

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
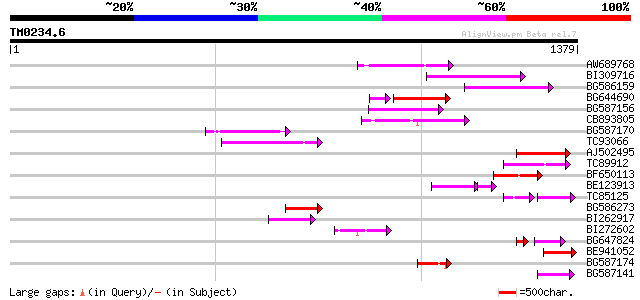
Query= TM0234.6
(1379 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsi... 204 3e-52
BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F2... 181 2e-45
BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2... 166 8e-41
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 122 2e-36
BG587156 similar to PIR|G85055|G8 probable polyprotein [imported... 142 7e-34
CB893805 similar to GP|10177935|d copia-type polyprotein {Arabid... 139 6e-33
BG587170 similar to PIR|F86470|F8 probable retroelement polyprot... 134 2e-31
TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative ret... 132 7e-31
AJ502495 weakly similar to GP|18071369|g putative gag-pol polypr... 128 1e-29
TC89912 weakly similar to PIR|B84512|B84512 probable retroelemen... 104 2e-22
BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vu... 100 3e-21
BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sati... 77 6e-21
TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-relate... 63 4e-20
BG586273 weakly similar to PIR|F86470|F8 probable retroelement p... 94 4e-19
BI262917 weakly similar to GP|19920130|g Putative retroelement {... 82 2e-15
BI272602 GP|11602812|db avenaIII {Gallus gallus}, partial (3%) 75 1e-13
BG647824 weakly similar to PIR|G96722|G96 hypothetical protein F... 55 4e-13
BE941052 weakly similar to PIR|B85188|B85 retrotransposon like p... 71 3e-12
BG587174 similar to PIR|A47759|A4775 retrovirus-related reverse ... 70 6e-12
BG587141 similar to PIR|H86461|H86 hypothetical protein AAF32440... 66 1e-10
>AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsis thaliana},
partial (9%)
Length = 675
Score = 204 bits (518), Expect = 3e-52
Identities = 110/233 (47%), Positives = 135/233 (57%), Gaps = 1/233 (0%)
Frame = +1
Query: 847 TRSMRGIYKPRKLFNLSVTIDDPTISPLPKN-PKLALSDPNWKSAMQSEFDALIRNNTWD 905
TR+ G KP+ P P++ L LSDP W AM++E+ ALI N TWD
Sbjct: 4 TRTKTGKLKPKVFLT----------EPCPQDCENLPLSDPRWLQAMKTEYKALIDNKTWD 153
Query: 906 LVPRPCDVNIIRCMWIFRHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSHVVKPATI 965
LVP P I C W++R K +G ++KARLV G SQ G D ETFS V+KP TI
Sbjct: 154 LVPLPPHKKAIGCKWVYRVKENPDGSVNKFKARLVAKGFSQTLGCDYTETFSPVIKPVTI 333
Query: 966 RTVLTIALSRSWPIHQLDVQNAFLHGDLHETVYMHQPLGFRDPNHPDYVCRLRKSLYGLK 1025
