

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0234.12
(607 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
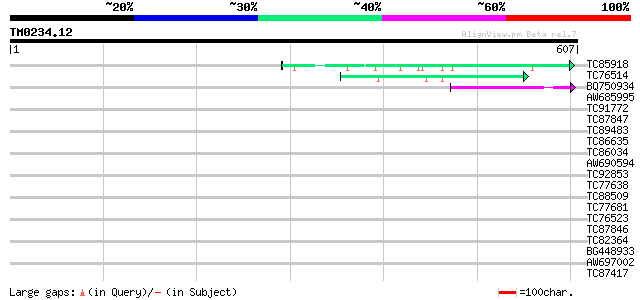
Score E
Sequences producing significant alignments: (bits) Value
TC85918 similar to GP|13562014|gb|AAK30610.1 fibroin 1 {Plectreu... 47 2e-05
TC76514 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - ... 45 1e-04
BQ750934 similar to PIR|T19673|T19 hypothetical protein C33B4.3 ... 42 9e-04
AW685995 similar to PIR|I51618|I516 nucleolar phosphoprotein - A... 40 0.003
TC91772 weakly similar to GP|17380998|gb|AAL36311.1 unknown prot... 40 0.003
TC87847 similar to GP|15929245|gb|AAH15068.1 Similar to adaptor-... 39 0.008
TC89483 similar to GP|18377662|gb|AAL66981.1 unknown protein {Ar... 38 0.010
TC86635 weakly similar to GP|9758600|dbj|BAB09233.1 gene_id:MDF2... 37 0.017
TC86034 similar to GP|6815067|dbj|BAA90354.1 ESTs AU082210(C5365... 37 0.022
AW690594 similar to GP|23498163|emb hypothetical protein {Plasmo... 35 0.064
TC92853 similar to GP|20268748|gb|AAM14077.1 unknown protein {Ar... 35 0.083
TC77638 similar to GP|20161455|dbj|BAB90379. ankyrin-like protei... 35 0.11
TC88509 similar to PIR|T01826|T01826 microfibril-associated prot... 35 0.11
TC77681 similar to GP|22597156|gb|AAN03465.1 nucleolar histone d... 35 0.11
TC76523 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - ... 34 0.14
TC87846 similar to GP|5821151|dbj|BAA83717.1 RNA binding protein... 34 0.19
TC82364 similar to GP|20268740|gb|AAM14073.1 unknown protein {Ar... 33 0.24
BG448933 similar to PIR|T09648|T09 nucleolin homolog nuM1 - alfa... 33 0.32
AW697002 similar to GP|10728183|gb CG11122 gene product {Drosoph... 33 0.32
TC87417 homologue to SP|O22607|MSI4_ARATH WD-40 repeat protein M... 33 0.32
>TC85918 similar to GP|13562014|gb|AAK30610.1 fibroin 1 {Plectreurys tristis},
partial (8%)
Length = 4117
Score = 47.0 bits (110), Expect = 2e-05
Identities = 61/336 (18%), Positives = 134/336 (39%), Gaps = 23/336 (6%)
Frame = -1
Query: 292 IIEKIQVAPKEDT--SDEVRQRDYPIDDFPIWSKKDNPACILEYVRMLRRQGDPITLTEF 349
++ KIQV KED ++ + D PI + DNP + + D IT +E
Sbjct: 3046 MMHKIQV*KKEDEVKAEPAVAAEKTDDSVPIETVSDNP----------KEEKDLITESET 2897
Query: 350 VN-TLPESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLK 408
++ T P + ++ +T S + + +++E + ++ I P E K
Sbjct: 2896 ISVTEPVNDTAISHHDETPSTESDAKETSQQ--IEKELVETNEENQPKIEDIPEAETTEK 2723
Query: 409 RTSAQQQQMSDSESSEDTWKDSSSEETEDEDM-PLAKRR-KIILEEEEDEQDDHE----- 461
+ A + + + + ++S+E+ E ++ PLA K+ E E++ Q+ E
Sbjct: 2722 SSDATETNQFEEKVKQAEISETSAEKEEKPELEPLATEEPKVTKEPEKESQEKSEEAEQP 2543
Query: 462 ----IIQNVIQGIRE---ASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTS 514
I+++ ++ E A+ +EE+ + P ++ P+ + E E+
Sbjct: 2542 KAIAIVESTVEANEEKTAAAISEETKIIEVEPTETVKEEPMVTEVATAEIVKEEPVVTEV 2363
