

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
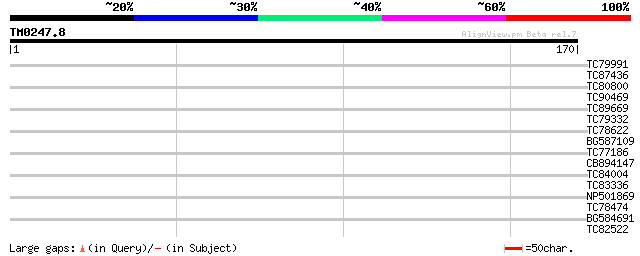
Query= TM0247.8
(170 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC79991 similar to GP|10177307|dbj|BAB10568. contains similarity... 29 0.80
TC87436 similar to PIR|T51575|T51575 2-hydroxyphytanoyl-CoA lyas... 29 1.0
TC80800 similar to GP|6606532|gb|AAF19195.1| pectate lyase 1 {Mu... 28 2.3
TC90469 similar to GP|11066213|gb|AAG28503.1 hexokinase {Citrus ... 28 2.3
TC89669 homologue to GP|1237082|emb|CAA65540.1 ADP-glucose pyrop... 27 3.0
TC79332 similar to PIR|JC6203|JC6203 SP8 binding protein homolog... 27 3.0
TC78622 weakly similar to GP|14587220|dbj|BAB61154. putative GTP... 27 4.0
BG587109 homologue to GP|22091671|em photosystem1 subunit A {Pin... 27 5.2
TC77186 phosphate transporter 27 5.2
CB894147 similar to GP|15375301|gb| putative c-myb-like transcri... 26 6.8
TC84004 type IIB calcium ATPase [Medicago truncatula] 26 6.8
TC83336 similar to GP|13938440|gb|AAH07361.1 Similar to protein ... 26 6.8
NP501869 NP501869|AY115844.1|AAM55305.1 auxin influx carrier pro... 26 8.9
TC78474 homologue to PIR|T09550|T09550 phosphoprotein phosphatas... 26 8.9
BG584691 similar to PIR|E87601|E876 OmpA family protein [importe... 26 8.9
TC82522 similar to PIR|A96708|A96708 hypothetical protein T2E12.... 26 8.9
>TC79991 similar to GP|10177307|dbj|BAB10568. contains similarity to zinc
finger protein~gene_id:MDC12.23 {Arabidopsis thaliana},
partial (8%)
Length = 1023
Score = 29.3 bits (64), Expect = 0.80
Identities = 21/78 (26%), Positives = 34/78 (42%), Gaps = 9/78 (11%)
Frame = -3
Query: 73 HALTRSHASVEF-----HAFHKVSRLSGVSSIHKVSRLSEASRLIPWMAN----KLLNER 123
H L H S++ H FH H++ ++S+L P + KLL+ R
Sbjct: 547 HQLVHHHQSLKLYPQFQHPFH-----------HQLVHHHQSSKLYPQFQSPFHQKLLHHR 401
Query: 124 RSTNILRTYQTPQHLDPT 141
S+ + +Q P HL P+
Sbjct: 400 *SSKLYHQFQQPLHLQPS 347
>TC87436 similar to PIR|T51575|T51575 2-hydroxyphytanoyl-CoA lyase-like
protein - Arabidopsis thaliana, partial (75%)
Length = 1592
Score = 28.9 bits (63), Expect = 1.0
Identities = 22/61 (36%), Positives = 33/61 (54%), Gaps = 2/61 (3%)
Frame = -3
Query: 70 VKFHALTRSHASVEFHAFHKVSRLSGVS--SIHKVSRLSEASRLIPWMANKLLNERRSTN 127
+ FH+ T+S + + F++ K R+S S +I K + S S I LLN+ RSTN
Sbjct: 1569 LNFHS-TKSISGIPFNSNDKTIRISDSSRNTITKTNPHSPPSTSIQPSPRLLLNQHRSTN 1393
Query: 128 I 128
I
Sbjct: 1392 I 1390
>TC80800 similar to GP|6606532|gb|AAF19195.1| pectate lyase 1 {Musa
acuminata}, partial (86%)
Length = 1216
Score = 27.7 bits (60), Expect = 2.3
Identities = 12/41 (29%), Positives = 21/41 (50%)
Frame = -3
Query: 75 LTRSHASVEFHAFHKVSRLSGVSSIHKVSRLSEASRLIPWM 115
+ +S + ++FH FH S S ++ S A+ L PW+
Sbjct: 1094 INKSPSDLQFHPFHSPSGASSCFVTSLLNLSSGATNLFPWL 972
>TC90469 similar to GP|11066213|gb|AAG28503.1 hexokinase {Citrus sinensis},
partial (33%)
Length = 636
Score = 27.7 bits (60), Expect = 2.3
Identities = 16/41 (39%), Positives = 23/41 (56%), Gaps = 1/41 (2%)
Frame = -1
Query: 98 SIHKVSRLSEASRLIPWMANKLLNERRST-NILRTYQTPQH 137
+IH +S+LS+ S + KLL R +T + RTY T H