R +LTIA++ W I Q+D+ NAFL+G L E VYM QP GF N VC+L KSLYGLK
Sbjct: 334 RLILTIAITYKWEIQQIDINNAFLNGFLQEEVYMSQPQGFEAAN-KSLVCKLNKSLYGLK 510
Query: 1026 QAPRAWYQCFADYVSTIGFQHSTSDHSLFIYRRGSDMAYLLLYVDDIILISSS 1078
QAPRAWY+ GF S D SL IY + YL +YVDDI++ SS
Sbjct: 511 QAPRAWYEXLTSAQIQFGFTKSRCDPSLLIYNQNGACIYLXIYVDDILITGSS 669
>BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (10%)
Length = 744
Score = 181 bits (459), Expect = 2e-45
Identities = 98/240 (40%), Positives = 137/240 (56%)
Frame = +2
Query: 1014 VCRLRKSLYGLKQAPRAWYQCFADYVSTIGFQHSTSDHSLFIYRRGSDMAYLLLYVDDII 1073
VC L+KS+YGLKQA R WY ++ + + G+ S+SD SLF + S LL+YVDDI+
Sbjct: 20 VCELQKSIYGLKQASRQWYSKLSESLISFGYLQSSSDFSLFTKFKDSSFTTLLVYVDDIV 199
Query: 1074 LISSSHDLRKSIMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYARDIIARAGM 1133
L + + + L F +KDLG L YFLG+ V R G+ L+Q Y +++ +G
Sbjct: 200 LAGNDISEIQHVKCFLIDRFKIKDLGSLRYFLGLEVARSKQGILLNQRKYTLELLEDSGN 379
Query: 1134 ASCHPSATPVDTKQKLSTSAGTPCDDPTLYRSLVGALQYLTFTRPDISYAVQQVCLHMHA 1193
+ + TP D KL S +D T YR L+G L YLT TRPDIS+AVQQ+ +
Sbjct: 380 LAVKSTLTPYDISLKLHNSDSPLYNDETQYRRLIGKLIYLTTTRPDISFAVQQLSQFVSK 559
Query: 1194 PRTEHMLALKRILRYVQGTLQLGLHLYPSPIEKLISYTDADWGGCLDTRRSTSGYCVFLG 1253
P+ H A R+L+Y++ GL + KL S+ D+DW C TR+S +GY VFLG
Sbjct: 560 PQQVHYQAAIRVLQYLKTAPAKGLFYSATSNLKLSSFADSDWATCPTTRKSVTGYWVFLG 739
>BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2O9.150 -
Arabidopsis thaliana, partial (11%)
Length = 732
Score = 166 bits (419), Expect = 8e-41
Identities = 85/217 (39%), Positives = 126/217 (57%)
Frame = +1
Query: 1107 IAVTRHAGGLFLSQSTYARDIIARAGMASCHPSATPVDTKQKLSTSAGTPCDDPTLYRSL 1166
+ V ++ G+++ Q Y D++ R GM + S P+ + KL D T Y+ +
Sbjct: 1 VEVIQNEEGIYICQRKYVTDLLERFGMEKSNLSRNPIAPRCKLIKDENGVKVDATKYKQI 180
Query: 1167 VGALQYLTFTRPDISYAVQQVCLHMHAPRTEHMLALKRILRYVQGTLQLGLHLYPSPIEK 1226
VG L YL TRPD+ Y + + M+ P HM A+KR+LRY+ GT+ LG+ + EK
Sbjct: 181 VGCLMYLAATRPDLMYVLSLISRFMNCPTELHMHAVKRVLRYLNGTINLGIMYKRNGSEK 360
Query: 1227 LISYTDADWGGCLDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSES 1286
L +YTD+D+ G LD R+STSGY L +SWSSK+QP ++ S+ +AE+ A +S
Sbjct: 361 LEAYTDSDYAGDLDDRKSTSGYVFMLSSGAVSWSSKKQPVVTLSTTKAEFIAAAFCACQS 540
Query: 1287 CWLRNLLLELHFPLS*ATLVYCDNVSAIYLSGNPVQH 1323
W+R +L +L + S + +YCDN S I LS NPV H
Sbjct: 541 VWMRRVLEKLGYTQSGSITMYCDNNSTIKLSKNPVLH 651
