

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0252a.5
(150 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
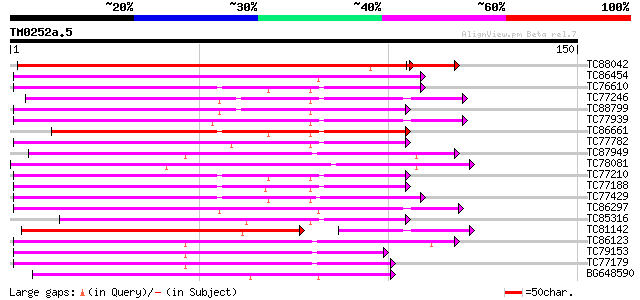
Score E
Sequences producing significant alignments: (bits) Value
TC88042 similar to GP|23297181|gb|AAN12515.1 phosphoenolpyruvate... 156 1e-39
TC86454 homologue to GP|4567091|gb|AAD23582.1| SNF-1-like serine... 89 7e-19
TC76610 similar to PIR|T50802|T50802 serine/threonine protein ki... 87 3e-18
TC77246 similar to PIR|C71408|C71408 probable protein kinase - A... 84 2e-17
TC88799 similar to PIR|C71408|C71408 probable protein kinase - A... 83 4e-17
TC77939 similar to GP|14194167|gb|AAK56278.1 At2g26980/T20P8.3 {... 81 2e-16
TC86661 similar to GP|9798599|dbj|BAB11737.1 serine/threonine pr... 81 2e-16
TC77782 similar to GP|13448037|gb|AAK26845.1 SOS2-like protein k... 81 2e-16
TC87949 similar to GP|16904224|gb|AAL30819.1 calcium/calmodulin-... 80 2e-16
TC78081 similar to GP|16904226|gb|AAL30820.1 calcium/calmodulin-... 80 3e-16
TC77210 similar to GP|21954717|gb|AAM83095.1 SOS2-like protein k... 80 3e-16
TC77188 similar to PIR|T04862|T04862 probable serine/threonine-s... 79 9e-16
TC77429 similar to GP|15215664|gb|AAK91377.1 AT5g25110/T11H3_120... 78 1e-15
TC86297 similar to GP|15912283|gb|AAL08275.1 At1g30270/F12P21_6 ... 78 1e-15
TC85316 weakly similar to PIR|E84707|E84707 probable protein kin... 78 1e-15
TC81142 similar to GP|8978255|dbj|BAA98146.1 serine/threonine pr... 69 1e-15
TC86123 homologue to PIR|S71770|S71770 calcium-dependent protein... 78 2e-15
TC79153 similar to SP|P28583|CDPK_SOYBN Calcium-dependent protei... 78 2e-15
TC77179 similar to GP|2665890|gb|AAB88537.1| calcium-dependent p... 77 2e-15
BG648590 similar to PIR|H96771|H96 hypothetical protein F1M20.1 ... 77 3e-15
>TC88042 similar to GP|23297181|gb|AAN12515.1 phosphoenolpyruvate
carboxylase kinase {Glycine max}, partial (97%)
Length = 1237
Score = 156 bits (395), Expect(2) = 1e-39
Identities = 76/124 (61%), Positives = 91/124 (73%), Gaps = 19/124 (15%)
Frame = +3
Query: 3 PIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGRR 62
P+ E + A+++K+LLEA+ HCHRLGVAH D+KPDN+LF + +LKLADFGSAEWFGDG +
Sbjct: 339 PLTEQQAAAIIKKLLEAVVHCHRLGVAHRDIKPDNILFDSEDNLKLADFGSAEWFGDGEK 518
Query: 63 MSGVVGTPYYVAPEVLMGWENGEKVDVWSCGV-------------------IFEAVIWGN 103
MSGVVGTPYYVAPEVL+G + EKVDVWSCGV IFE+VI N
Sbjct: 519 MSGVVGTPYYVAPEVLLGRDYTEKVDVWSCGVLLYIMLSGIPPFYGDSTAEIFESVIRAN 698
Query: 104 LRFP 107
LRFP
Sbjct: 699 LRFP 710
Score = 22.3 bits (46), Expect(2) = 1e-39
Identities = 11/14 (78%), Positives = 12/14 (85%)
Frame = +1
Query: 106 FPLESSGMFLLLLR 119
