

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0248.6
(573 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
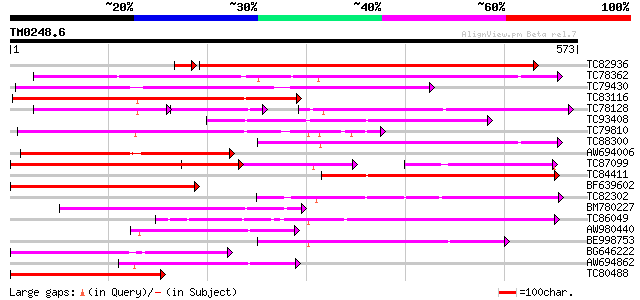
Score E
Sequences producing significant alignments: (bits) Value
TC82936 similar to PIR|G84829|G84829 probable PTR2 family peptid... 431 e-127
TC78362 similar to GP|15391731|emb|CAA93316. nitrite transporter... 343 1e-94
TC79430 similar to PIR|F86358|F86358 Similar to peptide transpor... 291 6e-79
TC83116 similar to GP|11933407|dbj|BAB19758. putative nitrate tr... 288 3e-78
TC78128 weakly similar to GP|21928025|gb|AAM78041.1 At1g69870/T1... 166 8e-73
TC93408 similar to GP|11933407|dbj|BAB19758. putative nitrate tr... 220 1e-57
TC79810 similar to GP|11994403|dbj|BAB02362. nitrate transporter... 204 7e-53
TC88300 similar to GP|9581817|emb|CAC00544.1 putative low-affini... 200 1e-51
AW694006 similar to PIR|F86358|F86 Similar to peptide transporte... 178 4e-45
TC87099 similar to PIR|G86449|G86449 hypothetical protein AAF813... 177 1e-44
TC84411 similar to GP|4102839|gb|AAD01600.1| LeOPT1 {Lycopersico... 175 4e-44
BF639602 similar to GP|4102839|gb| LeOPT1 {Lycopersicon esculent... 171 5e-43
TC82302 similar to PIR|F84663|F84663 probable nitrate transporte... 171 5e-43
BM780227 similar to PIR|E84798|E84 probable peptide/amino acid t... 171 5e-43
TC86049 weakly similar to GP|20466248|gb|AAM20441.1 putative tra... 167 1e-41
AW980440 similar to GP|17381202|gb putative peptide transporter ... 166 2e-41
BE998753 similar to PIR|T45958|T45 oligopeptide transporter-like... 163 2e-40
BG646222 similar to PIR|F86358|F86 Similar to peptide transporte... 159 3e-39
AW694862 similar to GP|10716600|gb putative peptide transport pr... 157 8e-39
TC80488 similar to GP|4102839|gb|AAD01600.1| LeOPT1 {Lycopersico... 148 6e-36
>TC82936 similar to PIR|G84829|G84829 probable PTR2 family peptide
transporter [imported] - Arabidopsis thaliana, partial
(53%)
Length = 1112
Score = 431 bits (1109), Expect(2) = e-127
Identities = 206/345 (59%), Positives = 268/345 (76%), Gaps = 2/345 (0%)
Frame = +3
Query: 192 GTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHKSRKARSSAKGFIRV 251
G L +TL +VYI+E GW LGY I G +++ + F++GTP+YRHK R ++S AK IRV
Sbjct: 78 GALIATLGLVYIQENLGWGLGYGIPTAGLILSLVIFYIGTPIYRHKVRTSKSPAKDIIRV 257
Query: 252 PVVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTPCFRFLDKAAITESNTDNSNP- 310
+VAF++RKLQ+PSNPSELHEF+MEH V G+R+++HTP RFLDKAAI E T S+
Sbjct: 258 FIVAFKSRKLQLPSNPSELHEFQMEHCVIRGKRQVYHTPTLRFLDKAAIKEDPTTGSSRR 437
Query: 311 -PCTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTMDRSLGPHFHFPAASL 369
P TV QVE AK+++GM IWL+ LIPS WA T+FVKQGTT+DR+LGP F PAASL
Sbjct: 438 VPMTVNQVEGAKLILGMLLIWLVTLIPSTIWAQINTLFVKQGTTLDRNLGPDFKIPAASL 617
Query: 370 WSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQIIGIAVTYAVEIQR 429
SF +M++S+P+YD F+PFMR+ TG+ RG+ LLQR+GIG +IQII IA+ YAVE++R
Sbjct: 618 GSFVTLSMLLSVPMYDRLFVPFMRQKTGHPRGITLLQRLGIGFSIQIIAIAIAYAVEVRR 797
Query: 430 MHVIRKHNIVGPKEVVPMSIFWLLPQNVIFGVANTFLASGLLEFFYDQSPEEMKVLGTTF 489
MHVI++++I GPK++VPMSIFWLLPQ V+ G+A+ F A GLLEFFYDQSPE+M+ LGTTF
Sbjct: 798 MHVIKENHIFGPKDIVPMSIFWLLPQYVLIGIADVFNAIGLLEFFYDQSPEDMQSLGTTF 977
Query: 490 YTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHLDHYY 534
+TS + GN+ NSFLVT+ DK+T + SWI DNLN+SHL +YY
Sbjct: 978 FTSGIGVGNFLNSFLVTMTDKITGRGDRKSWIADNLNDSHLXYYY 1112
Score = 43.9 bits (102), Expect(2) = e-127
Identities = 17/22 (77%), Positives = 18/22 (81%)
Frame = +2
Query: 167 FDDFKPGEKEHKVSFFNWWVFT 188
FDDF P EKE K SFFNWW+FT
Sbjct: 2 FDDFNPHEKELKASFFNWWMFT 67
>TC78362 similar to GP|15391731|emb|CAA93316. nitrite transporter {Cucumis
sativus}, partial (88%)
Length = 1871
Score = 343 bits (880), Expect = 1e-94
Identities = 205/550 (37%), Positives = 315/550 (57%), Gaps = 16/550 (2%)
Frame = +3
Query: 25 GKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHKNVVTSITSVSNWSGLSWITPILG 84