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 122 bits (306), Expect(2) = 2e-36
Identities = 59/139 (42%), Positives = 92/139 (65%)
Frame = -2
Query: 934 RYKARLVGDGRSQIAGVDCDETFSHVVKPATIRTVLTIALSRSWPIHQLDVQNAFLHGDL 993
R K++LV G +Q G+D DE FS V + IR ++ A + ++Q+DV++AF++GDL
Sbjct: 418 RNKSKLVVQGYNQKEGIDYDEAFSPVARMEAIRILIAFAAFMGFKLYQMDVKSAFINGDL 239
Query: 994 HETVYMHQPLGFRDPNHPDYVCRLRKSLYGLKQAPRAWYQCFADYVSTIGFQHSTSDHSL 1053
E V++ QP GF D P++V RL K+LYGLKQAPRAWY+ + ++ GF+ D++L
Sbjct: 238 KEEVFVKQPPGFEDAEVPNHVFRLNKTLYGLKQAPRAWYERLSKFLLKNGFKRGKIDNTL 59
Query: 1054 FIYRRGSDMAYLLLYVDDI 1072
F+ +R ++ + +YVDDI
Sbjct: 58 FLLKRE*ELLIIQVYVDDI 2
Score = 50.4 bits (119), Expect(2) = 2e-36
Identities = 23/51 (45%), Positives = 30/51 (58%)
Frame = -3
Query: 875 PKNPKLALSDPNWKSAMQSEFDALIRNNTWDLVPRPCDVNIIRCMWIFRHK 925
PKN K AL D +W ++MQ E R+ W LVPRP +I W+FR+K
Sbjct: 585 PKNVKEALRDADWINSMQEELHQFERSKVWYLVPRPEGKTVIGTRWVFRNK 433
>BG587156 similar to PIR|G85055|G8 probable polyprotein [imported] -
Arabidopsis thaliana, partial (17%)
Length = 618
Score = 142 bits (359), Expect = 7e-34
Identities = 75/181 (41%), Positives = 109/181 (59%)
Frame = -1
Query: 874 LPKNPKLALSDPNWKSAMQSEFDALIRNNTWDLVPRPCDVNIIRCMWIFRHKTKANGCFE 933
+P++ + A+ D WK ++ +E A+I+N+TW P + WIF K KA+G E
Sbjct: 561 IPRSYEEAMEDKEWKESVGAEAGAMIKNDTWYESELPKGKKAVSSRWIFTIKYKADGSIE 382
Query: 934 RYKARLVGDGRSQIAGVDCDETFSHVVKPATIRTVLTIALSRSWPIHQLDVQNAFLHGDL 993
R K RLV G + G D ETF+ V K TIR VL++A++ W + Q+DV+NAFL G+L
Sbjct: 381 RKKTRLVARGFTLTYGEDYIETFAPVAKLHTIRIVLSLAVNLGWGLWQMDVKNAFLQGEL 202
Query: 994 HETVYMHQPLGFRDPNHPDYVCRLRKSLYGLKQAPRAWYQCFADYVSTIGFQHSTSDHSL 1053
+ VYM+ P G V RL+K++YGLKQ+PRAWY + ++ GF+ S DH+L
Sbjct: 201 EDEVYMYPPPGLEHLVKRGNVLRLKKAIYGLKQSPRAWYNKLSTTLNGRGFRKSELDHTL 22
Query: 1054 F 1054
F
Sbjct: 21 F 19
>CB893805 similar to GP|10177935|d copia-type polyprotein {Arabidopsis
thaliana}, partial (14%)
Length = 778
Score = 139 bits (351), Expect = 6e-33
Identities = 89/271 (32%), Positives = 134/271 (48%), Gaps = 7/271 (2%)
Frame = +3
Query: 855 KPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNNTWDLVPRPCDVN 914
K + L L++T D T K+ K W+++M +E +A RNNTW+L
Sbjct: 3 KTQNLGMLTMTSDPTTFEEAVKSEK-------WRASMNNEMEATERNNTWELTDLRSGAK 161
Query: 915 IIRCMWIFRHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSHVVKPATIRTVLTIALS 974
I WIF+ K NG E+YKARLV G SQ GVD E F+ V + TIR V+ +A