Query: 515 EVQRERRSKRTSDPARAASLTRTSPIRSLSKVITESINSDSALV------VIPEQPMPIS 568
E + + + A + P + KV TE++ + + + E+P+
Sbjct: 2362 EATETLKEELVATEVEATETVKEEPAATEVKV-TETVKDEPVVTEVDPTETVKEEPLATE 2186
Query: 569 TSLPTSTQTEPIQPETQPQTQAEPQAPKSPIQTATT 604
+ + EP+ E +P + + + ++ T
Sbjct: 2185 AEATETVKEEPVATEVEPTETVKEEPVATEVEATET 2078
Score = 45.8 bits (107), Expect = 5e-05
Identities = 67/361 (18%), Positives = 131/361 (35%), Gaps = 49/361 (13%)
Frame = -1
Query: 293 IEKIQVAPKED----TSDEVRQRDYPIDDFPIWSKKDNPACILEYVRMLRRQGDPITLTE 348
IE + PKE+ T E P++D I + P+ + ++ E
Sbjct: 2956 IETVSDNPKEEKDLITESETISVTEPVNDTAISHHDETPSTESD-----AKETSQQIEKE 2792
Query: 349 FVNTLPESPPKL-------ATRKSRRATSSKPSDPKGKGI-LKEEPAKKK--------AA 392
V T E+ PK+ T KS AT + + K K + E A+K+ A
Sbjct: 2791 LVETNEENQPKIEDIPEAETTEKSSDATETNQFEEKVKQAEISETSAEKEEKPELEPLAT 2612
Query: 393 SKTVIIREPSPERPLKRTSAQQQQM-----SDSESSEDTWKDSSSEETE----------- 436
+ + +EP E K A+Q + S E++E+ + SEET+
Sbjct: 2611 EEPKVTKEPEKESQEKSEEAEQPKAIAIVESTVEANEEKTAAAISEETKIIEVEPTETVK 2432
Query: 437 DEDM-------PLAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVK 489
+E M + K ++ E E E E++ ++ + +T+ VK
Sbjct: 2431 EEPMVTEVATAEIVKEEPVVTEVEATETLKEELVATEVEATETVKEEPAATEVKVTETVK 2252
Query: 490 RRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTSDPARAASLTRTSPIRSLSKVITE 549
P+ + ET E+ +E + + + + P+ + + TE
Sbjct: 2251 DE--PVVTEVDPTETVKEEPLATEAEATETVKEEPVATEVEPTETVKEEPVATEVEA-TE 2081
Query: 550 SINSDSALV------VIPEQPMPISTSLPTSTQTEPIQPETQPQTQAEPQAPKSPIQTAT 603
++ + A + E+P+ T + + EP+ E + + ++ + ++
Sbjct: 2080 TVKDEPAATEVEATETVKEEPVATETEATETVKEEPVATEVEATETVKEESVATEVEATE 1901
Query: 604 T 604
T
Sbjct: 1900 T 1898
>TC76514 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - alfalfa,
partial (61%)
Length = 1288
Score = 44.7 bits (104), Expect = 1e-04
Identities = 42/225 (18%), Positives = 90/225 (39%), Gaps = 24/225 (10%)
Frame = +3
Query: 355 ESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAAS---KTVIIREPSPERPLKRTS 411
E K + K ++ K + KEE + ++++ + +++ P+P +
Sbjct: 198 EEEVKAVSAKKQKVEEVAAKQKALKVVKKEESSSEESSESEDEQPVVKAPAPSKKTPAKK 377
Query: 412 AQQQQMSDSESSEDTWKDSSSEETEDEDMPLAK----------RRKIILEEEEDEQDDHE 461
++ +SE++ DSSS + E+ P++K +K+ EEED ++ +
Sbjct: 378 GNVKKAQPETTSEESDSDSSSSDEEEVKKPVSKAVPSKNGSAPAKKVDTSEEEDSEESSD 557
Query: 462 ----------IIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRG-PLQQKETAAEQAS 510
+N ++A+ EE TD + + K + P + A++A
Sbjct: 558 EDKKPAAKAVPSKNGSAPAKKAASDEEDTDESSDEDEEDEKPAAKAVPSKNGSVPAKKAD 737
Query: 511 EPTSEVQRERRSKRTSDPARAASLTRTSPIRSLSKVITESINSDS 555
+S+ E + PA AS ++P + + E + +S
Sbjct: 738 TESSDEDSESSDEEDKKPAAKASKNVSAPTKKAASSSDEESDEES 872
Score = 37.0 bits (84), Expect = 0.022
Identities = 37/149 (24%), Positives = 62/149 (40%), Gaps = 16/149 (10%)
Frame = +3
Query: 357 PPKLATRKSRRA-TSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQ 415