F LESS +FLLLLR
Sbjct: 706 FLLESSVLFLLLLR 747
>TC86454 homologue to GP|4567091|gb|AAD23582.1| SNF-1-like serine/threonine
protein kinase {Glycine max}, partial (90%)
Length = 2194
Score = 89.0 bits (219), Expect = 7e-19
Identities = 45/110 (40%), Positives = 65/110 (58%), Gaps = 1/110 (0%)
Frame = +3
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G + E E S +Q++ + +CHR V H D+KP+N+L + +K+ADFG + DG
Sbjct: 393 GRLQEDEARSFFQQIISGVEYCHRNMVVHRDLKPENLLLDSKWSVKIADFGLSNIMRDGH 572
Query: 62 RMSGVVGTPYYVAPEVLMG-WENGEKVDVWSCGVIFEAVIWGNLRFPLES 110
+ G+P Y APEV+ G G +VDVWSCGVI A++ G L F E+
Sbjct: 573 FLKTSCGSPNYAAPEVISGKLYAGPEVDVWSCGVILYALLCGTLPFDDEN 722
>TC76610 similar to PIR|T50802|T50802 serine/threonine protein kinase-like
protein - Arabidopsis thaliana, partial (73%)
Length = 1799
Score = 87.0 bits (214), Expect = 3e-18
Identities = 50/115 (43%), Positives = 69/115 (59%), Gaps = 6/115 (5%)
Frame = +2
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G + E +QL+ A+ CH GV H D+KP+N+L DLK++DFG + D R
Sbjct: 602 GKMEENIARKYFQQLISAVDFCHSRGVTHRDLKPENLLLDENEDLKVSDFGLSA-LPDQR 778
Query: 62 RMSGVV----GTPYYVAPEVL--MGWENGEKVDVWSCGVIFEAVIWGNLRFPLES 110
R G++ GTP YVAPEVL +G++ G K D+WSCGVI A++ G L F E+
Sbjct: 779 RSDGMLLTPCGTPAYVAPEVLKKIGYD-GSKADIWSCGVILFALLCGYLPFQGEN 940
>TC77246 similar to PIR|C71408|C71408 probable protein kinase - Arabidopsis
thaliana, partial (65%)
Length = 1715
Score = 84.0 bits (206), Expect = 2e-17
Identities = 49/122 (40%), Positives = 72/122 (58%), Gaps = 5/122 (4%)
Frame = +3
Query: 5 PEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSA---EWFGDGR 61
PE +QL+ A+ CHR G+AH D+KP N+L A G+LK++DFG + E +G
Sbjct: 378 PEALARRYFQQLVSALCFCHRNGIAHRDLKPQNLLLDAEGNLKVSDFGLSALPEQLKNG- 554
Query: 62 RMSGVVGTPYYVAPEVL--MGWENGEKVDVWSCGVIFEAVIWGNLRFPLESSGMFLLLLR 119
+ GTP Y APE+L +G++ G K D WSCGVI ++ G+L P + S + + R
Sbjct: 555 LLHTACGTPAYTAPEILGRIGYD-GSKADAWSCGVILYVLLAGHL--PFDDSNIPAMCKR 725
Query: 120 IS 121
I+
Sbjct: 726 IT 731
>TC88799 similar to PIR|C71408|C71408 probable protein kinase - Arabidopsis
thaliana, partial (57%)
Length = 1648
Score = 83.2 bits (204), Expect = 4e-17
Identities = 47/110 (42%), Positives = 62/110 (55%), Gaps = 5/110 (4%)
Frame = +1
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSA---EWFG 58
G E +QL+ A+ CHR GVAH D+KP N+L A G+LK++DFG + E
Sbjct: 412 GRFSESTARFYFQQLVSALRFCHRNGVAHRDLKPQNLLLDADGNLKVSDFGLSALPEHLK 591
Query: 59 DGRRMSGVVGTPYYVAPEVL--MGWENGEKVDVWSCGVIFEAVIWGNLRF 106
DG + GTP Y APE+L G +G K D WSCG+I ++ G L F
Sbjct: 592 DG-LLHTACGTPAYTAPEILRRSGGYDGSKADAWSCGLILYVLLAGYLPF 738