Sbjct: 162 TIGLKWIFKTKLNENGEIEKYKARLVAKGYSQQYGVDYTEVFAPVARWDTIRMVIALAA- 338
Query: 975 RSWPIHQLDVQNAFL------HGDLHETVYMHQPLGFRDPNHPDYVCRLRKSLYGLKQAP 1028
Q+ + H + + + R R++++LYGLKQAP
Sbjct: 339 ------QIKRDGVCIS*M*KAHSCMEN*MRKFLLINHRVM*RRVIS*RVKRALYGLKQAP 500
Query: 1029 RAWYQCFADYVSTIGFQHSTSDHSLFI-YRRGSDMAYLLLYVDDIILISSSHDLRKSIMA 1087
RAWY Y + GF+ +H+LF+ G + + LYVDD+I I + ++ +
Sbjct: 501 RAWYSRIEAYFTKEGFEKCPYEHTLFVKLSEGGKILIISLYVDDLIFIGNDENMFEEFKK 680
Query: 1088 LLASEFAMKDLGPLSYFLGIAVTRHAGGLFL 1118
+ EF M DLG + YFLG+ VT++ G+++
Sbjct: 681 SMKKEFNMSDLGKMHYFLGVEVTQNEKGIYI 773
>BG587170 similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 718
Score = 134 bits (337), Expect = 2e-31
Identities = 69/206 (33%), Positives = 117/206 (56%)
Frame = -3
Query: 477 SLWHSRLGHPSSSALQSLRSNKFISYEHLNSSPVCESCVFGKHVRLPFVSSNNVTVMPFD 536
+LWH+RLGHP AL + + +E+ N CE+C+ GKH + F ++ V FD
Sbjct: 587 ALWHARLGHPHGRALNLMLPG--VVFENKN----CEACILGKHCKNVFPRTSTVYENCFD 426
Query: 537 ILHSDLWTSPVLSSAGHRFYVLFLDDFTDFLWTFPLSNKSQVFEMFISLSNQIRTHFSQT 596
++++DLWT+P LS H+++V F+D+ + + W + +K +V + F + + H+
Sbjct: 425 LIYTDLWTAPSLSRDNHKYFVTFIDEKSKYTWLTLIPSKDRVIDAFKNFQAYVTNHYHAK 246
Query: 597 IKCLQCDNGREFDNKSFHDYCAANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHA 656
IK L+ DNG E+ + +F + +G++ + SCP+T QNG A+RK + + + R+L+ A
Sbjct: 245 IKILRSDNGGEYTSYAFKSHLDHHGILHQTSCPYTPQQNGVAKRKNKHLMEVARSLMFQA 66
Query: 657 SVPPSFWHHALQMATYLLNIIPRKNL 682
+V A YL+N IP K L
Sbjct: 65 NV---------STACYLINWIPTKVL 15
>TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative retroelement
{Oryza sativa} [Oryza sativa (japonica cultivar-group)],
partial (10%)
Length = 823
Score = 132 bits (333), Expect = 7e-31
Identities = 82/249 (32%), Positives = 124/249 (48%), Gaps = 5/249 (2%)
Frame = +1
Query: 516 FGKHVRLPFVSSNNVTVMPFDILHSDLW-TSPVLSSAGHRFYVLFLDDFTDFLWTFPLSN 574
FG ++ F ++ + T D +HSDLW S V S G R+ + +DDF +W + L
Sbjct: 40 FGNRKKVSFSTATHRTKGILDYIHSDLWGPSKVTSYGGRRYMMTIIDDFPRKVWVYFLRY 219
Query: 575 KSQVFEMFISLSNQIRTHFSQTIKCLQCDNGREFDNKSFHDYCAANGLIFRFSCPHTSSQ 634
K++ F F + T + +K L DN EF + F+++C +G+ + P Q
Sbjct: 220 KNETFPTFKKWRILVETQTGKNVKKLITDN*LEFCSSDFNEFCTNHGIARHKTIPRNPQQ 399
Query: 635 NGKAERKIRTINNMIRTLLAHASV--PPSFWHHALQMATYLLNIIPRKNLSNLSPTQLLY 692
NG AER IRT+ R +L++A + W A A +L+N P L P +
Sbjct: 400 NGVAERMIRTLLERARCMLSNAGL*N*RDLWVEAASTACHLVNRSPHSALDFKVPEDIWS 579
