

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0239.11
(307 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
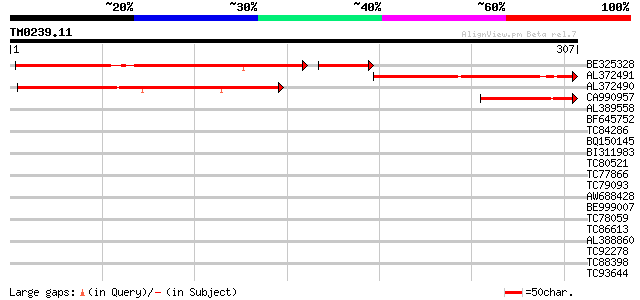
Score E
Sequences producing significant alignments: (bits) Value
BE325328 weakly similar to PIR|C84914|C849 hypothetical protein ... 184 3e-55
AL372491 130 5e-31
AL372490 similar to PIR|C84914|C849 hypothetical protein At2g473... 126 9e-30
CA990957 weakly similar to PIR|C84914|C849 hypothetical protein ... 82 3e-16
AL389558 similar to PIR|T30826|T30 nascent polypeptide-associate... 31 0.52
BF645752 homologue to GP|22713420|gb| Unknown (protein for MGC:3... 31 0.68
TC84286 similar to GP|9294063|dbj|BAB02020.1 beta-glucosidase {A... 31 0.68
BQ150145 31 0.68
BI311983 similar to PIR|E84671|E846 hypothetical protein At2g273... 30 0.89
TC80521 weakly similar to GP|18377712|gb|AAL67006.1 unknown prot... 30 1.2
TC77866 similar to GP|18266655|gb|AAL67601.1 putative cinnamoyl-... 30 1.5
TC79093 similar to GP|4102190|gb|AAD01431.1| 35 kDa seed maturat... 30 1.5
AW688428 29 2.0
BE999007 similar to GP|10176957|dbj gene_id:MHJ24.7~unknown prot... 29 2.6
TC78059 similar to GP|4206787|gb|AAD11808.1| syntaxin-related pr... 28 3.4
TC86613 similar to GP|22136866|gb|AAM91777.1 unknown protein {Ar... 28 3.4
AL388860 homologue to PIR|D86364|D863 hypothetical protein AAB72... 28 3.4
TC92278 similar to SP|O43093|KINH_SYNRA Kinesin heavy chain (Syn... 28 4.4
TC88398 similar to GP|9759187|dbj|BAB09724.1 importin beta {Arab... 28 4.4
TC93644 28 4.4
>BE325328 weakly similar to PIR|C84914|C849 hypothetical protein At2g47360
[imported] - Arabidopsis thaliana, partial (9%)
Length = 593
Score = 184 bits (466), Expect(2) = 3e-55
Identities = 101/160 (63%), Positives = 118/160 (73%), Gaps = 2/160 (1%)
Frame = +1
Query: 4 WQQKAKEFFSNHGGAPTPKATFHLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTLFLHS 63
WQ+K+KEFF N+ TP FHLLTIT+L+LLLPLSF LLASLS AK YLQ +HS
Sbjct: 13 WQKKSKEFFFNYIAIYTPTRIFHLLTITILTLLLPLSFLLLASLSGAKFYLQ-----IHS 177
Query: 64 PQQQQQPFSFLFTLALQINPCILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPSLY 123
QQP +LF+ A+ +PCILYVLVSIVS+ATLINGLMGKI L+NDSS S +LQ LY
Sbjct: 178 ----QQPLPYLFSFAIHTSPCILYVLVSIVSVATLINGLMGKINLMNDSSNSILLQTRLY 345
Query: 124 IA--WILLCAFQFCVGLGIEASIAAGIFEFDNSSSSSSSS 161
++ WILL FQ CVGLGIEASIAAG+F+ DN SS
Sbjct: 346 MSWKWILLFTFQICVGLGIEASIAAGVFDSDNDVFDDGSS 465
Score = 49.3 bits (116), Expect(2) = 3e-55
Identities = 22/30 (73%), Positives = 26/30 (86%)
Frame = +3
Query: 168 ERSLLSKVFFLLGLHETTQVWYRVVVRPVV 197
ERS +S++ FLLGLHETTQ+W RVVVR VV
Sbjct: 477 ERSFISRMIFLLGLHETTQIWSRVVVRTVV 566
>AL372491
Length = 481
Score = 130 bits (328), Expect = 5e-31
Identities = 66/112 (58%), Positives = 89/112 (78%), Gaps = 2/112 (1%)
Frame = +3
Query: 198 DDTVFGGA-KKEKWIERVAVAASLGVLWWWRLREEVESLVVMAEAKKSEQFGELELKDFI 256
DDTVFGG +KE+W+E++ V LG+LWWW+LR++VE+LVVM+EAKK +Q ++ + DF+