P K + +++A T S D + ++PA K ASK V P K+ ++
Sbjct: 699 PSKNGSVPAKKADTESSDEDSESSDEEDKKPAAK--ASKNV-------SAPTKKAASSSD 851
Query: 416 QMSDSESSED---------------TWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDH 460
+ SD ES ED + KDSS + ED+D + +K + ++ED+ +
Sbjct: 852 EESDEESDEDEDAKPVSKPAAVAKKSKKDSSDSDDEDDDSSSDEDKKPVAAKKEDKMNVD 1031
Query: 461 EIIQNVIQGIREASDAEESTDSDAIPVVK 489
+ + Q E+ D T I V+
Sbjct: 1032KDGSDSDQSEEESEDEPSKTPQKKIKDVE 1118
Score = 35.8 bits (81), Expect = 0.049
Identities = 50/271 (18%), Positives = 109/271 (39%), Gaps = 10/271 (3%)
Frame = +3
Query: 343 PITLTEFVNTLPESPPKLATRKSRRATSSKP--SDPKGKGILKEEPAKKKAASKTVIIRE 400
PI+ + + +P+S K AT+ P S KGK +EE K +A K +
Sbjct: 72 PISHSSLLFIMPKSSKKSATKVDAAPVVVSPVKSGKKGKRQAEEE-VKAVSAKKQKVEEV 248
Query: 401 PSPERPLKRTSAQQQQMSDSESSEDTWK----DSSSEETEDEDMPLAKRRKIILEEEEDE 456
+ ++ LK ++ +S SED + S++T + + K + EE D
Sbjct: 249 AAKQKALKVVKKEESSSEESSESEDEQPVVKAPAPSKKTPAKKGNVKKAQPETTSEESDS 428
Query: 457 QDDHEIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEV 516
+ V + + +A ++ + P ++ +T+ E+ SE +S+
Sbjct: 429 DSSSSDEEEVKKPVSKAVPSKNGS----------------APAKKVDTSEEEDSEESSDE 560
Query: 517 QRERRSKRTSDPARAASLTRTSPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQ 576
++ +K P++ S +P + K ++ ++D + E P + ++P+
Sbjct: 561 DKKPAAKAV--PSKNGS----APAK---KAASDEEDTDESSDEDEEDEKPAAKAVPSKNG 713
Query: 577 TEPIQ----PETQPQTQAEPQAPKSPIQTAT 603
+ P + + +++ + K P A+
Sbjct: 714 SVPAKKADTESSDEDSESSDEEDKKPAAKAS 806
Score = 32.7 bits (73), Expect = 0.41
Identities = 25/146 (17%), Positives = 61/146 (41%)
Frame = +3
Query: 462 IIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERR 521
++ V G + AEE + + K ++ + + E +SE +SE + E+
Sbjct: 153 VVSPVKSGKKGKRQAEEEVKAVSAKKQKVEEVAAKQKALKVVKKEESSSEESSESEDEQP 332
Query: 522 SKRTSDPARAASLTRTSPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQTEPIQ 581
+ P++ + + ++ + +E +SDS+ E P+S ++P+ + P +
Sbjct: 333 VVKAPAPSKKTPAKKGNVKKAQPETTSEESDSDSSSSDEEEVKKPVSKAVPSKNGSAPAK 512
Query: 582 PETQPQTQAEPQAPKSPIQTATTAIP 607
+ + ++ + A A+P
Sbjct: 513 KVDTSEEEDSEESSDEDKKPAAKAVP 590
>BQ750934 similar to PIR|T19673|T19 hypothetical protein C33B4.3 -
Caenorhabditis elegans, partial (1%)
Length = 627
Score = 41.6 bits (96), Expect = 9e-04
Identities = 35/134 (26%), Positives = 54/134 (40%), Gaps = 1/134 (0%)
Frame = +2
Query: 473 ASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQ-ASEPTSEVQRERRSKRTSDPARA 531
AS + E+T + P + ET ++ AS S + S ++ P+
Sbjct: 179 ASSSVETTAAAVFPTTSAVANGTLTAVTASETVTDEVASSSLSTLSATNSSVTSAAPSFT 358
Query: 532 ASLTRTSPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQTEPIQPETQPQTQAE 591
A TSP+ S S E++++ + V P STSLP P PQ
Sbjct: 359 AEAETTSPVSSASSTSLETLDATTTAVPSSSIPESSSTSLP---------PTLVPQNTTS 511
Query: 592 PQAPKSPIQTATTA 605