Query: 693 RRDPSYTHLRVFGCLCYPLVPSSTINKLQPRSTPCVFLGYPLHHRGYK--CFDLSHRKVI 750
Y++LR+FGC Y LV KL PR+ C+FL Y +GY+ C D +K+I
Sbjct: 580 GNLVDYSNLRIFGCPAYALVND---GKLAPRAGECIFLSYASESKGYRLWCSDPKSQKLI 750
Query: 751 ISRHVIFDE 759
+SR V F+E
Sbjct: 751 LSRDVTFNE 777
>AJ502495 weakly similar to GP|18071369|g putative gag-pol polyprotein {Oryza
sativa}, partial (9%)
Length = 542
Score = 128 bits (322), Expect = 1e-29
Identities = 62/132 (46%), Positives = 87/132 (64%)
Frame = +2
Query: 1233 ADWGGCLDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWLRNL 1292
+DW G +TR+STSGY LG ISWSSK+QP ++ S+AEAEY + +++ WLR +
Sbjct: 2 SDWAGDTETRKSTSGYAFHLGTGAISWSSKKQPVVAFSTAEAEYIASTSCATQTVWLRRI 181
Query: 1293 LLELHFPLS*ATLVYCDNVSAIYLSGNPVQHQRTKHIEMDIHFVREKVARGQARVLHVPS 1352
L +H + T +YCDN SAI LS NPV H R+KHI++ H +RE +A + + + P+
Sbjct: 182 LEVMHHEQNTPTKIYCDNKSAIALSKNPVFHGRSKHIDIQFHKIRELIAEKEVVIEYCPT 361
Query: 1353 RHQIADIFTKGL 1364
+IADIFTK L
Sbjct: 362 EEKIADIFTKPL 397
>TC89912 weakly similar to PIR|B84512|B84512 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(10%)
Length = 814
Score = 104 bits (260), Expect = 2e-22
Identities = 57/164 (34%), Positives = 95/164 (57%), Gaps = 2/164 (1%)
Frame = +1
Query: 1201 ALKRILRYVQGTLQLGLHLYPSPIEK--LISYTDADWGGCLDTRRSTSGYCVFLGDNLIS 1258
ALK +L+Y+ +L+ L + E+ L Y DAD+ G +DTR+S SG+ L IS
Sbjct: 4 ALKWVLKYLNESLKSSLKYTKAAQEEDALEGYVDADYAGNVDTRKSLSGFVFTLYGTTIS 183
Query: 1259 WSSKRQPTLSRSSAEAEYRGVANVVSESCWLRNLLLELHFPLS*ATLVYCDNVSAIYLSG 1318
W + +Q ++ S+ +AEY V ++ WL+ ++ EL ++CD+ SAI+L+
Sbjct: 184 WKANQQSVVTLSTTQAEYIAFVEGVKDAIWLKGMIGELGITQE-YVKIHCDSQSAIHLAN 360
Query: 1319 NPVQHQRTKHIEMDIHFVREKVARGQARVLHVPSRHQIADIFTK 1362
+ V H+RTKHI++ +HF+R+ + + V + S AD+FTK
Sbjct: 361 HQVYHERTKHIDIRLHFIRDMIESKEIVVEKMASEENPADVFTK 492
>BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vulgaris},
partial (13%)
Length = 494
Score = 100 bits (250), Expect = 3e-21
Identities = 56/124 (45%), Positives = 76/124 (61%), Gaps = 4/124 (3%)
Frame = +1
Query: 1177 RPDISYAVQQVCLHMHAPRTEHMLALKRILRYVQGTLQLGLHLYP----SPIEKLISYTD 1232
RPDI Y+V + MH PR H++A RILRYV+GT++ GL L+P S + +LI Y+D
Sbjct: 121 RPDICYSVSVISKFMHDPRKPHLIAANRILRYVRGTMEYGL-LFPYGAKSEVYELICYSD 297
Query: 1233 ADWGGCLDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWLRNL 1292
+DW G RRSTSGY D ISW +K+QP + SS EAEY ++ WL ++
