASENSKEDNSGKSSNKLDAPLIEDILAVSE 466
ND A+ G T + + + NC S + ED K +N LD+ +I D+LA+ E
Sbjct: 288 NDRHDLKALRAG-TQLLSEFVNNCRSGMADLSEDTFIKLANLLDSEVIGDVLAIME 452
>BQ144307 weakly similar to GP|6523547|emb| hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (24%)
Length = 1059
Score = 32.0 bits (71), Expect = 0.80
Identities = 19/48 (39%), Positives = 23/48 (47%), Gaps = 4/48 (8%)
Frame = -2
Query: 347 SPTPPRSPFAPLSSLLNHISRLKPSV----DPFSAHDIDHLSTTKYSP 390
SPTPPR PF+PL+ H+SR P H TT Y+P
Sbjct: 746 SPTPPRPPFSPLTP---HLSRTAYHFI*PDSPGPNQTYGHFPTTTYTP 612
>AL373052 similar to GP|21593014|gb| putative aldolase {Arabidopsis
thaliana}, partial (19%)
Length = 459
Score = 31.6 bits (70), Expect = 1.0
Identities = 17/64 (26%), Positives = 29/64 (44%), Gaps = 4/64 (6%)
Frame = +3
Query: 333 PLRQDAGSPAEQLASPTPPRSPFAPLSSLLNHISRLKPSVDPFSAHDIDHL----STTKY 388
P + SP+ L+ +PP LSS N ++ P++ P S H + S ++
Sbjct: 267 PSTASSSSPSPPLSPKSPPSPATTSLSSTWNTVTAASPTLSPVSTHSPPQIPQQSSVSQK 446
Query: 389 SPVH 392
P+H
Sbjct: 447 PPLH 458
>TC89336 weakly similar to GP|21554135|gb|AAM63215.1 unknown {Arabidopsis
thaliana}, partial (7%)
Length = 1241
Score = 30.8 bits (68), Expect = 1.8
Identities = 48/214 (22%), Positives = 89/214 (41%), Gaps = 15/214 (7%)
Frame = +1
Query: 71 EENPGLFNSQDDERNDVVDVDVDVAEDGQDFPRKRRPGLGLKRARFSLKPSTSVSLESLL 130
EE L N + + V ++ + + K+ L L++ + LE+ +
Sbjct: 541 EEIIKLKNQNEKSEGQLDSVQKELTLNMDELEHKKGQVLELQKQK--------AELETHV 696
Query: 131 PNLDIENLKDPVEFFK-AHERMENARREIQ------KQMGVSVEQNQDDVSTRPRQRRPG 183
PNL +E L+ E K +++ + R+E++ +Q+ +E Q++V+ Q G
Sbjct: 697 PNL-VEQLEVANEHLKISNDEVARLRKELESNLAETRQLQDQLEVAQENVTKLEWQLDSG 873
Query: 184 ---LRGNDQRSIKYRHRYST---EISDKND--HAMASQEAFGSDSLDPDTEKIENGEACL 235
+R + R ++ + E+ D H + +Q +F D L D + + L
Sbjct: 874 RKQIRELEDRITWFKTNETNLEVEVQKLKDEMHDVQAQFSFEKDQLHSDIASLSEIKTQL 1053
Query: 236 TSLENEVTDSSATEGNKICEILDGLLRCNSEDLE 269
TS E S + NK L RC +E+LE
Sbjct: 1054TSTLEEWESRSHSLANK-------LRRCEAENLE 1134
>TC85840 similar to SP|P35016|ENPL_CATRO Endoplasmin homolog precursor (GRP94
homolog). [Rosy periwinkle Madagascar periwinkle],
partial (92%)
Length = 2791
Score = 30.8 bits (68), Expect = 1.8
Identities = 17/61 (27%), Positives = 33/61 (53%)
Frame = +1
Query: 33 SSALPLSVADDLDAIHNHLRSMALRSPAKLVEQTKAILEENPGLFNSQDDERNDVVDVDV 92
S LPL+V+ ++ H+ L+++ + K ++ + + EE+P S D E+ + DV
Sbjct: 1393 SDTLPLNVSREMLQQHSSLKTIKKKLIRKALDMIRRLAEEDPD--ESTDREKKEETSSDV 1566
Query: 93 D 93