G + FIL +V +RFA G A+++ Y+T L+ +V++ ++SN+SGLS +TP+LG
Sbjct: 87 GGIRTLPFILANEVCDRFASAGFHANMITYLTQQLNMPLVSASNTLSNFSGLSSLTPLLG 266
Query: 85 AYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKG--LGPTCENGV-CKEASNFQLAFF 141
A+IADS+ GRFWTI + LIY +GL + + TS++ P C V C+EA + QL
Sbjct: 267 AFIADSFAGRFWTIVFATLIYELGL-ITITTSAIVPHFRPPPCPTQVNCQEAKSSQLWIL 443
Query: 142 YLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVSFFNWWVFTTACGTLASTLFVV 201
YL+++ ++GSG ++ + F DQFD K G + K + FNW+ F +L++ VV
Sbjct: 444 YLALFLTSLGSGGIRSCVVPFSGDQFDMTKKGVESRKWNLFNWYFFCMGFASLSALTIVV 623
Query: 202 YIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHKSRKARSSAKGFIR---VPVVAFRN 258
YI++ GW G I I +A IAF LG+ +Y + + S F+R V V AFR
Sbjct: 624 YIQDHMGWGWGLGIPTIAMFIAIIAFLLGSRLY----KTLKPSGSPFLRLAQVIVAAFRK 791
Query: 259 RKLQVPSNPSELHE-FEMEHYVSSGRRKLHHTPCFRFLDKAAI-TESNTDNSNP------ 310
R +P++P L++ E++ +S R L HT +++LDKAAI T+ N N
Sbjct: 792 RNDALPNDPKLLYQNLELDSSISLEGR-LSHTDQYKWLDKAAIVTDEEAKNLNKLPNLWN 968
Query: 311 PCTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTMDRSLGPHFHFPAASLW 370
TV +VE+ K ++ M IW ++ + + + + Q TMDR L F AS+
Sbjct: 969 LATVHRVEELKCLVRMLPIWASGILLITASSSQHSFVIVQARTMDRHLSHTFEISPASMA 1148
Query: 371 SFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQIIGIAVTYAVEIQRM 430
F+V TM+ + +Y+ F+PF+RR T N G+ LQR+GIG I II V+ +EI+R
Sbjct: 1149IFSVLTMMTGVILYERLFVPFIRRFTKNPAGITCLQRMGIGFVINIIATIVSALLEIKRK 1328
Query: 431 HVIRKHNIV-GPKEVVPMSIFWLLPQNVIFGVANTFLASGLLEFFYDQSPEEMKVLGTTF 489
V K++++ PK ++P+S+FWL+PQ + GVA F+ G LEF YDQSPE M+ T
Sbjct: 1329KVASKYHLLDSPKAIIPISVFWLVPQYFLHGVAEVFMNVGHLEFLYDQSPESMRSSATAL 1508
Query: 490 YTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGD-NLNESHLDHYYAFLFVIAIFNFGVF 548
Y +A G++ + LVT+V K T K + +W+ D NLN L++YY + I + NF +
Sbjct: 1509YCIAIAIGHFIGTLLVTLVHKYTGK--ERNWLPDRNLNRGRLEYYYFLVCGIQVINFIYY 1682
Query: 549 LWVSNGYIYK 558
+ + Y YK
Sbjct: 1683VICAWIYNYK 1712
>TC79430 similar to PIR|F86358|F86358 Similar to peptide transporter
[imported] - Arabidopsis thaliana, partial (55%)
Length = 1303
Score = 291 bits (744), Expect = 6e-79
Identities = 157/427 (36%), Positives = 234/427 (54%), Gaps = 4/427 (0%)
Frame = +2
Query: 7 TLDGTVDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHKNVVTS 66
T+DG VD GRPV+ S +G W+A FI+ +V ERFAYYGI ++L+NY+T L ++ T+
Sbjct: 80 TVDGVVDFKGRPVLRSSSGGWRAAAFIISVEVAERFAYYGINSNLINYLTGPLGQSTATA 259
Query: 67 ITSVSNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGLGPTCE 126
+V+ WSG + + P+LGA++ADS+LGR+ TI ++ L+Y + L LL L+++L G
Sbjct: 260 AENVNIWSGTASLLPLLGAFLADSFLGRYRTIIIASLVYILALSLLTLSATLPSDG---- 427
Query: 127 NGVCKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVSFFNWWV 186
+ Q F+ S+Y +A G KP + FGADQFD P E+ + SFFNWW
Sbjct: 428 --------DAQAILFFFSLYLVAFAQGGHKPCVQAFGADQFDINHPQERRSRSSFFNWWY 583
Query: 187 FTTACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHK--SRKARSS 244
FT G L + + Y+++ GW LG+ I I ++A F LGT YR + R
Sbjct: 584 FTFTAGLLVTVSILNYVQDNVGWVLGFGIPWIAMIIALSLFSLGTWTYRFSIPGNQQRGP 763
Query: 245 AKGFIRVPVVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTPC--FRFLDKAAITE 302
RV + A RN + P R L H P F FL+KA I
Sbjct: 764 FSRIGRVFITALRNFRTTEEEEP---------------RPSLLHQPSQQFSFLNKALIAS 898
Query: 303 SNTDNSNPPCTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTMDRSLGPHF 362
+ + C+V++VE+AK ++ + IW LI + ++ T F KQG T+DR + P F
Sbjct: 899 DGSKENGKVCSVSEVEEAKAILRLVPIWATSLIFAIVFSQSSTFFTKQGVTLDRKILPGF 1078
Query: 363 HFPAASLWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQIIGIAVT 422
+ P ASL SF ++V+ +P+YD +P R TG G+ +LQR+G GI +I + +
Sbjct: 1079YVPPASLQSFISLSIVLFIPVYDRIIVPIARTFTGKPSGITMLQRIGAGILFSVISMVIA 1258
Query: 423 YAVEIQR 429
VE++R
Sbjct: 1259AFVEMKR 1279
>TC83116 similar to GP|11933407|dbj|BAB19758. putative nitrate transporter
NRT1-3 {Glycine max}, partial (50%)
Length = 1192
Score = 288 bits (738), Expect = 3e-78