Sbjct: 298 SDWCG---DRRSTSGYVFKFNDAAISWCTKKQPITALSSYEAEYIAGTFATFQALWLDSV 468
Query: 1293 LLEL 1296
+ EL
Sbjct: 469 IKEL 480
>BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 503
Score = 77.4 bits (189), Expect(2) = 6e-21
Identities = 44/118 (37%), Positives = 59/118 (49%), Gaps = 1/118 (0%)
Frame = +1
Query: 1026 QAPRAWYQCFADYVSTIGFQHSTSDHSLFIYRRGS-DMAYLLLYVDDIILISSSHDLRKS 1084
Q+PR W+ F V G+ +DH++FI + A L++YVDDI L K
Sbjct: 1 QSPRDWFDRFT*VVKKFGYIQCQTDHAMFIKHSSTVKKAILIVYVDDIFLTGDHGK*IKR 180
Query: 1085 IMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYARDIIARAGMASCHPSATP 1142
+ LLA EF +KDLG L YFLG+ V R G +SQ Y D++ M C P
Sbjct: 181 LKNLLAEEFEIKDLGNLKYFLGMEVARWKKGSSISQRKYVLDLLKETRMIGCKTIRDP 354
Score = 43.1 bits (100), Expect(2) = 6e-21
Identities = 23/47 (48%), Positives = 27/47 (56%)
Frame = +2
Query: 1138 PSATPVDTKQKLSTSAGTPCDDPTLYRSLVGALQYLTFTRPDISYAV 1184
PS TP+D KL T D Y+ LVG L YL+ TRPDIS+ V
Sbjct: 341 PSETPMDATVKLGTLDNGTLVDKGRYQRLVGKLIYLSHTRPDISFVV 481
>TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
(EC 3.4.23.-);, partial (7%)
Length = 705
Score = 62.8 bits (151), Expect(2) = 4e-20
Identities = 35/91 (38%), Positives = 50/91 (54%)
Frame = +3
Query: 1285 ESCWLRNLLLELHFPLS*ATLVYCDNVSAIYLSGNPVQHQRTKHIEMDIHFVREKVARGQ 1344
E+ W++ L+ EL T VYCD+ SA++++ NP H RTKHI + HFVRE V G
Sbjct: 249 EAIWMQRLMEELGHKQEQIT-VYCDSQSALHIARNPAFHSRTKHIGIQYHFVREVVEEGS 425
Query: 1345 ARVLHVPSRHQIADIFTKGLPRVLFDDFRSS 1375
+ + + +AD TK + F RSS
Sbjct: 426 VDMQKIHTNDNLADAMTKSINTDKFIWCRSS 518
Score = 55.1 bits (131), Expect(2) = 4e-20
Identities = 29/75 (38%), Positives = 45/75 (59%)
Frame = +1
Query: 1202 LKRILRYVQGTLQLGLHLYPSPIEKLISYTDADWGGCLDTRRSTSGYCVFLGDNLISWSS 1261
+KRI+RY++GT + + S + + Y D+D+ G D R+ST+GY L +SW S
Sbjct: 1 VKRIMRYIKGTSGVAVCFGGSELT-VRGYVDSDFAGDHDKRKSTTGYVFTLAGGAVSWLS 177
Query: 1262 KRQPTLSRSSAEAEY 1276
K Q ++ S+ EAEY
Sbjct: 178 KLQTVVALSTTEAEY 222
>BG586273 weakly similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 705
Score = 94.0 bits (232), Expect = 4e-19
Identities = 44/90 (48%), Positives = 61/90 (66%)
Frame = -2
Query: 670 ATYLLNIIPRKNLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSSTINKLQPRSTPCVF 729
A YL+N IP + L + +P ++L +R PS T++RVFGCLCY LVP NKL+ RS +F
Sbjct: 704 ACYLINRIPTRVLKDQAPFEVLNQRKPSLTYMRVFGCLCYVLVPGELRNKLEARSRKAMF 525