P + S I T+TT+
Sbjct: 512 PASSPSSITTSTTS 553
>AW685995 similar to PIR|I51618|I516 nucleolar phosphoprotein - African
clawed frog, partial (5%)
Length = 516
Score = 40.0 bits (92), Expect = 0.003
Identities = 29/121 (23%), Positives = 52/121 (42%), Gaps = 13/121 (10%)
Frame = +3
Query: 354 PESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSP---ERPLKRT 410
P+ K+ K ++A ++ K + +E+PA + + E S E P K+
Sbjct: 63 PKKVEKVEKTKGKKAAKAEEEKKKQQKKKEEKPAPMEVEEDSSSSEESSSSEDEAPAKKA 242
Query: 411 SAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAK----------RRKIILEEEEDEQDDH 460
+ ++ + ESS + SS E+E+E+ P K EEE DE++ +
Sbjct: 243 APAKKAAAKKESSSEEESSSSESESEEEEKPAKKAAAKKESSSEEESSSSEEESDEEEKN 422
Query: 461 E 461
E
Sbjct: 423 E 425
Score = 37.4 bits (85), Expect = 0.017
Identities = 31/107 (28%), Positives = 44/107 (40%), Gaps = 9/107 (8%)
Frame = +3
Query: 355 ESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSP---------ER 405
E P + + ++ S K PAKK AA K E S E+
Sbjct: 153 EKPAPMEVEEDSSSSEESSSSEDEAPAKKAAPAKKAAAKKESSSEEESSSSESESEEEEK 332
Query: 406 PLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEE 452
P K+ +A+++ S+ ESS SSEE DE+ RR+ EE
Sbjct: 333 PAKKAAAKKESSSEEESS-------SSEEESDEEEKNEARRR*EQEE 452
>TC91772 weakly similar to GP|17380998|gb|AAL36311.1 unknown protein
{Arabidopsis thaliana}, partial (17%)
Length = 1237
Score = 39.7 bits (91), Expect = 0.003
Identities = 48/213 (22%), Positives = 91/213 (42%), Gaps = 15/213 (7%)
Frame = +2
Query: 358 PKL--ATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQ 415
PKL +T K + T S P KG + + K + +EP PER ++R +
Sbjct: 20 PKLERSTIKKQDGTDS-PVLSKGNNLSGNDERDDMDTEK-IDSKEPKPERSVRRKGKKAS 193
Query: 416 QMSDSESSE--DTWKDSSSEETED-----EDMPLAKRRKIILEEEEDEQDDHEI---IQN 465
++ S+ + + +E+T D +++P++ ++E ++D EI I +
Sbjct: 194 SSKSTKPSKKSNVVSEKEAEKTADSKSSKKEVPISLNEDSVVEATGTSENDKEIKAKISS 373
Query: 466 VIQGIREASDAEESTDSDAIPVVKRRKLPLR---GPLQQKETAAEQASEPTSEVQRERRS 522
G E+ A + S++ R K +R KE AAE S+ SE + +
Sbjct: 374 PKAGGLESDAAGSPSPSESNHDENRSKKRVRTKKNDSSAKEVAAEDISKKVSEGTSDSKV 553
Query: 523 KRTSDPARAASLTRTSPIRSLSKVITESINSDS 555
K A+ + R+S ++++ + + S S
Sbjct: 554 KPARPSAKKGPI-RSSDVKTVVHAVMADVGSSS 649
>TC87847 similar to GP|15929245|gb|AAH15068.1 Similar to adaptor-related
protein complex AP-3 beta 1 subunit {Mus musculus},
partial (2%)
Length = 931
Score = 38.5 bits (88), Expect = 0.008
Identities = 48/203 (23%), Positives = 75/203 (36%), Gaps = 4/203 (1%)
Frame = +1
Query: 356 SPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQ 415
S PK S S P K ++ + P AS + + PS ++P + +Q
Sbjct: 13 SLPKTLKP*SHPKFSMAPKPKKRLALVDDPPT----ASSSDEEQPPSTQKPADKVVNEQD 180
Query: 416 QMSDSESSEDTWKD-SSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREA- 473
+ S SE E +D SSSE+ E+ED + + I N E+
Sbjct: 181 EDSSSEEEEGDEEDGSSSEDEEEEDNQTQQTPSKDSKPASKNPPPSTPISNPKPAESESG 360