>TC77939 similar to GP|14194167|gb|AAK56278.1 At2g26980/T20P8.3 {Arabidopsis
thaliana}, partial (70%)
Length = 1353
Score = 80.9 bits (198), Expect = 2e-16
Identities = 48/125 (38%), Positives = 70/125 (55%), Gaps = 5/125 (4%)
Frame = +1
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFG---SAEWFG 58
G + E E +QL+ + +CH GV H D+KP+N+L A G+LK++DFG ++
Sbjct: 832 GRMGEPEARRYFQQLINVVDYCHSRGVYHRDLKPENLLLDAYGNLKVSDFGLSALSQQVR 1011
Query: 59 DGRRMSGVVGTPYYVAPEVL--MGWENGEKVDVWSCGVIFEAVIWGNLRFPLESSGMFLL 116
D + GTP YVAPEVL G++ G D+WSCGVI ++ G L P + + L
Sbjct: 1012DDGLLHTTCGTPNYVAPEVLNDRGYD-GATADLWSCGVILFVLVAGYL--PFDDPNLMEL 1182
Query: 117 LLRIS 121
+IS
Sbjct: 1183YKKIS 1197
>TC86661 similar to GP|9798599|dbj|BAB11737.1 serine/threonine protein
kinase {Arabidopsis thaliana}, partial (58%)
Length = 1840
Score = 80.9 bits (198), Expect = 2e-16
Identities = 44/101 (43%), Positives = 66/101 (64%), Gaps = 6/101 (5%)
Frame = +2
Query: 12 LMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGRRMSGVV---- 67
L +QL+ A+ +CH GV H D+KP+N+L G+LK++DFG + + R+ G++
Sbjct: 614 LFQQLISAVGYCHSHGVFHRDLKPENLLLDDKGNLKVSDFGLSA-VKEQVRIDGMLHTLC 790
Query: 68 GTPYYVAPEVL--MGWENGEKVDVWSCGVIFEAVIWGNLRF 106
GTP YVAPE+L G++ G KVD+WSCG+I ++ G L F
Sbjct: 791 GTPAYVAPEILAKRGYD-GAKVDIWSCGIILFVLVAGYLPF 910
>TC77782 similar to GP|13448037|gb|AAK26845.1 SOS2-like protein kinase PKS6
{Arabidopsis thaliana}, partial (77%)
Length = 1236
Score = 80.9 bits (198), Expect = 2e-16
Identities = 45/108 (41%), Positives = 63/108 (57%), Gaps = 3/108 (2%)
Frame = +2
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWF-GDG 60
G + E E S +QL+ A+ +CH GV H D+KP+N+L G LK++DFG + + +
Sbjct: 470 GRLKEDEARSYFQQLINAVDYCHSRGVYHRDLKPENLLLDTNGVLKVSDFGLSTYSQQED 649
Query: 61 RRMSGVVGTPYYVAPEVL--MGWENGEKVDVWSCGVIFEAVIWGNLRF 106
+ GTP YVAPEVL G+ G D+WSCGVI ++ G L F
Sbjct: 650 ELLRTACGTPNYVAPEVLNDRGYV-GSSSDIWSCGVILFVLMAGYLPF 790
>TC87949 similar to GP|16904224|gb|AAL30819.1 calcium/calmodulin-dependent
protein kinase CaMK2 {Nicotiana tabacum}, partial (62%)
Length = 1413
Score = 80.5 bits (197), Expect = 2e-16
Identities = 45/118 (38%), Positives = 69/118 (58%), Gaps = 4/118 (3%)
Frame = +1
Query: 6 EVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGD---LKLADFGSAEWFGDGRR 62
E + A +++Q+L+ CH G+ H D+KP+N LF + + LK DFG +++ G+R
Sbjct: 145 EKDAAVVVRQMLKVAAQCHLHGLVHRDMKPENFLFKSNKEDSALKATDFGLSDFIKPGKR 324
Query: 63 MSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWGNLRF-PLESSGMFLLLLR 119
+VG+ YYVAPEVL ++G + DVWS GVI ++ G F G+F +LR
Sbjct: 325 FQDIVGSAYYVAPEVLKR-KSGPESDVWSIGVITYILLCGRRPFWDKTEDGIFKEVLR 495