Query: 730 LGYPLHHRGYKCFDLSHRKVIISRHVIFDE 759
+GY +GYKC+D R+V++SR V F E
Sbjct: 524 IGYSTTQKGYKCYDPEARRVLVSRDVKFIE 435
>BI262917 weakly similar to GP|19920130|g Putative retroelement {Oryza
sativa} [Oryza sativa (japonica cultivar-group)],
partial (8%)
Length = 426
Score = 81.6 bits (200), Expect = 2e-15
Identities = 42/114 (36%), Positives = 61/114 (52%)
Frame = +1
Query: 630 HTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKNLSNLSPTQ 689
HT QNG AER RT+ R +L A + SFW A++ A Y++N P + +P +
Sbjct: 82 HTPQQNGVAERMNRTLLERTRAMLKTAGMAKSFWAEAVKTACYVINRSPSTVIDLKTPME 261
Query: 690 LLYRRDPSYTHLRVFGCLCYPLVPSSTINKLQPRSTPCVFLGYPLHHRGYKCFD 743
+ + Y+ L VFGC Y + S KL P+S C+FLGY + +GY +D
Sbjct: 262 MWKGKPVDYSSLHVFGCPVYVMYNSQERTKLDPKSRKCIFLGYADNVKGYXLWD 423
>BI272602 GP|11602812|db avenaIII {Gallus gallus}, partial (3%)
Length = 488
Score = 75.5 bits (184), Expect = 1e-13
Identities = 51/148 (34%), Positives = 71/148 (47%), Gaps = 10/148 (6%)
Frame = +1
Query: 791 TTQTPSPDLQPTPVAPSATATPPTSTASSSSPSDPSPSSSTPQSPPQPAPPVRT------ 844
+TQT SP +P + S+ +S + ++ + +++P PP P P R
Sbjct: 7 STQTSSPS---SPNSNEVAQNIQISSPASPTSNELANITTSPTPPPPPPPTARPHNNHTT 177
Query: 845 ----MATRSMRGIYKPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIR 900
+ TRS I K K NL V P P N AL D +W+S MQ EFDAL
Sbjct: 178 NVGGIITRSQNNIVKTIKKLNLHVR---PLCPIEPSNITQALRDHDWRSIMQDEFDALQN 348
Query: 901 NNTWDLVPRPCDVNIIRCMWIFRHKTKA 928
NNTWDLV R N++ C F ++TK+
Sbjct: 349 NNTWDLVSRSFAQNLVGCKLGFSYQTKS 432
>BG647824 weakly similar to PIR|G96722|G96 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (5%)
Length = 721
Score = 55.5 bits (132), Expect(2) = 4e-13
Identities = 30/78 (38%), Positives = 44/78 (55%), Gaps = 1/78 (1%)
Frame = -3
Query: 1276 YRGVANVVSESCWLRNLLLELHFPLS*ATLVYCDNVSAI-YLSGNPVQHQRTKHIEMDIH 1334
YR + + + E WL LL +L F ++YCDN SA +++ N +RTKHIE+D H
Sbjct: 374 YRSI*STICEIKWLTYLLNDLKFTFIKPAMLYCDNQSAARHIAANSSFLERTKHIELDCH 195
Query: 1335 FVREKVARGQARVLHVPS 1352
VR K+ +LH+ S
Sbjct: 194 IVRVKLQLKLFHILHILS 141
Score = 38.5 bits (88), Expect(2) = 4e-13
Identities = 16/28 (57%), Positives = 20/28 (71%)
Frame = -2
Query: 1234 DWGGCLDTRRSTSGYCVFLGDNLISWSS 1261
D CLDT +S S +C+FLGD+LI W S
Sbjct: 486 D*SSCLDT*KSISYFCIFLGDSLICWKS 403
>BE941052 weakly similar to PIR|B85188|B85 retrotransposon like protein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 480