Query: 474 --SDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTSDPARA 531
S +E T+SD+ P K P P + + ++ ++ Q ++ A
Sbjct: 361 SESGSESGTESDSEPEHKSEPTPPPNPKVKPLASKPMKTQTQAQAQSTPVPIKSGTKRVA 540
Query: 532 ASLTRTSPIRSLSKVITESINSD 554
S T RS K T SD
Sbjct: 541 ESSTGNDSKRSKKKTTTAGGGSD 609
>TC89483 similar to GP|18377662|gb|AAL66981.1 unknown protein {Arabidopsis
thaliana}, partial (35%)
Length = 1711
Score = 38.1 bits (87), Expect = 0.010
Identities = 41/164 (25%), Positives = 73/164 (44%), Gaps = 10/164 (6%)
Frame = +1
Query: 338 RRQGDPITLTEFVNTLPESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVI 397
R+ GD +E + PE+P + RKSR + + + +L+ ++ A + I
Sbjct: 61 RQTGDKKPTSE--HDRPETPDRRHDRKSRERDRERELKREKERVLERY---EREAERDRI 225
Query: 398 IREPSPERPLKRTSAQ-QQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDE 456
+E +R ++ Q + Q+ + E E + E E E KRRK IL +EED+
Sbjct: 226 RKEREQKRRIEEVERQFELQLKEWEYREREKEKERQYEKEKEKDRERKRRKEILYDEEDD 405
Query: 457 QDD-------HEIIQNVIQGIREASD--AEESTDSDAIPVVKRR 491
D + I + + +RE D A++ + + I K+R
Sbjct: 406 DGDSRKRWRXNAIEEKRNKRLREKEDDLADKQKEEEEIAEAKKR 537
>TC86635 weakly similar to GP|9758600|dbj|BAB09233.1
gene_id:MDF20.10~pir||T08929~similar to unknown protein
{Arabidopsis thaliana}, partial (32%)
Length = 1963
Score = 37.4 bits (85), Expect = 0.017
Identities = 30/111 (27%), Positives = 54/111 (48%), Gaps = 1/111 (0%)
Frame = +2
Query: 413 QQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIRE 472
++ ++ + E +D K S EETE+++ K + EEE+E D+ I ++ + E
Sbjct: 26 EKTEVEEKEEGDDNEK--SEEETEEKEEGDEKEKTEAETEEEEEADEDTIDKSKEEDKAE 199
Query: 473 ASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSE-VQRERRS 522
S E+ + A V K+ ++ +KE + PTS+ RER+S
Sbjct: 200 GSKGEKGSKKRARGKVNEEKVKVK----KKELKLPEPKTPTSDRPVRERKS 340
Score = 28.5 bits (62), Expect = 7.8
Identities = 49/228 (21%), Positives = 86/228 (37%), Gaps = 43/228 (18%)
Frame = +2
Query: 351 NTLPESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASK-------------TVI 397
N +PE +KS R S ++ P+K +A K T
Sbjct: 953 NDIPEKSEDETPQKSEREDKSDSGSESEDVKKRKRPSKTSSAKKESAGRSKTEKSTVTNK 1132
Query: 398 IREPSPERPLKRTSAQQQQMSD--------------SE-------SSEDTWKDSSSEETE 436
R P P+R K++S + + D SE S+ K SS + +
Sbjct: 1133 SRSPPPKRAPKKSSNTRPKSDDDINESPKVFSRKKKSEQGGKQKISTPTPTKSSSKSKEK 1312
Query: 437 DEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKRR----K 492
E + K +K + H+ I +++ + D +T +D + ++ ++
Sbjct: 1313 TEKVTKGKGKKKETSSSPTDDQLHDAICEILKEV----DFNTATFTDILKLLAKQFDVDL 1480
Query: 493 LPLRGPLQ---QKETA--AEQASEPTSEVQRERRSKRTSDPARAASLT 535
P + ++ QKE A AE+A E SE E +++ D + SLT
Sbjct: 1481 TPKKSAIKLMIQKELARLAEEADE--SEEDEEEDNEKDEDRS*TLSLT 1618
>TC86034 similar to GP|6815067|dbj|BAA90354.1 ESTs AU082210(C53655)
AU056546(S20671) AU056545(S20671) correspond to a
region of the predicted, partial (42%)