>TC78081 similar to GP|16904226|gb|AAL30820.1 calcium/calmodulin-dependent
protein kinase CaMK3 {Nicotiana tabacum}, partial (58%)
Length = 1066
Score = 80.1 bits (196), Expect = 3e-16
Identities = 47/127 (37%), Positives = 70/127 (55%), Gaps = 4/127 (3%)
Frame = +2
Query: 1 GGPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLF---GAGGDLKLADFGSAEWF 57
GG E + ++M Q+L + CH GV H D+KP+N L+ +LK DFG +++
Sbjct: 221 GGKYSEDDAKAVMVQILNVVAFCHLQGVVHRDLKPENFLYTTKDESSELKAIDFGLSDFV 400
Query: 58 GDGRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWGNLRF-PLESSGMFLL 116
R++ +VG+ YYVAPEVL + E DVWS GVI ++ G+ F SG+F
Sbjct: 401 RPDERLNDIVGSAYYVAPEVLHRSYSTE-ADVWSIGVIAYILLCGSRPFWARTESGIFRA 577
Query: 117 LLRIS*G 123
+L+ G
Sbjct: 578 VLKADPG 598
>TC77210 similar to GP|21954717|gb|AAM83095.1 SOS2-like protein kinase
{Glycine max}, partial (88%)
Length = 1831
Score = 80.1 bits (196), Expect = 3e-16
Identities = 45/111 (40%), Positives = 65/111 (58%), Gaps = 6/111 (5%)
Frame = +1
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G + E +QL+ A+ CH GV H D+KP+N+L G+LK++DFG F +
Sbjct: 631 GRLKEDVARVYFQQLISAVDFCHSRGVYHRDLKPENLLLDEDGNLKVSDFGLCT-FSEHL 807
Query: 62 RMSGVV----GTPYYVAPEVL--MGWENGEKVDVWSCGVIFEAVIWGNLRF 106
R G++ GTP YV+PEV+ G++ G K D+WSCGVI ++ G L F
Sbjct: 808 RQDGLLHTTCGTPAYVSPEVIAKKGYD-GAKADIWSCGVILYVLLAGFLPF 957
>TC77188 similar to PIR|T04862|T04862 probable serine/threonine-specific
protein kinase (EC 2.7.1.-) F28A21.110 - Arabidopsis
thaliana, partial (79%)
Length = 1881
Score = 78.6 bits (192), Expect = 9e-16
Identities = 46/111 (41%), Positives = 65/111 (58%), Gaps = 6/111 (5%)
Frame = +1
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G + E +QL+ A+ CH GV H D+KP+N+L G+LK++DFG + D
Sbjct: 466 GRLKEEVARKYFQQLICAVGFCHARGVFHRDLKPENLLLDEKGNLKVSDFGLSA-VSDEI 642
Query: 62 RMSGV----VGTPYYVAPEVL--MGWENGEKVDVWSCGVIFEAVIWGNLRF 106
+ G+ GTP YVAPEVL G++ G KVD+WSCGV+ ++ G L F
Sbjct: 643 KQDGLFHTFCGTPAYVAPEVLSRKGYD-GAKVDIWSCGVVLFVLMAGYLPF 792
>TC77429 similar to GP|15215664|gb|AAK91377.1 AT5g25110/T11H3_120
{Arabidopsis thaliana}, partial (79%)
Length = 1869
Score = 78.2 bits (191), Expect = 1e-15
Identities = 46/115 (40%), Positives = 67/115 (58%), Gaps = 6/115 (5%)
Frame = +3
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G E +QL+ A+ +CH GV+H D+KP+N+L +LK++DFG + +
Sbjct: 546 GKFKEELARKYFQQLISAVDYCHSRGVSHRDLKPENLLLDENENLKVSDFGLSA-LPEHL 722
Query: 62 RMSGVV----GTPYYVAPEVL--MGWENGEKVDVWSCGVIFEAVIWGNLRFPLES 110
R G++ GTP YVAPEV+ G+ +G K D WSCGVI A++ G L F E+
Sbjct: 723 RQDGLLHTQCGTPAYVAPEVVRKRGY-SGFKADTWSCGVILYALLAGFLPFQHEN 884
>TC86297 similar to GP|15912283|gb|AAL08275.1 At1g30270/F12P21_6