Sbjct: 315 NIHSISQLSKLSNWCLTLFPKLLQNRHATCPLSRTYHTVAH 193
>TC89669 homologue to GP|1237082|emb|CAA65540.1 ADP-glucose
pyrophosphorylase {Pisum sativum}, partial (66%)
Length = 1438
Score = 27.3 bits (59), Expect = 3.0
Identities = 26/93 (27%), Positives = 40/93 (42%), Gaps = 3/93 (3%)
Frame = -3
Query: 38 SILRQNTIIIKRVQPLTMTLRVINTLTKSHASVKFHALTRSHASVEFH---AFHKVSRLS 94
SI+R ++ I R + + + N T ASV A S+ + A K S+
Sbjct: 551 SIIRPSSSIFIRPKAVARFSSIGNAATVISASVSLCAWINFS*SIRYKWSPARIKYSKTL 372
Query: 95 GVSSIHKVSRLSEASRLIPWMANKLLNERRSTN 127
S HK + A +PW + LLN+R S +
Sbjct: 371 CSSKSHKYCLTASA---VPWKHHTLLNKRHSNH 282
>TC79332 similar to PIR|JC6203|JC6203 SP8 binding protein homolog -
cucumber, partial (49%)
Length = 1486
Score = 27.3 bits (59), Expect = 3.0
Identities = 27/91 (29%), Positives = 41/91 (44%)
Frame = +2
Query: 11 TTSPTSLDTSPRVLAKLMSLKAYPKVKSILRQNTIIIKRVQPLTMTLRVINTLTKSHASV 70
TT+P+S DTSP ++L P V++ N V P MTL +++ + ++
Sbjct: 50 TTTPSSADTSPPPPRPTITLPPRPSVEAFFTSNA-----VSPGPMTL--VSSFFATESAT 208
Query: 71 KFHALTRSHASVEFHAFHKVSRLSGVSSIHK 101
L + AS AF S L+G S K
Sbjct: 209 FSQLLAGAMASP--LAFSSSSSLAGEYSFGK 295
>TC78622 weakly similar to GP|14587220|dbj|BAB61154. putative GTPase {Oryza
sativa (japonica cultivar-group)}, partial (32%)
Length = 1025
Score = 26.9 bits (58), Expect = 4.0
Identities = 29/125 (23%), Positives = 55/125 (43%), Gaps = 5/125 (4%)
Frame = +2
Query: 42 QNTIIIKRVQPLTMTLRVINTLTKSHAS--VKFHALTRSHASVEFHAFHKVSRLSGVSSI 99
QN + + R M + + N + + A +K H + + + ++ + + + G
Sbjct: 422 QNKMKVLRTLMTMMRMNLWNVVKRMLARKRLKVHLVNKMRSYIQRKVY--LIQN*GEQRR 595
Query: 100 HKVSRLSEASRLIPW---MANKLLNERRSTNILRTYQTPQHLDPTTLLVFPKLWFDFQCL 156
K RL++ + LIPW M +KL+ +R+ R T + T +L+ K CL
Sbjct: 596 RKRRRLTKPAHLIPWMVIMTSKLITSKRT----RWMLTGVMVVTTMMLMTNKSTLRLPCL 763
Query: 157 PSRLL 161
RL+
Sbjct: 764 VLRLM 778
>BG587109 homologue to GP|22091671|em photosystem1 subunit A {Pinus
parviflora}, partial (29%)
Length = 791
Score = 26.6 bits (57), Expect = 5.2
Identities = 18/81 (22%), Positives = 34/81 (41%), Gaps = 7/81 (8%)
Frame = +2
Query: 74 ALTRSHASVEFHA-----FHKVSRLSGVSSIHKV--SRLSEASRLIPWMANKLLNERRST 126
AL+R + H H+V + S+ KV + + S + W++ + R +
Sbjct: 77 ALSRDSCNGSLHTSEV*TMHRVIPVIWRKSLEKVFSAHFGQLSIIFLWLSGMYFHGARFS 256
Query: 127 NILRTYQTPQHLDPTTLLVFP 147
N P H+ P+ +V+P
Sbjct: 257 NYEACLSDPTHIGPSAQVVWP 319
>TC77186 phosphate transporter
Length = 2314
Score = 26.6 bits (57), Expect = 5.2
Identities = 13/48 (27%), Positives = 24/48 (49%)
Frame = -1
Query: 46 IIKRVQPLTMTLRVINTLTKSHASVKFHALTRSHASVEFHAFHKVSRL 93
I+ +QP T TL I + TK H + + + A ++ + K S++
Sbjct: 2224 ILNSIQPKTYTLHSIESYTKKHRTASLL*IQQPVAKMQTEKWFKSSKM 2081
>CB894147 similar to GP|15375301|gb| putative c-myb-like transcription
factor MYB3R-5 {Arabidopsis thaliana}, partial (7%)
Length = 809
Score = 26.2 bits (56), Expect = 6.8
Identities = 22/68 (32%), Positives = 29/68 (42%), Gaps = 6/68 (8%)
Frame = +1
Query: 8 ISQTTSPTS------LDTSPRVLAKLMSLKAYPKVKSILRQNTIIIKRVQPLTMTLRVIN 61
I TT P + +T +L K + YP SI R+ VQPLT +V+
Sbjct: 28 IGYTTPPRARVSELCTETPESILRK--AANTYPNTPSIFRRRRT---GVQPLTSPSKVLK 192