Length = 2412
Score = 37.0 bits (84), Expect = 0.022
Identities = 33/143 (23%), Positives = 61/143 (42%)
Frame = +3
Query: 343 PITLTEFVNTLPESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPS 402
P++L+ N P A K R+ SS P EP K+ EP
Sbjct: 42 PLSLSVSPNPNPNFDSITAMAKKRKLRSSDP-----------EPTKQA---------EPE 161
Query: 403 PERPLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEI 462
P++ A+++ + E + +D E E E+ P ++ EEE+D+Q + E
Sbjct: 162 PQQ---EEVAEEEPTTQEEENTPIEEDQPQNEGEGEEEP---EEEVAEEEEDDQQGEDET 323
Query: 463 IQNVIQGIREASDAEESTDSDAI 485
++N ++ D E++++ A+
Sbjct: 324 LEN-----QQQQDGGEASNNAAV 377
>AW690594 similar to GP|23498163|emb hypothetical protein {Plasmodium
falciparum 3D7}, partial (10%)
Length = 633
Score = 35.4 bits (80), Expect = 0.064
Identities = 23/80 (28%), Positives = 34/80 (41%)
Frame = +3
Query: 404 ERPLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEII 463
E KR ++ + E + +D EE EDE + + EEEEDE DD E
Sbjct: 18 ESSWKRRKVEEHNEEEDEGEVEEEEDEHDEEEEDEHDEEEEEEEE--EEEEDEHDDEEEE 191
Query: 464 QNVIQGIREASDAEESTDSD 483
+ + E D EE + +
Sbjct: 192 EEEEEEEEEEDDDEEGEEDE 251
>TC92853 similar to GP|20268748|gb|AAM14077.1 unknown protein {Arabidopsis
thaliana}, partial (21%)
Length = 761
Score = 35.0 bits (79), Expect = 0.083
Identities = 28/108 (25%), Positives = 41/108 (37%), Gaps = 13/108 (12%)
Frame = +3
Query: 365 SRRATSSKPSDPKGKGILKEEPAKKKAAS-------------KTVIIREPSPERPLKRTS 411
SR SS PS+ + K + +++ K+ T+ PSP +P
Sbjct: 150 SREGLSSNPSEKEIKEVNRKKARAKELEGIDMSNIVSSTRRRATISFVAPSPPKPKIPVE 329
Query: 412 AQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDD 459
+ S++ D + EE EDED EEEED DD
Sbjct: 330 TSGKDTKGSDNDNDDEGKENEEEEEDED-----------EEEEDTSDD 440
>TC77638 similar to GP|20161455|dbj|BAB90379. ankyrin-like protein {Oryza
sativa (japonica cultivar-group)}, partial (62%)
Length = 2345
Score = 34.7 bits (78), Expect = 0.11
Identities = 34/148 (22%), Positives = 66/148 (43%), Gaps = 10/148 (6%)
Frame = +1
Query: 413 QQQQMSDSESSEDTWKDSSSEETEDE----------DMPLAKRRKIILEEEEDEQDDHEI 462
+ ++ S +SSEDT + ++TEDE + K + E++DE +++
Sbjct: 604 KSEENSTEKSSEDTKTEDEGKKTEDEGSNTENNKDGEEASTKESESDESEKKDESEENN- 780
Query: 463 IQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRS 522
+ + +++SD+ E+TDS+ V++ Q + + E ASE
Sbjct: 781 KSDSDESEKKSSDSNETTDSNVEEKVEQ---------SQNKESDENASE----------- 900
Query: 523 KRTSDPARAASLTRTSPIRSLSKVITES 550
K T D A+ S P + S+++ E+
Sbjct: 901 KNTDDNAKDQSSNEVFPSGAQSELLNET 984
>TC88509 similar to PIR|T01826|T01826 microfibril-associated protein homolog
T15F16.8 - Arabidopsis thaliana, partial (83%)
Length = 1306
Score = 34.7 bits (78), Expect = 0.11
Identities = 36/168 (21%), Positives = 70/168 (41%), Gaps = 11/168 (6%)
Frame = +1
Query: 398 IREPSPERPLKRTSAQQQQMSDSESSEDTW-------KDSSSEETEDEDMPLAKRRKIIL 450
IR+ ++ + +Q+ + E ED K+ + ++E +P + +