Identities = 141/298 (47%), Positives = 193/298 (64%), Gaps = 6/298 (2%)
Frame = +3
Query: 4 QAHTLDGTVDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHKNV 63
+ +T DGTV+L G+PV+ S TG WKAC+F++VY+VFER AYYGI ++L+ T LH+
Sbjct: 72 EEYTEDGTVNLKGKPVLRSKTGGWKACSFVVVYEVFERMAYYGISSNLILLFTKKLHQGT 251
Query: 64 VTSITSVSNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGL-G 122
T+ +V+NW G W+TPILGAY+AD++LGRFWT ++ IY G+ LL L SL L
Sbjct: 252 XTAXNNVTNWVGTIWMTPILGAYVADAHLGRFWTFLIASFIYLSGMSLLTLAVSLPTLKP 431
Query: 123 PTCEN---GVCKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKV 179
P C CK+ S QLA FY ++YT+ IG+G KPN+ST GADQFDDF P EK HK+
Sbjct: 432 PECHELXVTKCKKLSTLQLAVFYGALYTLXIGTGGTKPNISTIGADQFDDFHPKEKSHKL 611
Query: 180 SFFNWWVFTTACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHKSR 239
SFFNWW+F+ GTL + +VYI++ GW LGYA+ +G ++ + F GTP YRHK
Sbjct: 612 SFFNWWMFSIFFGTLFANTVLVYIQDNVGWTLGYALPTLGLAISIMIFLGGTPFYRHK-L 788
Query: 240 KARSSAKGFIRVPVVAFRNRKLQVPSNPSELHEFEMEHYVSSGRR--KLHHTPCFRFL 295
A S+ RV V + R K+ VP + +L+E +ME Y G ++ TP R++
Sbjct: 789 PAGSTFTRMARVIVASLRKWKVPVPDDTKKLYELDMEEYAKKGSNTYRIDSTPTLRYI 962
>TC78128 weakly similar to GP|21928025|gb|AAM78041.1 At1g69870/T17F3_10
{Arabidopsis thaliana}, partial (77%)
Length = 2022
Score = 166 bits (420), Expect(3) = 8e-73
Identities = 98/286 (34%), Positives = 141/286 (49%), Gaps = 9/286 (3%)
Frame = +2
Query: 293 RFLDKAAITESNTDNSNPPCTVT---------QVEKAKVVIGMFHIWLLMLIPSNFWAVE 343
R LDKAA+ + NP T+ QVE+ K + IW ++ A +
Sbjct: 1058 RVLDKAALVMEG--DLNPDGTIVNQWNLVSIQQVEEVKCLARTLPIWAAGILGFTAMAQQ 1231
Query: 344 VTIFVKQGTTMDRSLGPHFHFPAASLWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVK 403
T V Q MDR LGP F PA SL ++ + + +P YD +P +R+ T N G+
Sbjct: 1232 GTFIVSQAMKMDRHLGPKFQIPAGSLGVISLIVIGLWVPFYDRICVPSLRKITKNEGGIT 1411
Query: 404 LLQRVGIGIAIQIIGIAVTYAVEIQRMHVIRKHNIVGPKEVVPMSIFWLLPQNVIFGVAN 463
LLQR+GIG+ II + V VE R V + P+ + PMS+ WL PQ V+ G
Sbjct: 1412 LLQRIGIGMVFSIISMIVAGLVEKVRRDVANSNPT--PQGIAPMSVMWLFPQLVLMGFCE 1585
Query: 464 TFLASGLLEFFYDQSPEEMKVLGTTFYTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGD 523
F GL+EFF Q P+ M+ + ++ + A NY +S LV V VT+ W+ +
Sbjct: 1586 AFNIIGLIEFFNRQFPDHMRSIANALFSCSFALANYVSSILVITVHSVTKTHNHPDWLTN 1765
Query: 524 NLNESHLDHYYAFLFVIAIFNFGVFLWVSNGYIYKKECTSTTEPND 569
N+NE LD++Y L + + N FL+VS Y YK +P D
Sbjct: 1766 NINEGRLDYFYYLLAGVGVLNLVYFLYVSQRYHYKGSVDIQEKPMD 1903
Score = 87.8 bits (216), Expect(3) = 8e-73
Identities = 49/145 (33%), Positives = 79/145 (53%), Gaps = 6/145 (4%)
Frame = +2
Query: 25 GKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHKNVVTSITSVSNWSGLSWITPILG 84
G WKA FIL + FER A +G+ A+ + Y+T H V + ++ W G+S P+LG
Sbjct: 233 GGWKAMPFILGNESFERLAAFGLLANFMVYLTREFHLEQVHASNILNIWGGISNFAPLLG 412
Query: 85 AYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGL-GPTCEN-----GVCKEASNFQL 138
A+I+D+Y GRF TI + +G+ + LT+ L L P+C C A++ Q+
Sbjct: 413 AFISDTYTGRFKTIAFASFFSLLGMTAVTLTAWLPKLQPPSCTPQQQALNQCVTANSSQV 592
Query: 139 AFFYLSIYTIAIGSGAVKPNMSTFG 163
F ++ + ++IGS ++P FG
Sbjct: 593 GFLFMGLIFLSIGSSGIRPCSIPFG 667
Score = 60.1 bits (144), Expect(3) = 8e-73
Identities = 33/98 (33%), Positives = 49/98 (49%)
Frame = +3
Query: 163 GADQFDDFKPGEKEHKVSFFNWWVFTTACGTLASTLFVVYIEEKFGWALGYAISAIGFLV 222
G DQFD K+ SFFNW+ + L + +VYI++ W G+AI + L
Sbjct: 666 GVDQFDPTTEKGKKGINSFFNWYYTSFTVVLLFTQTVIVYIQDSISWKFGFAIPTLCMLX 845
Query: 223 AFIAFFLGTPMYRHKSRKARSSAKGFIRVPVVAFRNRK 260
+ I FF+GT +Y H + S +V V +F+ RK
Sbjct: 846 SIILFFIGTKIYVHVKPEG-SIFSSIAQVLVASFKKRK 956
>TC93408 similar to GP|11933407|dbj|BAB19758. putative nitrate transporter
NRT1-3 {Glycine max}, partial (44%)
Length = 909
Score = 220 bits (560), Expect = 1e-57
Identities = 120/293 (40%), Positives = 176/293 (59%), Gaps = 4/293 (1%)
Frame = +1
Query: 200 VVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHKSRKARSSAKGFIRVPVVAFRNR 259
+VYI++ GW LGYA+ +G ++ + F GTP YRHK A S+ RV V + R