Sbjct: 3 DDTVFGGVIRKERWVEKLGVDMCLGILWWWKLRDDVENLVVMSEAKK-DQLMDVGINDFV 179
Query: 257 GWWLYYVTVTIGMVRIVKGLMWMF-MVALCRRRVTEVPVSPVVESSLNDDKV 307
GW LYY+TVTIGMV++VKGLMW+ M+ CRR E+ VS +VE S NDDKV
Sbjct: 180 GWCLYYLTVTIGMVKVVKGLMWILAMICPCRR---EMGVS-MVEHSENDDKV 323
>AL372490 similar to PIR|C84914|C849 hypothetical protein At2g47360
[imported] - Arabidopsis thaliana, partial (5%)
Length = 451
Score = 126 bits (317), Expect = 9e-30
Identities = 73/151 (48%), Positives = 93/151 (61%), Gaps = 7/151 (4%)
Frame = +1
Query: 5 QQKAKEFFSNHGGAPTPKATFHLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTLFLHSP 64
+ K K F + TP+ F+ T TLLSL LPLSF LLA L ++YLQ F H
Sbjct: 1 EHKTKNLFHLFASSITPREIFYFFTHTLLSLFLPLSFLLLAKLLEIQYYLQRIN-FWHQK 177
Query: 65 QQQQQP--FSFLFTLALQINPCILYVLVSIVSIATLINGLMGKITLLNDSS-----TSPV 117
P FS++ LAL INP ILY LV I+SI++LI+ L KI L+++S +S +
Sbjct: 178 YYYSSPQNFSYILKLALHINPFILYFLVFIISISSLIHALTNKIINLSNNSLTSSCSSNI 357
Query: 118 LQPSLYIAWILLCAFQFCVGLGIEASIAAGI 148
++P LYIAWILLC FQ CVGLGIE SI G+
Sbjct: 358 VRPRLYIAWILLCVFQVCVGLGIEGSILVGL 450
>CA990957 weakly similar to PIR|C84914|C849 hypothetical protein At2g47360
[imported] - Arabidopsis thaliana, partial (9%)
Length = 361
Score = 81.6 bits (200), Expect = 3e-16
Identities = 37/52 (71%), Positives = 45/52 (86%)
Frame = +1
Query: 256 IGWWLYYVTVTIGMVRIVKGLMWMFMVALCRRRVTEVPVSPVVESSLNDDKV 307
+GWWLYY+TVTIGMVRIVKGLMW+FM++LCRRRVT++ +E S NDDKV
Sbjct: 1 VGWWLYYLTVTIGMVRIVKGLMWIFMISLCRRRVTQIN-EVELELSQNDDKV 153
>AL389558 similar to PIR|T30826|T30 nascent polypeptide-associated complex
alpha chain muscle splice form gp220 - mouse, partial
(3%)
Length = 493
Score = 31.2 bits (69), Expect = 0.52
Identities = 21/59 (35%), Positives = 31/59 (51%)
Frame = +3
Query: 64 PQQQQQPFSFLFTLALQINPCILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPSL 122
P + Q+ S F + + INP IL + + + I L K+TLLN+S P L+P L
Sbjct: 309 PSEDQKTSSLSFLILMFINPPILTATLCSAKLKSRILILKLKLTLLNNSKL-PKLRPLL 482
>BF645752 homologue to GP|22713420|gb| Unknown (protein for MGC:33586) {Homo
sapiens}, partial (1%)
Length = 321
Score = 30.8 bits (68), Expect = 0.68
Identities = 14/30 (46%), Positives = 19/30 (62%)
Frame = -1
Query: 133 QFCVGLGIEASIAAGIFEFDNSSSSSSSSF 162
Q+C+ I + + F F+NS SSSSSSF
Sbjct: 156 QYCLPTSIIWKVGSNCFSFNNSRSSSSSSF 67
>TC84286 similar to GP|9294063|dbj|BAB02020.1 beta-glucosidase {Arabidopsis
thaliana}, partial (9%)
Length = 844
Score = 30.8 bits (68), Expect = 0.68
Identities = 18/44 (40%), Positives = 23/44 (51%)
Frame = -2
Query: 85 ILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPSLYIAWIL 128
I+YV SI+ ING + KI L PV SL+ +WIL
Sbjct: 255 IIYVECSILIDTNKINGALHKIHFLLSKLWKPVPNNSLHSSWIL 124
>BQ150145
Length = 538
Score = 30.8 bits (68), Expect = 0.68
Identities = 15/44 (34%), Positives = 24/44 (54%)
Frame = +1
Query: 19 PTPKATFHLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTLFLH 62
P+P +FHL + LSLL L F +A + A+ Y+ + F +