Sbjct: 346 IRQAEIVSTIEEEAKRQEGLDLEEQDEDAMAERRRRIKEKLRQRDQEEALPQEEEEEEEE 525
Query: 451 EEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAE--- 507
EEEE+E+ D+E +D++E + + +VK P+ P +++T AE
Sbjct: 526 EEEEEEESDYE------------TDSDE--EYTGVAMVK----PVFVPKSERDTIAERER 651
Query: 508 -QASEPTSEVQRERRSKRTSDPARAASLTRTSPIRSLSKVITESINSD 554
+A E E R+RR + + R + + + K I N D
Sbjct: 652 LEAEEEALEEARKRRMEERRNETRQIVVEEIRKDQEIQKNIELEANID 795
>TC77681 similar to GP|22597156|gb|AAN03465.1 nucleolar histone deacetylase
HD2-P39 {Glycine max}, partial (47%)
Length = 1233
Score = 34.7 bits (78), Expect = 0.11
Identities = 21/73 (28%), Positives = 31/73 (41%)
Frame = +2
Query: 367 RATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQMSDSESSEDT 426
+A K S+PK PAK A +K V + +P E + + S + E+
Sbjct: 461 KAEELKVSEPKKADAKNAAPAKNAATAKHVKVVDPKDEDSDDDDESDDEIGSSDDEMENA 640
Query: 427 WKDSSSEETEDED 439
DS E+ DED
Sbjct: 641 DSDSEDEDDSDED 679
>TC76523 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - alfalfa,
partial (69%)
Length = 1615
Score = 34.3 bits (77), Expect = 0.14
Identities = 24/100 (24%), Positives = 46/100 (46%), Gaps = 22/100 (22%)
Frame = +3
Query: 406 PLKRTSAQQQQMSDSESSED---------------TWKDSSSEETEDED-------MPLA 443
P ++ ++ + SD ES ED + KDSS + ED+D P+A
Sbjct: 144 PQRKPASSSDEESDEESDEDEDAKPVSKPAAVAKKSKKDSSDSDDEDDDSSSDEDKKPVA 323
Query: 444 KRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSD 483
++++ E E + D E NV + ++ ++EE ++ +
Sbjct: 324 AKKEV--SESESDSSDDEDKMNVDKDGSDSDESEEESEDE 437
>TC87846 similar to GP|5821151|dbj|BAA83717.1 RNA binding protein {Homo
sapiens}, partial (2%)
Length = 550
Score = 33.9 bits (76), Expect = 0.19
Identities = 32/127 (25%), Positives = 53/127 (41%), Gaps = 4/127 (3%)
Frame = +1
Query: 376 PKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQMSDSESSEDTWKD-SSSEE 434
PK + L ++P ++ + + PS ++P + +Q + S SE E +D SSSE+
Sbjct: 52 PKKRLALVDDPPTASSSDEE---QPPSTQKPADKVVNEQDEDSSSEEEEGDEEDGSSSED 222
Query: 435 TEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREA---SDAEESTDSDAIPVVKRR 491
E+ED + + I N E+ S +E T+SD+ P K
Sbjct: 223 EEEEDNQTQQTPSKDSKPASKNPPPSTPISNPKPAESESGSESGSESGTESDSEPEHKSE 402
Query: 492 KLPLRGP 498
P P
Sbjct: 403 PTPPPNP 423
>TC82364 similar to GP|20268740|gb|AAM14073.1 unknown protein {Arabidopsis
thaliana}, partial (39%)
Length = 1174
Score = 33.5 bits (75), Expect = 0.24
Identities = 22/60 (36%), Positives = 31/60 (51%), Gaps = 2/60 (3%)
Frame = +3
Query: 550 SINSDSALVVIPEQPMPIS-TSLPTSTQTEPI-QPETQPQTQAEPQAPKSPIQTATTAIP 607
S +S S + +QP+PI S P Q P +PE+QP + +P P P Q + T IP
Sbjct: 102 SFSSSSESNPLFQQPIPIQPVSYPVKPQDPPPPEPESQPSSDTQP--PPPPPQASNTPIP 275
>BG448933 similar to PIR|T09648|T09 nucleolin homolog nuM1 - alfalfa, partial
(29%)
Length = 663
Score = 33.1 bits (74), Expect = 0.32