Query: 62 TLTKSHAS 69
SHAS
Sbjct: 193VDNDSHAS 216
>TC84004 type IIB calcium ATPase [Medicago truncatula]
Length = 3655
Score = 26.2 bits (56), Expect = 6.8
Identities = 11/28 (39%), Positives = 17/28 (60%)
Frame = -3
Query: 2 SACNAVISQTTSPTSLDTSPRVLAKLMS 29
++CNA S SP + TSP + KL++
Sbjct: 2534 ASCNAGASFVPSPVTATTSPNIFLKLVT 2451
>TC83336 similar to GP|13938440|gb|AAH07361.1 Similar to protein phosphatase
1G (formerly 2C) magnesium-dependent gamma isoform
{Homo sapiens}, partial (5%)
Length = 655
Score = 26.2 bits (56), Expect = 6.8
Identities = 15/39 (38%), Positives = 22/39 (55%), Gaps = 2/39 (5%)
Frame = +1
Query: 93 LSGVSSIHKVSRLSEASRLI--PWMANKLLNERRSTNIL 129
LS +S +S LS L+ PWM NKL + +RS ++
Sbjct: 4 LSANASFFTLSLLSLLLLLLHHPWMCNKLFSNQRSLRLI 120
>NP501869 NP501869|AY115844.1|AAM55305.1 auxin influx carrier protein
[Medicago truncatula]
Length = 1449
Score = 25.8 bits (55), Expect = 8.9
Identities = 17/46 (36%), Positives = 23/46 (49%)
Frame = -3
Query: 88 HKVSRLSGVSSIHKVSRLSEASRLIPWMANKLLNERRSTNILRTYQ 133
HKV L SS V R+SE LIP+ ER++ +L T +
Sbjct: 304 HKVH*LDKQSSFQ*VHRISETLCLIPYQV-----ERKNMEVLTTLE 182
>TC78474 homologue to PIR|T09550|T09550 phosphoprotein phosphatase (EC
3.1.3.16) 1 catalytic epsilon chain - alfalfa, complete
Length = 1597
Score = 25.8 bits (55), Expect = 8.9
Identities = 15/44 (34%), Positives = 24/44 (54%)
Frame = +1
Query: 120 LNERRSTNILRTYQTPQHLDPTTLLVFPKLWFDFQCLPSRLLSL 163
+N ++ N+L+ +Q + T+LV +W C PSR LSL
Sbjct: 934 MNFLQTDNLLQYFQLQTTVVNLTMLVQ**VWTRT*CAPSRFLSL 1065
>BG584691 similar to PIR|E87601|E876 OmpA family protein [imported] -
Caulobacter crescentus, partial (10%)
Length = 871
Score = 25.8 bits (55), Expect = 8.9
Identities = 18/54 (33%), Positives = 24/54 (44%), Gaps = 6/54 (11%)
Frame = +3
Query: 117 NKLLNERRSTNILRTYQT----PQHLDPTTLLVFPKL--WFDFQCLPSRLLSLL 164
NK NER+ +LRT+ T L+FP+L W F C L L+
Sbjct: 588 NKNTNERKQVGLLRTFHTIGLGTSPSPKKEQLLFPRLRNWCIFLCFFGNLALLV 749
>TC82522 similar to PIR|A96708|A96708 hypothetical protein T2E12.9
[imported] - Arabidopsis thaliana, partial (33%)
Length = 727
Score = 25.8 bits (55), Expect = 8.9
Identities = 22/71 (30%), Positives = 32/71 (44%), Gaps = 4/71 (5%)
Frame = -3
Query: 52 PLTMTLRVINTLTKSHASV----KFHALTRSHASVEFHAFHKVSRLSGVSSIHKVSRLSE 107
PL++T N+ T S + V K L R+H S++ FH VS +S + +SE
Sbjct: 203 PLSLTPN*SNSKTHSFSQVY*ISKSKTLKRAHTSIQKCLFHTVSTYLHTNSDFFPNLISE 24
Query: 108 ASRLIPWMANK 118
W NK
Sbjct: 23 N----VWNVNK 3
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.131 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,300,908
Number of Sequences: 36976
Number of extensions: 69099
Number of successful extensions: 452
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 451
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 452
length of query: 170
length of database: 9,014,727
effective HSP length: 89
effective length of query: 81
effective length of database: 5,723,863
effective search space: 463632903
effective search space used: 463632903
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0247.8