Sbjct: 13 LVYIQDNVGWTLGYALPTLGLAISIMIFLGGTPFYRHKL-PAGSTFTRMARVIVASLRKW 189
Query: 260 KLQVPSNPSELHEFEMEHYVSSGRR--KLHHTPCFRFLDKAAITESNTDNSNP--PCTVT 315
K+ VP + +L+E +ME Y G ++ TP RFLDKA++ T +++P CTVT
Sbjct: 190 KVPVPDDTKKLYELDMEEYAKKGSNTYRIDSTPTLRFLDKASV---KTGSTSPWMLCTVT 360
Query: 316 QVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTMDRSLGPHFHFPAASLWSFAVF 375
VE+ K ++ M I + +PS A T+FVKQGTT+DR +G F P ASL +F
Sbjct: 361 HVEETKQMLRMIPILVATFVPSTMMAQVNTLFVKQGTTLDRHIGS-FKIPPASLAAFVTL 537
Query: 376 TMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQIIGIAVTYAVEIQRMHVIRK 435
++++ + +YD FF+ M+R T N RG+ LLQR+GIG+ + I + V E R+ V ++
Sbjct: 538 SLLVCVVLYDRFFVRIMQRLTKNPRGITLLQRMGIGLVLHTIIMVVASVTENYRLRVAKE 717
Query: 436 HNIVGPKEVVPMSIFWLLPQNVIFGVANTFLASGLLEFFYDQSPEEMKVLGTT 488
H +V VP+SIF LLPQ ++ G A+ FL +EFFYDQ+P MK +GT+
Sbjct: 718 HGLVESGGQVPLSIFILLPQFILMGTADAFLEVAKIEFFYDQAPXSMKSIGTS 876
>TC79810 similar to GP|11994403|dbj|BAB02362. nitrate transporter
{Arabidopsis thaliana}, partial (60%)
Length = 1236
Score = 204 bits (519), Expect = 7e-53
Identities = 135/387 (34%), Positives = 212/387 (53%), Gaps = 16/387 (4%)
Frame = +3
Query: 9 DGTVDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHKNVVTSIT 68
+ VD G P + +G W A IL ++ ER GI +LV Y+ LH + +S T
Sbjct: 96 NAAVDFRGHPADKTKSGGWLAAGLILGTELAERICVMGISMNLVTYLVGDLHLHSASSAT 275
Query: 69 SVSNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGLGPT-C-- 125
V+N+ G + +LG ++AD+ LGR+ T+ +S I +G+ +L L +++ + P C
Sbjct: 276 IVTNFMGTLNLLGLLGGFLADAKLGRYLTVAISATIATVGVCMLTLATTIPSMTPPPCSE 455
Query: 126 ---ENGVCKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVSFF 182
++ C +AS QL+ + ++YTIA+G G +K N+S FG+DQFD P E+++ + FF
Sbjct: 456 VRRQHHQCIQASGKQLSLLFAALYTIALGGGGIKSNVSGFGSDQFDTNDPKEEKNMIFFF 635
Query: 183 NWWVFTTACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHKSRKAR 242
N + F + G+L S + +VY+++ G GY ISA LVA GTP+YR K K +
Sbjct: 636 NRFYFFISIGSLFSVVVLVYVQDNIGRGWGYGISAGTMLVAVGILLCGTPLYRFK--KPQ 809
Query: 243 SSAKGFI-RVPVVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTPCFRFLDKAAI- 300
S I RV +A++ R L +PS+P+ L+ + K+ +T FR LDKAAI
Sbjct: 810 GSPLTIIWRVLFLAWKKRTLPIPSDPTLLNGYL--------EAKVTYTDKFRSLDKAAIL 965
Query: 301 --TESNTDNSNPP---CTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEV---TIFVKQGT 352
T+S N+ P T+TQVE+ K+VI + IW ++ FW V T ++Q T
Sbjct: 966 DETKSKDGNNENPWLVSTMTQVEEVKMVIKLLPIWSTCIL---FWTVYSQMNTFTIEQAT 1136
Query: 353 TMDRSLGPHFHFPAASLWSFAVFTMVI 379
M+R +G PA SL +F T+++
Sbjct: 1137FMNRKVG-SLEIPAGSLSAFLFITILL 1214
>TC88300 similar to GP|9581817|emb|CAC00544.1 putative low-affinity nitrate
transporter {Nicotiana plumbaginifolia}, partial (56%)
Length = 1169
Score = 200 bits (508), Expect = 1e-51
Identities = 109/318 (34%), Positives = 178/318 (55%), Gaps = 10/318 (3%)
Frame = +1
Query: 251 VPVVAFRNRKLQVPSNPSELHEFE--MEHYVSSGRRKLHHTPCFRFLDKAAITESNTDNS 308
V V A+R R +++P + S L + + + ++ L H+ FRFLDKAAI + TD +
Sbjct: 85 VYVAAWRKRNMELPYDSSLLFNVDDIEDEMLRKKKQVLPHSKQFRFLDKAAIKDPKTDGN 264
Query: 309 NPPC-------TVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTMDRSLGPH 361
T+T VE+ K+V+ M IW ++ +A T V Q TT++R +G
Sbjct: 265 EINVVRKWYLSTLTDVEEVKLVLRMLPIWATTIMFWTVYAQMTTFSVSQATTLNRHIGKS 444
Query: 362 FHFPAASLWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQIIGIAV 421
F P ASL +F + ++++++PIYD +P R+ N +G+ LQR+G+G+ I +
Sbjct: 445 FQIPPASLTAFFIGSILLTIPIYDRVIVPITRKIFKNPQGLTPLQRIGVGLVFSIFAMVA 624
Query: 422 TYAVEIQRMHVIRKHNIV-GPKEVVPMSIFWLLPQNVIFGVANTFLASGLLEFFYDQSPE 480
E++RM + HN+ P +PMS+FWL+PQ G F G L+FF + P+
Sbjct: 625 AALTELKRMRMAHLHNLTHNPNSEIPMSVFWLIPQFFFVGSGEAFTYIGQLDFFLRECPK 804
Query: 481 EMKVLGTTFYTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHLDHYYAFLFVI 540
MK + T + ST++ G +F+S LVT+V KVT + W+ DNLNE+ L ++Y L ++
Sbjct: 805 GMKTMSTGLFLSTLSLGFFFSSLLVTLVHKVTSQ--HKPWLADNLNEAKLYNFYWLLALL 978