Identities = 27/144 (18%), Positives = 62/144 (42%), Gaps = 14/144 (9%)
Frame = +2
Query: 355 ESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAAS---KTVIIREPSPERPLKRTS 411
E K + K ++ K + KEE + ++++ + +++ P+P +
Sbjct: 176 EEEVKAVSAKKQKVEEVAAKQKALKVVKKEESSSEESSESEDEQPVVKAPAPSKKTPAKK 355
Query: 412 AQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKR---------RKIILEEEEDEQDDHEI 462
++ +SE++ DSSS + E+ P++K K + +ED++ +
Sbjct: 356 GNVKKAQPETTSEESDSDSSSSDEEEVKKPVSKAVPSKNGSAPAKKVDTSDEDKKPAAKA 535
Query: 463 I--QNVIQGIREASDAEESTDSDA 484
+ +N ++A+ EE TD +
Sbjct: 536 VPSKNGSAPAKKAASDEEDTDESS 607
Score = 32.0 bits (71), Expect = 0.71
Identities = 25/137 (18%), Positives = 56/137 (40%)
Frame = +2
Query: 462 IIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERR 521
++ V G + AEE + + K ++ + + E +SE +SE + E+
Sbjct: 131 VVSPVKSGKKGKRQAEEEVKAVSAKKQKVEEVAAKQKALKVVKKEESSSEESSESEDEQP 310
Query: 522 SKRTSDPARAASLTRTSPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQTEPIQ 581
+ P++ + + ++ + +E +SDS+ E P+S ++P+ + P +
Sbjct: 311 VVKAPAPSKKTPAKKGNVKKAQPETTSEESDSDSSSSDEEEVKKPVSKAVPSKNGSAPAK 490
Query: 582 PETQPQTQAEPQAPKSP 598
+P A P
Sbjct: 491 KVDTSDEDKKPAAKAVP 541
>AW697002 similar to GP|10728183|gb CG11122 gene product {Drosophila
melanogaster}, partial (0%)
Length = 857
Score = 33.1 bits (74), Expect = 0.32
Identities = 20/74 (27%), Positives = 32/74 (43%)
Frame = +3
Query: 355 ESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQ 414
E+PP K + KP + K +EEP++ K+ RE E+ ++ QQ
Sbjct: 57 ETPPSTGAVKKKETRGPKPKPKEDK---REEPSQVKSP------RESKKEKQQQQLHQQQ 209
Query: 415 QQMSDSESSEDTWK 428
QQ + + WK
Sbjct: 210 QQQASVDEKYSEWK 251
>TC87417 homologue to SP|O22607|MSI4_ARATH WD-40 repeat protein MSI4.
[Mouse-ear cress] {Arabidopsis thaliana}, partial (36%)
Length = 743
Score = 33.1 bits (74), Expect = 0.32
Identities = 20/74 (27%), Positives = 32/74 (43%)
Frame = +2
Query: 355 ESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQ 414
E+PP K + KP + K +EEP++ K+ RE E+ ++ QQ
Sbjct: 65 ETPPSTGAVKKKETRGRKPKPKEDK---REEPSQVKSP------RESKKEKQQQQLHQQQ 217
Query: 415 QQMSDSESSEDTWK 428
QQ + + WK
Sbjct: 218 QQQASVDEKYSQWK 259
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.312 0.129 0.363
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,715,070
Number of Sequences: 36976
Number of extensions: 208271
Number of successful extensions: 1354
Number of sequences better than 10.0: 85
Number of HSP's better than 10.0 without gapping: 1275
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1329
length of query: 607
length of database: 9,014,727
effective HSP length: 102
effective length of query: 505
effective length of database: 5,243,175
effective search space: 2647803375
effective search space used: 2647803375
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0234.12