D
Sbjct: 1567 D 1569
>TC79129 similar to PIR|A96725|A96725 hypothetical protein F20P5.7
[imported] - Arabidopsis thaliana, partial (64%)
Length = 1328
Score = 30.4 bits (67), Expect = 2.3
Identities = 16/46 (34%), Positives = 23/46 (49%)
Frame = -3
Query: 404 AKPSNELNDHITKDAVAIGETNAVPDTLRNCASASENSKEDNSGKS 449
+KP LN K A+ ++A D NC E+ E++SGKS
Sbjct: 318 SKPGTNLNSCSMKTAIDSSSSSANNDDDGNCDDDDESRSEEHSGKS 181
>BQ150994 similar to GP|22122173|dbj preproneuropeptide B {Homo sapiens},
partial (15%)
Length = 1036
Score = 30.4 bits (67), Expect = 2.3
Identities = 24/83 (28%), Positives = 38/83 (44%)
Frame = +3
Query: 578 ANSIDKRTTCTNEGSEQCLQEERDGSRAPVEQKRVKSRSQKDTKSKRLAKRQSLAAAGTS 637
AN+ KRTT + + R+ SRA E++R ++ + K R K+++ G
Sbjct: 252 ANTNTKRTTPHGHRQRKHTRRRRETSRAEGEREREGTKKEARRKRAR-RKKETPTPDGPR 428
Query: 638 WTSGIRRSTRIRTRPLEYWKGER 660
GI R R TR E + E+
Sbjct: 429 PRRGITRPERQATRRAERRRAEK 497
>TC90175 weakly similar to GP|6002488|gb|AAF00009.1| hypothetical protein
{Penicillium chrysogenum}, partial (87%)
Length = 1290
Score = 30.0 bits (66), Expect = 3.0
Identities = 21/77 (27%), Positives = 29/77 (37%)
Frame = +2
Query: 584 RTTCTNEGSEQCLQEERDGSRAPVEQKRVKSRSQKDTKSKRLAKRQSLAAAGTSWTSGIR 643
R C ++C + G RA R + R+ + R A R+S AAG + R
Sbjct: 518 RRPCPRRDRDRC-RRRGSGRRAAARGWRRRRRASAGRRGPRTAGRRSRRAAGRGRSPRPR 694
Query: 644 RSTRIRTRPLEYWKGER 660
R R W G R
Sbjct: 695 RPPPCRAGRFRAWTGRR 745
>TC79471 similar to GP|21536629|gb|AAM60961.1 seed maturation-like protein
{Arabidopsis thaliana}, partial (50%)
Length = 1464
Score = 28.9 bits (63), Expect = 6.7
Identities = 18/61 (29%), Positives = 33/61 (53%)
Frame = +1
Query: 340 SPAEQLASPTPPRSPFAPLSSLLNHISRLKPSVDPFSAHDIDHLSTTKYSPVHKMNQELN 399
+P+ ASPT + + S L+ S + P+VDPFS+ + ++T+ + ++NQ
Sbjct: 109 NPSSSHASPTTHLTHLSTTLSPLSSSSAVPPTVDPFSSR-LPVTTSTETTLFQRVNQRST 285
Query: 400 V 400
V
Sbjct: 286 V 288
>BQ141743 homologue to GP|21112788|gb| CTP synthetase {Xanthomonas campestris
pv. campestris str. ATCC 33913}, partial (3%)
Length = 1303
Score = 28.9 bits (63), Expect = 6.7
Identities = 16/55 (29%), Positives = 28/55 (50%), Gaps = 3/55 (5%)
Frame = -1
Query: 599 ERDGSRAPVEQKRVKSRSQKDTKSKRL---AKRQSLAAAGTSWTSGIRRSTRIRT 650
E + R+ +K V + +++ + R A RQ+ A GT++ G+R IRT
Sbjct: 439 EEEKKRSETGEKTVLGKKKQEGRKARRPSRANRQNTAGRGTNYEGGVRAIISIRT 275
>TC78403 similar to PIR|E96603|E96603 unknown protein F14G9.26 [imported] -
Arabidopsis thaliana, partial (19%)