Query: 541 AIFNFGVFLWVSNGYIYK 558
++ N ++L +N Y+YK
Sbjct: 979 SVLNLVIYLLCANWYVYK 1032
>AW694006 similar to PIR|F86358|F86 Similar to peptide transporter [imported]
- Arabidopsis thaliana, partial (33%)
Length = 641
Score = 178 bits (452), Expect = 4e-45
Identities = 86/216 (39%), Positives = 135/216 (61%)
Frame = +1
Query: 12 VDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHKNVVTSITSVS 71
VD G+P + S +G W++ FI+ +V ER +Y GI +L++Y+T L ++ T+ +V+
Sbjct: 1 VDYRGQPAVRSKSGYWRSAWFIIGVEVAERISYIGIKGNLISYLTGPLKQSTATAAKNVN 180
Query: 72 NWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGLGPTCENGVCK 131
W+G + + P+LGA++ADS+LGR+ TI L+ LIY +GLGLL L++ L L +C
Sbjct: 181 IWAGTASLLPLLGAFVADSFLGRYHTIILASLIYILGLGLLTLSTILPSLA-SC------ 339
Query: 132 EASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVSFFNWWVFTTAC 191
++ Q+ F++S+Y +AIG G KP + FGADQFD+ P E + SFFNWW FT
Sbjct: 340 -STQSQVILFFISLYLVAIGQGGHKPCVQAFGADQFDEKYPEEHRARSSFFNWWYFTMVA 516
Query: 192 GTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAF 227
G A+ + YI++ + W LG+ I + ++A I F
Sbjct: 517 GATATLPILTYIQDNYSWVLGFGIPCVIMIIALIIF 624
>TC87099 similar to PIR|G86449|G86449 hypothetical protein AAF81343.1
[imported] - Arabidopsis thaliana, partial (86%)
Length = 2205
Score = 177 bits (449), Expect = 1e-44
Identities = 96/238 (40%), Positives = 145/238 (60%), Gaps = 3/238 (1%)
Frame = +2
Query: 2 EAQAHTLDGTVDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHK 61
E + TLDG+VD GRP I + +G+W A T IL+ Q A++G+G +LV ++T L +
Sbjct: 128 ETEELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVLGQ 307
Query: 62 NVVTSITSVSNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGL 121
+ + +VS W+G +I ++GA+++DSY GR+ T + I+ GL L +T+ L L
Sbjct: 308 DNAAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLALL 487
Query: 122 GPT-CENGV--CKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHK 178
P C NG C E S+ ++ FYLSIY IA+G+G +PN++TFGADQFD+ E K
Sbjct: 488 RPKGCGNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSKESYSK 667
Query: 179 VSFFNWWVFTTACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRH 236
V+FF+++ G+L S + Y E++ WALG+ SA +A + F +GTP YRH
Sbjct: 668 VAFFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPKYRH 841
Score = 90.9 bits (224), Expect = 1e-18
Identities = 49/158 (31%), Positives = 84/158 (53%), Gaps = 4/158 (2%)
Frame = +2
Query: 400 RGVKLLQRVGIGIAIQIIGIAVTYAVEIQRMHVIRKHNIVGPKEVVPMSIFWLLPQNVIF 459
+G+ LQR+GIG+ I II + VE R+ ++ + +SIFW +PQ +
Sbjct: 1355 KGLTELQRMGIGLVIAIIAMVTAGIVECYRLKYAKQG------DTSSLSIFWQIPQYALI 1516
Query: 460 GVANTFLASGLLEFFYDQSPEEMKVLGTTFYTSTMAGGNYFNSFLVTVVDKVTRKMCDHS 519
G + F+ G LEFF Q+P+ +K G+ ++++ GNY +S +V++V K++ +
Sbjct: 1517 GASEVFMYVGQLEFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPG 1696
Query: 520 WIGDNLNESHLDHYYAFLFVIAIFNFGVFL----WVSN 553
WI NLN HLD ++ L V+ + ++ W N
Sbjct: 1697 WIPGNLNRGHLDRFFFLLAVLTSLDLIAYIACAKWFQN 1810
Score = 64.3 bits (155), Expect = 1e-10
Identities = 51/186 (27%), Positives = 85/186 (45%), Gaps = 8/186 (4%)
Frame = +3
Query: 174 EKEHKVSFFNWWVFTTACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPM 233
+K H ++ F W + TLF ++ K LG+ + +G + FFL P
Sbjct: 663 QKWHFLATFTWH*TLVH---FSQTLFWAILKMKDYGLLGFGL-LLGLPFWLLYFFLLAPQ 830
Query: 234 YRHKSRKARSSAKGFIRVPVVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTPCFR 293
+ + F +V A R +Q+ SN +L+ + + ++ RK+ HT F+
Sbjct: 831 NIDTFKPCGNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILHTHGFK 1010
Query: 294 FLDKAAITESNT------DNSNP--PCTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVT 345
FLD+AA S NP C +TQVE+ K ++ + IWL +I S + +
Sbjct: 1011FLDRAAYITSRDLEVQKGGQHNPWXLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMAS 1190
Query: 346 IFVKQG 351
+FV+QG
Sbjct: 1191LFVEQG 1208
>TC84411 similar to GP|4102839|gb|AAD01600.1| LeOPT1 {Lycopersicon
esculentum}, partial (38%)
Length = 804
Score = 175 bits (444), Expect = 4e-44