{Arabidopsis thaliana}, partial (90%)
Length = 2043
Score = 78.2 bits (191), Expect = 1e-15
Identities = 45/123 (36%), Positives = 66/123 (53%), Gaps = 4/123 (3%)
Frame = +2
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSA---EWFG 58
G + E E +QL+ A+ +CH GV H D+KP+N+L G LK++DFG + +
Sbjct: 728 GRLKEDEARKYFQQLICAVDYCHSRGVCHRDLKPENLLLDTNGTLKVSDFGLSALPQQVR 907
Query: 59 DGRRMSGVVGTPYYVAPEVLMG-WENGEKVDVWSCGVIFEAVIWGNLRFPLESSGMFLLL 117
+ + GTP YVAPEV+ +G D+WSCGVI ++ G L P E + L
Sbjct: 908 EDGLLHTTCGTPNYVAPEVIQNKGYDGAIADLWSCGVILFVLMAGYL--PFEEDNLVALY 1081
Query: 118 LRI 120
+I
Sbjct: 1082KKI 1090
>TC85316 weakly similar to PIR|E84707|E84707 probable protein kinase
[imported] - Arabidopsis thaliana, partial (66%)
Length = 1528
Score = 78.2 bits (191), Expect = 1e-15
Identities = 45/98 (45%), Positives = 57/98 (57%), Gaps = 5/98 (5%)
Frame = +1
Query: 14 KQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR---RMSGVVGTP 70
+QL+ A+ HCH GV H D+K DN+L +LK+ DFG + R + V GTP
Sbjct: 580 RQLISAVKHCHSRGVFHRDLKLDNLLLDENDNLKVTDFGLSAVKNQIRPDGLLHTVCGTP 759
Query: 71 YYVAPEVL--MGWENGEKVDVWSCGVIFEAVIWGNLRF 106
YVAPE+L G+E G K DVWSCGV+ V G L F
Sbjct: 760 SYVAPEILAKKGYE-GAKADVWSCGVVLFTVTAGYLPF 870
>TC81142 similar to GP|8978255|dbj|BAA98146.1 serine/threonine protein
kinase SOS2 {Arabidopsis thaliana}, partial (54%)
Length = 784
Score = 68.6 bits (166), Expect(2) = 1e-15
Identities = 34/76 (44%), Positives = 48/76 (62%), Gaps = 1/76 (1%)
Frame = +1
Query: 4 IPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDG-RR 62
+ E E +QL++A+ HCH+ GV H D+KP+N+L A G+LK++DFG + G
Sbjct: 304 LSENESRRYFQQLIDAVAHCHKKGVYHRDLKPENLLLDAYGNLKVSDFGLSALTKQGDEL 483
Query: 63 MSGVVGTPYYVAPEVL 78
+ GTP YVAPEVL
Sbjct: 484 LHTTCGTPNYVAPEVL 531
Score = 29.6 bits (65), Expect(2) = 1e-15
Identities = 16/36 (44%), Positives = 21/36 (57%)
Frame = +2
Query: 88 DVWSCGVIFEAVIWGNLRFPLESSGMFLLLLRIS*G 123
DVWSCG+I ++ G L P E + + L RIS G
Sbjct: 563 DVWSCGIILYVLMAGYL--PFEEADLPTLFRRISAG 664
>TC86123 homologue to PIR|S71770|S71770 calcium-dependent protein kinase (EC
2.7.1.-) - mung bean, complete
Length = 2448
Score = 77.8 bits (190), Expect = 2e-15
Identities = 48/122 (39%), Positives = 69/122 (56%), Gaps = 4/122 (3%)
Frame = +3
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGD---LKLADFGSAEWFG 58
G E + A L + ++ + CH LGV H D+KP+N L D LK DFG + +F
Sbjct: 816 GHYTERKAAELTRIIVGVVEACHSLGVMHRDLKPENFLLVNKDDDFSLKAIDFGLSVFFK 995
Query: 59 DGRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWGNLRFPLES-SGMFLLL 117
G+ + VVG+PYYVAPEVL+ G + DVW+ GVI ++ G F E+ G+F +
Sbjct: 996 PGQVFTDVVGSPYYVAPEVLLK-HYGPEADVWTAGVILYILLSGVPPFWAETQQGIFDAV 1172