Score = 71.2 bits (173), Expect = 3e-12
Identities = 36/80 (45%), Positives = 51/80 (63%)
Frame = +2
Query: 1299 PLS*ATLVYCDNVSAIYLSGNPVQHQRTKHIEMDIHFVREKVARGQARVLHVPSRHQIAD 1358
P+S + L+ CD +SA YL+ NPV H R KHI +DIHFVR+ V +G+ +V HV + Q+AD
Sbjct: 2 PISQSGLLRCDYLSATYLTHNPVYHSRMKHISIDIHFVRDLVQQGKLKVQHVCTVDQLAD 181
Query: 1359 IFTKGLPRVLFDDFRSSLSV 1378
TK L + R+ + V
Sbjct: 182 CLTKPLSKSRHQLLRNKIGV 241
>BG587174 similar to PIR|A47759|A4775 retrovirus-related reverse transcriptase
homolog - rape retrotransposon copia-like (fragment),
partial (84%)
Length = 249
Score = 70.1 bits (170), Expect = 6e-12
Identities = 39/87 (44%), Positives = 53/87 (60%), Gaps = 5/87 (5%)
Frame = -1
Query: 992 DLHETVYMHQPLGFRDPNHPDYVCRLRKSLYGLKQAPRAWYQCFADYVSTIGFQHSTSDH 1051
+L E +YM QP GF P D+VC+LRKSLYGLKQ+PR WY+ F Y S+ +T+
Sbjct: 249 ELEEKIYMTQPEGFLFPGKEDHVCKLRKSLYGLKQSPRQWYKRFDSYRSS----WATTGV 82
Query: 1052 SLFIYR-----RGSDMAYLLLYVDDII 1073
+ + R S YL+LYVDD++
Sbjct: 81 LMTVVST*TR*RMSRYIYLVLYVDDML 1
>BG587141 similar to PIR|H86461|H86 hypothetical protein AAF32440.1 [imported]
- Arabidopsis thaliana, partial (20%)
Length = 731
Score = 65.9 bits (159), Expect = 1e-10
Identities = 36/89 (40%), Positives = 49/89 (54%)
Frame = +3
Query: 1285 ESCWLRNLLLELHFPLS*ATLVYCDNVSAIYLSGNPVQHQRTKHIEMDIHFVREKVARGQ 1344
++ WL++LL E+ + ++ DN S I L+ NPV H R HI HF+RE V GQ
Sbjct: 135 QAMWLQDLLSEVTWEPCEEVVIRIDNQSVIALTRNPVFHGRGNHIHKRYHFIRECVENGQ 314
Query: 1345 ARVLHVPSRHQIADIFTKGLPRVLFDDFR 1373
V HVP A I TK L R++F + R
Sbjct: 315 VEVEHVPGEKHRAYI*TKALGRIIFREIR 401
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.329 0.141 0.453
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 54,260,297
Number of Sequences: 36976
Number of extensions: 1037038
Number of successful extensions: 15077
Number of sequences better than 10.0: 503
Number of HSP's better than 10.0 without gapping: 7513
Number of HSP's successfully gapped in prelim test: 325
Number of HSP's that attempted gapping in prelim test: 1598
Number of HSP's gapped (non-prelim): 11059
length of query: 1379
length of database: 9,014,727
effective HSP length: 108
effective length of query: 1271
effective length of database: 5,021,319
effective search space: 6382096449
effective search space used: 6382096449
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0234.6