Identities = 92/240 (38%), Positives = 145/240 (60%)
Frame = +1
Query: 316 QVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTMDRSLGPHFHFPAASLWSFAVF 375
QVE+ K++I MF IW +I S+ +A T+FV+QGT M+ S+G F ASL +F V
Sbjct: 1 QVEELKILIRMFPIWATGIIFSSVYAQMSTLFVEQGTMMNTSIGS-FKLSPASLSTFEVA 177
Query: 376 TMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQIIGIAVTYAVEIQRMHVIRK 435
++V+ +P+YD +P +++ TG RG+ + QR+GIG I + + AVEI+R+ + R+
Sbjct: 178 SVVMWVPVYDKILVPIVKKFTGKKRGISVFQRIGIGPFISGLCMLAAAAVEIKRLQLARE 357
Query: 436 HNIVGPKEVVPMSIFWLLPQNVIFGVANTFLASGLLEFFYDQSPEEMKVLGTTFYTSTMA 495
++V VP+S+ W +PQ +I G A F G LEFFY++SP+ M+ + +
Sbjct: 358 LDLVDKPVGVPLSVLWQIPQYLILGAAEIFTFVGQLEFFYEESPDAMRTICGALPLLNFS 537
Query: 496 GGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHLDHYYAFLFVIAIFNFGVFLWVSNGY 555
GNY +SF++T+V T K WI DNLN+ HLD+Y+ L +++ N VF+ + Y
Sbjct: 538 LGNYLSSFILTIVTYFTTKGGRLGWIPDNLNKGHLDYYFLLLSGLSLLNMLVFIVAAKIY 717
>BF639602 similar to GP|4102839|gb| LeOPT1 {Lycopersicon esculentum}, partial
(32%)
Length = 684
Score = 171 bits (434), Expect = 5e-43
Identities = 83/192 (43%), Positives = 124/192 (64%), Gaps = 1/192 (0%)
Frame = +2
Query: 2 EAQAHTLDGTVDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHK 61
E HT DG+V++ G + TG WK+C FIL ER AYYGI A+LV Y+T LH+
Sbjct: 98 EDSVHTGDGSVNVKGELSVKRETGTWKSCPFILGTFFCERLAYYGIAANLVTYLTTRLHE 277
Query: 62 NVVTSITSVSNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGL 121
V++ +V+ + G ++TP++G++ AD+YLGR+WTI + IY +G+ +L+L++++ L
Sbjct: 278 GTVSAAKNVTTFQGTCYLTPLIGSFFADTYLGRYWTIAVFYGIYLIGICILILSATIAAL 457
Query: 122 GP-TCENGVCKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVS 180
P C + VC A+ Q A F+L ++ IA+G+G +KP + FGADQFDD E+ K S
Sbjct: 458 KPMECVSSVCPSATLAQRAVFFLGLFLIAVGTGGIKPCIWPFGADQFDDTDHKERASKGS 637
Query: 181 FFNWWVFTTACG 192
FFNW FT+ G
Sbjct: 638 FFNWNYFTSNIG 673
>TC82302 similar to PIR|F84663|F84663 probable nitrate transporter
[imported] - Arabidopsis thaliana, partial (51%)
Length = 1133
Score = 171 bits (434), Expect = 5e-43
Identities = 105/319 (32%), Positives = 177/319 (54%), Gaps = 9/319 (2%)
Frame = +3
Query: 250 RVPVVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTPCFRFLDKAAIT-ESNTDNS 308
+V V + + RK+++P N L+E E ++ + FRFL+KAAI E + D
Sbjct: 9 QVIVASIKKRKMELPYNVGSLYEDTPED------SRIEQSEQFRFLEKAAIVVEGDFDKD 170
Query: 309 ------NP--PCTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTMDRSLGP 360
NP C++T+VE+ K+++ + IW +I +A +T V+Q TM+R++G
Sbjct: 171 LYGSGPNPWKLCSLTRVEEVKMMVRLLPIWATTIIFWTTYAQMITFSVEQAATMERNVG- 347
Query: 361 HFHFPAASLWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQIIGIA 420
+F PA SL F V ++++L + D +P ++ G G LQR IG+ + IG+A
Sbjct: 348 NFQIPAGSLTVFFVVAILLTLAVNDRIIMPLWKKLNGKP-GFSNLQRNAIGLLLSTIGMA 524
Query: 421 VTYAVEIQRMHVIRKHNIVGPKEVVPMSIFWLLPQNVIFGVANTFLASGLLEFFYDQSPE 480
+E++R+ V + + G + +P+S+F L+PQ + G F+ +G L+FF QSP+
Sbjct: 525 AASLIEVKRLSVAK--GVKGNQTTLPISVFLLVPQFFLVGSGEAFIYTGQLDFFITQSPK 698
Query: 481 EMKVLGTTFYTSTMAGGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHLDHYYAFLFVI 540
MK + T + +T++ G + +SFLV+VV KVT W+ D++N+ LD +YA L ++
Sbjct: 699 GMKTMSTGLFLTTLSLGFFVSSFLVSVVKKVTGTRDGQGWLADHINKGRLDLFYALLTIL 878
Query: 541 AIFNFGVFLWVSNGYIYKK 559
+ NF FL + Y KK
Sbjct: 879 SFINFVAFLVCAFWYKPKK 935
>BM780227 similar to PIR|E84798|E84 probable peptide/amino acid transporter
[imported] - Arabidopsis thaliana, partial (36%)
Length = 750
Score = 171 bits (434), Expect = 5e-43
Identities = 90/250 (36%), Positives = 140/250 (56%)
Frame = +3
Query: 51 LVNYMTVHLHKNVVTSITSVSNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLG 110
LV Y+ + +H+++ T+ +V+ W+G++ + P+ G +IAD+YLGR+ + S ++Y MGL
Sbjct: 6 LVLYLILVIHQDLKTAARNVNYWAGVTTLMPLFGGFIADAYLGRYSAVVASSIVYLMGLI 185
Query: 111 LLVLTSSLKGLGPTCENGVCKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDF 170
LL L+ L L P C E F+L+IY I+I +G KP++ +FGADQFD+