Query: 118 LR 119
L+
Sbjct: 1173LK 1178
>TC79153 similar to SP|P28583|CDPK_SOYBN Calcium-dependent protein kinase
SK5 (EC 2.7.1.-) (CDPK). [Soybean] {Glycine max},
partial (95%)
Length = 1751
Score = 77.8 bits (190), Expect = 2e-15
Identities = 43/102 (42%), Positives = 59/102 (57%), Gaps = 3/102 (2%)
Frame = +2
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGD---LKLADFGSAEWFG 58
G E + A L++ ++E + CH LGV H D+KP+N LF + + LK DFG + ++
Sbjct: 431 GHYSERQAAKLIRTIVEVVEACHSLGVMHRDLKPENFLFDSVDEDALLKTIDFGLSVFYK 610
Query: 59 DGRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVI 100
G S VVG+PYYVAPEVL G + DVWS F +I
Sbjct: 611 PGEIFSDVVGSPYYVAPEVLHK-HYGPEADVWSAWCYFVHLI 733
>TC77179 similar to GP|2665890|gb|AAB88537.1| calcium-dependent protein
kinase {Fragaria x ananassa}, partial (92%)
Length = 2243
Score = 77.4 bits (189), Expect = 2e-15
Identities = 40/104 (38%), Positives = 61/104 (58%), Gaps = 3/104 (2%)
Frame = +1
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGD---LKLADFGSAEWFG 58
G E A+++K +++ + CH GV H D+KP+N LF + LK DFG + F
Sbjct: 676 GHYTERAAATVVKTIVQVVQMCHEHGVMHRDLKPENFLFANKKETSPLKAIDFGLSITFK 855
Query: 59 DGRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWG 102
G + + +VG+PYY+APEVL G ++D+WS GVI ++ G
Sbjct: 856 PGDKFNEIVGSPYYMAPEVLKR-NYGPEIDIWSAGVILYILLCG 984
>BG648590 similar to PIR|H96771|H96 hypothetical protein F1M20.1 [imported] -
Arabidopsis thaliana, partial (40%)
Length = 805
Score = 76.6 bits (187), Expect = 3e-15
Identities = 40/99 (40%), Positives = 59/99 (59%), Gaps = 3/99 (3%)
Frame = +1
Query: 7 VEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGRR--MS 64
V++ MKQLL + HCH GV H D+K N+L G LK+ADFG A + G++ ++
Sbjct: 82 VQIKCYMKQLLSGLEHCHSRGVMHRDIKGSNLLVNNEGILKVADFGLANFTNSGKKQPLT 261
Query: 65 GVVGTPYYVAPEVLMG-WENGEKVDVWSCGVIFEAVIWG 102
V T +Y PE+L+G + G VD+WS G +F ++ G
Sbjct: 262 SRVVTLWYRPPELLLGSTDYGPSVDLWSVGCVFAELLVG 378
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.334 0.150 0.505
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,240,810
Number of Sequences: 36976
Number of extensions: 69194
Number of successful extensions: 1186
Number of sequences better than 10.0: 474
Number of HSP's better than 10.0 without gapping: 953
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 974
length of query: 150
length of database: 9,014,727
effective HSP length: 87
effective length of query: 63
effective length of database: 5,797,815
effective search space: 365262345
effective search space used: 365262345
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0252a.5