Length = 999
Score = 28.9 bits (63), Expect = 6.7
Identities = 18/51 (35%), Positives = 25/51 (48%), Gaps = 9/51 (17%)
Frame = -1
Query: 325 LNRRKNKTPLRQDAGSPAEQLASPTPPRSPF---------APLSSLLNHIS 366
++ R ++TP + SP E+L P PP PF PLSS L +S
Sbjct: 600 MSLRMSRTPRIPKSTSPPEKLPPPPPPLPPFPSSSRGGNPGPLSSSLIFLS 448
>BQ144455 homologue to PIR|B64826|B648 probable membrane protein ybjE -
Escherichia coli, partial (28%)
Length = 794
Score = 28.9 bits (63), Expect = 6.7
Identities = 13/38 (34%), Positives = 20/38 (52%)
Frame = +3
Query: 327 RRKNKTPLRQDAGSPAEQLASPTPPRSPFAPLSSLLNH 364
+R +K P+R A P + ++P PRS P+S H
Sbjct: 657 QRHDKYPVRNTATPPIPECSAPFVPRSILVPISPRARH 770
>TC86834 similar to PIR|T46158|T46158 hypothetical protein T4D2.50 -
Arabidopsis thaliana, partial (77%)
Length = 1222
Score = 28.5 bits (62), Expect = 8.8
Identities = 36/162 (22%), Positives = 60/162 (36%), Gaps = 6/162 (3%)
Frame = +1
Query: 340 SPAEQLASPTPPRSPFAPLSSLLNHISRLKPSVDPFSAHDIDHLSTTKYSPVHKMNQELN 399
+PA ASP PP S P + S SVD ++ L + K Q LN
Sbjct: 271 TPASSSASPRPPSSHVPPAEAAGTIASLKDKSVD-----ELRKLLSDK----DAYQQFLN 423
Query: 400 VVGSAKPSNELNDHITKDAVAIGETNAVPD----TLRNCASASENSKEDNSGKSSNKLDA 455
+ K L D + K+ + E N + LRN ++ + + N+L+
Sbjct: 424 SLEQVKIQTNLKDELAKENRQLAEENLQKEPRMMELRNQCRIIRTTELATANEKLNELEK 603
Query: 456 PLIEDILAVSESCLIE--DAVMNSTSTSQMPMEDNSMEPEFD 495
E + S + L++ +N T + ++ E D
Sbjct: 604 QKEEMLKMNSPASLLQRIQESVNQTDEESENLHQQLLDREVD 729
>TC83167 similar to GP|10177502|dbj|BAB10896. peroxidase ATP26a homolog
{Arabidopsis thaliana}, partial (17%)
Length = 429
Score = 28.5 bits (62), Expect = 8.8
Identities = 15/41 (36%), Positives = 20/41 (48%)
Frame = +2
Query: 323 NLLNRRKNKTPLRQDAGSPAEQLASPTPPRSPFAPLSSLLN 363
N N + PL + + A L + TPP S PLS+ LN
Sbjct: 188 NRSNLQPPLPPLSVSSSTTASSLTAATPPSSSLQPLSTKLN 310
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.309 0.128 0.354
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,856,970
Number of Sequences: 36976
Number of extensions: 208156
Number of successful extensions: 1132
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 1111
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1123
length of query: 677
length of database: 9,014,727
effective HSP length: 103
effective length of query: 574
effective length of database: 5,206,199
effective search space: 2988358226
effective search space used: 2988358226
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0253.9