Sbjct: 186 LLTLSWFLPSLKPCDHTTTCNEPRKIHEVVFFLAIYLISIATGGHKPSLESFGADQFDED 365
Query: 171 KPGEKEHKVSFFNWWVFTTACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLG 230
E++ K+SFFNWW G + +VYI++ W I ++ + F +G
Sbjct: 366 HVEERKQKMSFFNWWNCALCSGLILGVTLIVYIQDNINWGAADIIFTGVMALSLLIFIIG 545
Query: 231 TPMYRHKSRKARSSAKGFIRVPVVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTP 290
P YR++ + S ++V V AF RKL PSNP +L+E H + R+ L HT
Sbjct: 546 RPFYRYRV-PSGSPLTPMLQVLVAAFSKRKLPYPSNPDQLYEVSKSH--GNKRKFLCHTK 716
Query: 291 CFRFLDKAAI 300
RFL +AAI
Sbjct: 717 KLRFL*QAAI 746
>TC86049 weakly similar to GP|20466248|gb|AAM20441.1 putative transport
protein {Arabidopsis thaliana}, partial (43%)
Length = 1836
Score = 167 bits (422), Expect = 1e-41
Identities = 117/422 (27%), Positives = 203/422 (47%), Gaps = 14/422 (3%)
Frame = +2
Query: 148 IAIGSGAVKPNMSTFGADQFD--DFKPGEKEHKVSFFNWWVFTTACGTLASTLFVVYIEE 205
IAIG+G + ++S FGA Q + D G + ++ FF+W+ T + + +VYI++
Sbjct: 11 IAIGNGGITCSLS-FGAYQVNRKDNPDGYRVLEI-FFSWYYAFTTIAVIIALTGIVYIQD 184
Query: 206 KFGWALGYAISAIGFLVAFIAFFLGTPMYRHKSRKARSSAKGFIRVPVVAFRNRKLQVPS 265
GW +G+ + AI L++ + FFL +P+Y K ++ S GF +V V A++NRKL +P
Sbjct: 185 HLGWKVGFGVPAILMLISTVLFFLASPLYV-KIKQKTSLFTGFAQVSVAAYKNRKLPLP- 358
Query: 266 NPSELHEFEMEHYVSSGRRKLHHTPCFRFLDKAAI---------TESNTDNSNPPCTVTQ 316
P EF ++ + T RFL+KA + ++ + N CTV Q
Sbjct: 359 -PKTSPEF---YHQQKDSELVVPTDKLRFLNKACVIKDHEQDIASDGSAINRWSLCTVDQ 526
Query: 317 VEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTMDRSL--GPHFHFPAASLWSFAV 374
VE+ K +I + +W + S + + + Q ++DR + +F PA S +
Sbjct: 527 VEELKAIIKVIPLWSTAITMSI--NIGGSFGLLQAKSLDRHIISSSNFEVPAGSFSVILI 700
Query: 375 FTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQIIGIAVTYAVE-IQRMHVI 433
++I + IYD IP + G + +R+GIG+ + + E ++R I
Sbjct: 701 VAILIWIIIYDRVLIPLASKIRGKPVIISPKKRMGIGLFFNFLHLITAAIFETVRRKEAI 880
Query: 434 RKHNIVGPKEVVPMSIFWLLPQNVIFGVANTFLASGLLEFFYDQSPEEMKVLGTTFYTST 493
++ + V+ MS WL PQ + G+A F G EF+Y + P+ M + +
Sbjct: 881 KEGYLNDTHGVLKMSAMWLAPQLCLAGIAEMFNVIGQNEFYYKEFPKSMSSVAASLSGLA 1060
Query: 494 MAGGNYFNSFLVTVVDKVTRKMCDHSWIGDNLNESHLDHYYAFLFVIAIFNFGVFLWVSN 553
M GN +S ++++++ T + W+ DN+N+ H D YY + I N +L S
Sbjct: 1061MGVGNLVSSLVLSIIESTTPSGGNEGWVSDNINKGHFDKYYWVIVGINALNLLYYLVCSW 1240
Query: 554 GY 555
Y
Sbjct: 1241AY 1246
>AW980440 similar to GP|17381202|gb putative peptide transporter protein
{Arabidopsis thaliana}, partial (23%)
Length = 688
Score = 166 bits (421), Expect = 2e-41
Identities = 82/176 (46%), Positives = 105/176 (59%), Gaps = 5/176 (2%)
Frame = +1
Query: 123 PTCENGV-----CKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEH 177
P C+ GV C +AS Q F L++Y I +G+G KPN+ST GADQFDDF P EK
Sbjct: 22 PECDIGVVAFENCPKASPLQKGIFCLALYIIVLGTGGTKPNISTMGADQFDDFDPKEKSD 201
Query: 178 KVSFFNWWVFTTACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIAFFLGTPMYRHK 237
K+SFFNWW F+ G L +T F+VYI++ GW LGY + IG + + F LGTP YRHK
Sbjct: 202 KLSFFNWWFFSILIGVLFATTFLVYIQDNIGWELGYGLPTIGLAFSILVFLLGTPYYRHK 381
Query: 238 SRKARSSAKGFIRVPVVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTPCFR 293
S ++V V A R K +VP + ELHE ME Y +GR ++ HT FR
Sbjct: 382 LPPG-SPITRMLQVFVAAIRKWKARVPEDKKELHELSMEEYTCNGRTRIDHTSFFR 546
>BE998753 similar to PIR|T45958|T45 oligopeptide transporter-like protein -
Arabidopsis thaliana, partial (44%)
Length = 809
Score = 163 bits (412), Expect = 2e-40
Identities = 92/261 (35%), Positives = 146/261 (55%), Gaps = 6/261 (2%)
Frame = +3
Query: 251 VPVVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKLHHTPCFRFLDKAAI---TESNTDN 307
V V + R ++ P++ S L+E G RKL HT RF DKAA+ +++ ++
Sbjct: 30 VIVASIRKYRVDAPTDKSLLYEIADTESAIKGSRKLDHTNELRFFDKAAVQGESDNLKES 209
Query: 308 SNP--PCTVTQVEKAKVVIGMFHIWLLMLIPSNFWAVEVTIFVKQGTTMDRSLG-PHFHF 364
NP CTVTQVE+ K ++ + +W +I + + T+FV QG TM+ +G F
Sbjct: 210 INPWRLCTVTQVEELKSILRLLPVWATGIIFATVYGQMSTLFVLQGQTMNTHVGNSSFKI 389