Sbjct: 13 PSPYRSFHLFALLFLSLLCALVFLSIAHILYARIYVISSRFFTY 144
>BI311983 similar to PIR|E84671|E846 hypothetical protein At2g27320
[imported] - Arabidopsis thaliana, partial (16%)
Length = 593
Score = 30.4 bits (67), Expect = 0.89
Identities = 17/46 (36%), Positives = 26/46 (55%)
Frame = -3
Query: 148 IFEFDNSSSSSSSSFGESAAERSLLSKVFFLLGLHETTQVWYRVVV 193
+F+FD S+ S S G S A+ S F ++ LHET W +++V
Sbjct: 273 LFDFDRSNIFCS*SNGSSGAQDRSKSIFFVMIDLHET---WTKLIV 145
>TC80521 weakly similar to GP|18377712|gb|AAL67006.1 unknown protein
{Arabidopsis thaliana}, partial (57%)
Length = 1025
Score = 30.0 bits (66), Expect = 1.2
Identities = 13/33 (39%), Positives = 19/33 (57%)
Frame = -3
Query: 11 FFSNHGGAPTPKATFHLLTITLLSLLLPLSFFL 43
F + + T K+ FH + T + LLLPLSF +
Sbjct: 813 FIAKASTSETAKSGFHFINSTTIELLLPLSFLI 715
>TC77866 similar to GP|18266655|gb|AAL67601.1 putative cinnamoyl-CoA
reductase {Oryza sativa}, partial (58%)
Length = 1300
Score = 29.6 bits (65), Expect = 1.5
Identities = 25/109 (22%), Positives = 50/109 (44%)
Frame = +2
Query: 20 TPKATFHLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTLFLHSPQQQQQPFSFLFTLAL 79
T K +HLL L++L + + +L+ K + + F+H ++ S F L L
Sbjct: 194 THKRIWHLLLTLLVTLFVSWMLQVN*ALALFKDFFKEAIPFMHPFNNMEKIHSMGFQLIL 373
Query: 80 QINPCILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPSLYIAWIL 128
+ L+ I++ L+ + T LN T+P++ ++++ W L
Sbjct: 374 INSRFFDQTLLIIIA*LMLLEVALDCFTHLNLPLTNPIMM-NIWLTWKL 517
>TC79093 similar to GP|4102190|gb|AAD01431.1| 35 kDa seed maturation protein
{Glycine max}, partial (57%)
Length = 1127
Score = 29.6 bits (65), Expect = 1.5
Identities = 23/59 (38%), Positives = 25/59 (41%), Gaps = 2/59 (3%)
Frame = -3
Query: 15 HGGAPTPKATFHLLTITLLS--LLLPLSFFLLASLSAAKHYLQTFTLFLHSPQQQQQPF 71
H P+P L T L S LLP S L SL + YLQ FL QQ Q F
Sbjct: 660 HPSLPSPSLQHSLYTPLLYSQWYLLPPSLSLQHSLCIPRLYLQLCPSFLWLSQQHNQHF 484
>AW688428
Length = 404
Score = 29.3 bits (64), Expect = 2.0
Identities = 11/26 (42%), Positives = 13/26 (49%)
Frame = -3
Query: 203 GGAKKEKWIERVAVAASLGVLWWWRL 228
GG + W R V S V+WWW L
Sbjct: 114 GGCGGDGWRRRRVVCGSPAVVWWWML 37
>BE999007 similar to GP|10176957|dbj gene_id:MHJ24.7~unknown protein
{Arabidopsis thaliana}, partial (20%)
Length = 484
Score = 28.9 bits (63), Expect = 2.6
Identities = 23/46 (50%), Positives = 28/46 (60%), Gaps = 1/46 (2%)
Frame = +2
Query: 20 TPKATFHLLTITLL-SLLLPLSFFLLASLSAAKHYLQTFTLFLHSP 64
TP++ LL+ LL SLLLPL F L +SLS A L+ F L SP
Sbjct: 299 TPRSLSTLLSHRLLRSLLLPLRFPLRSSLSLA---LRHFPLLRSSP 427
>TC78059 similar to GP|4206787|gb|AAD11808.1| syntaxin-related protein Nt-syr1
{Nicotiana tabacum}, partial (94%)
Length = 1595
Score = 28.5 bits (62), Expect = 3.4
Identities = 18/76 (23%), Positives = 38/76 (49%)
Frame = -2
Query: 14 NHGGAPTPKATFHLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTLFLHSPQQQQQPFSF 73
N+ G +P + HL+ + + LL P S +++ A + +Q FTL L+ + +