Query: 365 PAASLWSFAVFTMVISLPIYDNFFIPFMRRHTGNHRGVKLLQRVGIGIAIQIIGIAVTYA 424
P ASL F +++ +P+YD +P R+ TG+ G+ LQR+G+G+ I I +
Sbjct: 390 PPASLSIFDTISVIFWVPVYDRIIVPIARKFTGHKNGLTQLQRMGVGLFISIFSMVAAAF 569
Query: 425 VEIQRMHVIRKHNIVGPKEVVPMSIFWLLPQNVIFGVANTFLASGLLEFFYDQSPEEMKV 484
+E+ R+ +R++N +E +PM+IFW +PQ + G A F G LEFFY+Q+P+ M+
Sbjct: 570 LELVRLRTVRRNNYYELEE-IPMTIFWQVPQYFLIGCAEVFTFIGQLEFFYEQAPDAMRS 746
Query: 485 LGTTFYTSTMAGGNYFNSFLV 505
L + T+A G Y +S LV
Sbjct: 747 LCSALSLLTVAFGQYLSSLLV 809
>BG646222 similar to PIR|F86358|F86 Similar to peptide transporter [imported]
- Arabidopsis thaliana, partial (28%)
Length = 749
Score = 159 bits (402), Expect = 3e-39
Identities = 88/226 (38%), Positives = 134/226 (58%), Gaps = 1/226 (0%)
Frame = +2
Query: 1 MEAQAHTLDGTVDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLH 60
+EAQ +DG VD G+P + +G W++ FI+ +V ER +YYGI +L++Y+T L
Sbjct: 95 LEAQLSLVDGAVDYNGQPAVRFKSGYWRSAWFIIGVEVAERVSYYGIQGNLISYLTGPLK 274
Query: 61 KNVVTSITSVSNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKG 120
++ T+ +V+ W+G + + P+LGA++ADS+LGR+ TI L+ LIY +GLGLL L++ L
Sbjct: 275 QSTATAAKNVNVWAGTASLLPLLGAFVADSFLGRYRTIILASLIYILGLGLLTLSALLPS 454
Query: 121 LGPTCENGVCKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQFDDFKPGEKEHKVS 180
L C S Q+ F++S+Y +AIG G KP + FGADQFD+ P E E + S
Sbjct: 455 L------TFCSPQS--QVILFFISLYLVAIGQGGHKPCVQAFGADQFDEKHPKEHEARSS 610
Query: 181 FFNWWVFTTACGTLASTL-FVVYIEEKFGWALGYAISAIGFLVAFI 225
FFN VF C ++T +Y LG+ I + ++A I
Sbjct: 611 FFNLVVFYYDCRLHSNTYDLELYTR*TLVGFLGFGIPCVVMIIALI 748
>AW694862 similar to GP|10716600|gb putative peptide transport protein {Oryza
sativa}, partial (26%)
Length = 645
Score = 157 bits (398), Expect = 8e-39
Identities = 81/188 (43%), Positives = 109/188 (57%), Gaps = 4/188 (2%)
Frame = +1
Query: 111 LLVLTSSLKGLGPT----CENGVCKEASNFQLAFFYLSIYTIAIGSGAVKPNMSTFGADQ 166
LL L+ SL L P + CKEAS L FY+++Y IA+G+G KPN+ST GADQ
Sbjct: 4 LLTLSVSLPSLKPPKCHEMDVTKCKEASTQTLVVFYVALYIIAVGTGGTKPNISTIGADQ 183
Query: 167 FDDFKPGEKEHKVSFFNWWVFTTACGTLASTLFVVYIEEKFGWALGYAISAIGFLVAFIA 226
FDDF P EK K+SFFNWW+ G+L + +VYI++ GW LGYA+ +G ++ I
Sbjct: 184 FDDFDPKEKSLKLSFFNWWMSCIVFGSLFAFTVIVYIQDNVGWTLGYALPTLGLAISIIT 363
Query: 227 FFLGTPMYRHKSRKARSSAKGFIRVPVVAFRNRKLQVPSNPSELHEFEMEHYVSSGRRKL 286
F GTP YRHK K S +V V A R VP +P EL+E +E Y G+ ++
Sbjct: 364 FLAGTPFYRHKLIKG-SPFISMAKVIVAAIRKFDAVVPDDPKELYELSLEEYTKKGKFRI 540
Query: 287 HHTPCFRF 294
T F++
Sbjct: 541 DLTQTFKY 564
>TC80488 similar to GP|4102839|gb|AAD01600.1| LeOPT1 {Lycopersicon
esculentum}, partial (28%)
Length = 664
Score = 148 bits (373), Expect = 6e-36
Identities = 70/157 (44%), Positives = 103/157 (65%), Gaps = 1/157 (0%)
Frame = +3
Query: 2 EAQAHTLDGTVDLGGRPVISSVTGKWKACTFILVYQVFERFAYYGIGADLVNYMTVHLHK 61
E++ +T DG+VD GRPV+ TG WKAC FIL + ER AYYGI +LV Y+T LH+
Sbjct: 180 ESRRYTGDGSVDFKGRPVLKQNTGNWKACPFILGNECCERLAYYGIAKNLVTYLTRKLHQ 359
Query: 62 NVVTSITSVSNWSGLSWITPILGAYIADSYLGRFWTITLSLLIYAMGLGLLVLTSSLKGL 121
++ +V+ W G ++TP++GA +ADSYLGR+WTI + IY +G+ L L++S+ L
Sbjct: 360 GNASAARNVTTWQGTCYLTPLIGAILADSYLGRYWTIAVFSTIYFIGMCTLTLSASVPAL 539
Query: 122 GPT-CENGVCKEASNFQLAFFYLSIYTIAIGSGAVKP 157
P C + C A+ Q A F++ +Y IA+G+G +KP
Sbjct: 540 KPADCFSSACPPATPSQYAAFFVGLYLIALGTGGIKP 650
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.326 0.140 0.441
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,223,428
Number of Sequences: 36976
Number of extensions: 339761
Number of successful extensions: 2526
Number of sequences better than 10.0: 116
Number of HSP's better than 10.0 without gapping: 2418
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2452
length of query: 573
length of database: 9,014,727
effective HSP length: 101
effective length of query: 472
effective length of database: 5,280,151
effective search space: 2492231272
effective search space used: 2492231272
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0248.6