Sbjct: 1021 NYDGNTSPFSGIHLVLPSNMQLLDPTSNIRISTCHVAFNIIQLFTL*LNQNRHVHKHLVK 842
Query: 74 LFTLALQINPCILYVL 89
L ++ + C++ +L
Sbjct: 841 LH*ISFYLFHCVMPLL 794
>TC86613 similar to GP|22136866|gb|AAM91777.1 unknown protein {Arabidopsis
thaliana}, partial (75%)
Length = 2134
Score = 28.5 bits (62), Expect = 3.4
Identities = 19/58 (32%), Positives = 26/58 (44%), Gaps = 4/58 (6%)
Frame = +2
Query: 21 PKATFHLLTITLLSLLLPLSFFLLASLSAAK----HYLQTFTLFLHSPQQQQQPFSFL 74
P T +L + +L LL F AS+ + H+LQ L H P+ Q P S L
Sbjct: 1034 PAVTSFMLPVWILGLLRGEPFAQFASVMQERVWLIHHLQNRHLCCHQPRHLQHPLSCL 1207
>AL388860 homologue to PIR|D86364|D863 hypothetical protein AAB72157.1
[imported] - Arabidopsis thaliana, partial (3%)
Length = 526
Score = 28.5 bits (62), Expect = 3.4
Identities = 19/41 (46%), Positives = 22/41 (53%)
Frame = -2
Query: 137 GLGIEASIAAGIFEFDNSSSSSSSSFGESAAERSLLSKVFF 177
GLG+ +S A IF F SS SSSS SA SL + F
Sbjct: 342 GLGVCSSSIASIFAFRLSSLSSSSFNAPSALSISLSISLIF 220
>TC92278 similar to SP|O43093|KINH_SYNRA Kinesin heavy chain (Synkin).
{Syncephalastrum racemosum}, partial (10%)
Length = 709
Score = 28.1 bits (61), Expect = 4.4
Identities = 23/76 (30%), Positives = 38/76 (49%), Gaps = 3/76 (3%)
Frame = -1
Query: 89 LVSIVSIATLINGLMGKITLLNDSS---TSPVLQPSLYIAWILLCAFQFCVGLGIEASIA 145
+VS++SI L+ I L N+SS +S V + S +++ C+F C L + +
Sbjct: 358 IVSVISI--LLFASSSAIVLSNNSSYSSSSVVKRISSSSLFLVKCSFS*CNSLIVCSCCV 185
Query: 146 AGIFEFDNSSSSSSSS 161
+F+ DNS SS
Sbjct: 184 TKLFDSDNSRRKVESS 137
>TC88398 similar to GP|9759187|dbj|BAB09724.1 importin beta {Arabidopsis
thaliana}, partial (41%)
Length = 1208
Score = 28.1 bits (61), Expect = 4.4
Identities = 19/50 (38%), Positives = 24/50 (48%)
Frame = +1
Query: 18 APTPKATFHLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTLFLHSPQQQ 67
+P T L + LL LL LSF + L+ HY Q FT FL +Q
Sbjct: 421 SPLLYLTLGLQQLKLLLKLLELSFLKNSGLN**DHYYQIFTKFLLMSSKQ 570
>TC93644
Length = 488
Score = 28.1 bits (61), Expect = 4.4
Identities = 25/70 (35%), Positives = 31/70 (43%), Gaps = 16/70 (22%)
Frame = +2
Query: 83 PCILYVLVSIVSIATLINGLMGKITLLNDSS------------TSPVLQPSLYIAW---- 126
P L VLVS+VSI T+I L K N SS T+ L + W
Sbjct: 245 PY*LRVLVSLVSIYTVIF*LFAKYKSTNTSSMM*QACVFGFLLTARFLVELKCLRWRYGV 424
Query: 127 ILLCAFQFCV 136
+LLC FQF +
Sbjct: 425 LLLCGFQFSI 454
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.137 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,566,952
Number of Sequences: 36976
Number of extensions: 202276
Number of successful extensions: 2481
Number of sequences better than 10.0: 61
Number of HSP's better than 10.0 without gapping: 2242
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2387
length of query: 307
length of database: 9,014,727
effective HSP length: 96
effective length of query: 211
effective length of database: 5,465,031
effective search space: 1153121541
effective search space used: 1153121541
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0239.11


