

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0230.1
(420 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
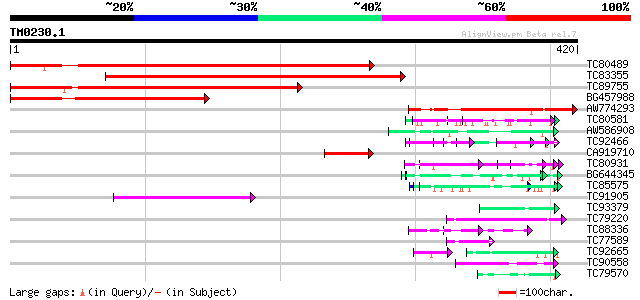
Score E
Sequences producing significant alignments: (bits) Value
TC80489 similar to GP|9294341|dbj|BAB02238.1 contains similarity... 332 2e-91
TC83355 similar to GP|6016726|gb|AAF01552.1| unknown protein {Ar... 296 1e-80
TC89755 similar to GP|10177798|dbj|BAB11289. emb|CAB87778.1~gene... 251 3e-67
BG457988 similar to PIR|C84635|C84 hypothetical protein At2g2432... 175 3e-44
AW774293 similar to GP|13881624|gb PPE family protein {Mycobacte... 159 2e-39
TC80581 similar to GP|19683021|gb|AAL92644.1 hypothetical protei... 77 1e-14
AW586908 weakly similar to GP|2104814|emb| pleiotropic effects o... 66 3e-11
TC92466 similar to GP|11141605|gb|AAG32022.1 LEUNIG {Arabidopsis... 63 2e-10
CA919710 similar to GP|10177798|dbj emb|CAB87778.1~gene_id:MXA21... 58 8e-09
TC80931 similar to GP|14190783|gb|AAF65166.2 putative phloem tra... 55 1e-08
BG644345 weakly similar to GP|19697333|gb putative protein poten... 57 2e-08
TC85575 similar to PIR|D86410|D86410 protein F3M18.16 [imported]... 49 4e-06
TC91905 similar to GP|19347881|gb|AAL85997.1 unknown protein {Ar... 47 1e-05
TC93379 similar to GP|19569994|gb|AAL92295.1 hypothetical protei... 45 4e-05
TC79220 similar to GP|2149570|gb|AAB58577.1| MAP kinase kinase p... 44 2e-04
TC88336 similar to PIR|T51477|T51477 glutamine-rich protein - Ar... 43 2e-04
TC77589 similar to GP|1087015|gb|AAB35283.1| arabinogalactan-pro... 43 2e-04
TC92665 homologue to GP|15291583|gb|AAK93060.1 GH28569p {Drosoph... 38 2e-04
TC90558 similar to GP|9758584|dbj|BAB09197.1 gb|AAC80623.1~gene_... 43 3e-04
TC79570 similar to PIR|F86427|F86427 auxin response factor 6 (AR... 43 3e-04
>TC80489 similar to GP|9294341|dbj|BAB02238.1 contains similarity to unknown
protein~gb|AAF01552.1~gene_id:T6J22.18 {Arabidopsis
thaliana}, partial (68%)
Length = 1399
Score = 332 bits (851), Expect = 2e-91
Identities = 166/273 (60%), Positives = 202/273 (73%), Gaps = 3/273 (1%)
Frame = +3
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQL---KKRDDGKLTVVSAASPSVPIPHADDSSLDVLLV 57
MG + S +AAKFAFFPP+PP+Y L + K+T VS ++DVL +
Sbjct: 96 MGAVTSSMAAKFAFFPPNPPSYGLGVDESTGKNKITGVSTRE-----------NVDVLKL 242
Query: 58 DTKHGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGA 117
TK GN IVA Y+KN A +T+LYSHGNAADLGQ+Y+LF +L ++LRVNL+ YDYSGYG
Sbjct: 243 CTKRGNNIVALYIKNSSASITILYSHGNAADLGQMYELFSELSIHLRVNLLCYDYSGYGQ 422
Query: 118 STGKPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLH 177
S+GKPSE +TYADIEA Y+CL YG +EDIILYGQSVGSGPT LAA+LP LR V+LH
Sbjct: 423 SSGKPSEQNTYADIEAAYKCLVEMYGSKEEDIILYGQSVGSGPTTDLAARLPNLRAVILH 602
Query: 178 SGILSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARE 237
S ILSGLRV+ VK ++ FDIYKNI+KI V CPVLVIHGT DDVV+ HG LW+ +E
Sbjct: 603 SPILSGLRVMYPVKRTYWFDIYKNIDKIPMVNCPVLVIHGTADDVVDCSHGKQLWEHCKE 782
Query: 238 SYDPLWIKGGGHCNLELYPDYIRHLCKFIQEME 270
Y+PLW+KGG HC+LELYP YI+HL KFI +E
Sbjct: 783 KYEPLWVKGGNHCDLELYPQYIKHLKKFIAAIE 881
>TC83355 similar to GP|6016726|gb|AAF01552.1| unknown protein {Arabidopsis
thaliana}, partial (68%)
Length = 1032
Score = 296 bits (757), Expect = 1e-80
Identities = 139/222 (62%), Positives = 176/222 (78%)
Frame = +1
Query: 72 NPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGASTGKPSESSTYADI 131
+P A TLL+SHGNAADLGQ+Y+LF++L ++LRVNLMGYDYSGYG S+GKPSE +TY+DI
Sbjct: 1 SPMATSTLLHSHGNAADLGQMYELFIELSIHLRVNLMGYDYSGYGQSSGKPSEQNTYSDI 180
Query: 132 EAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHSGILSGLRVLCHVK 191
EA+Y+CLE +G QEDIILYGQSVGSGPTL LAA+LP+LR VVLHS ILSGLRV+ VK
Sbjct: 181 EAVYKCLEESFGAKQEDIILYGQSVGSGPTLDLAARLPQLRAVVLHSPILSGLRVMYPVK 360
Query: 192 FSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARESYDPLWIKGGGHCN 251
S+ FDIYKNI+KI V CPVL++HGT D+VV+ HG LW++ +E Y+PLW+KGG HC+
Sbjct: 361 RSYWFDIYKNIDKIPLVNCPVLIVHGTSDEVVDCSHGKQLWELCKEKYEPLWLKGGNHCD 540
Query: 252 LELYPDYIRHLCKFIQEMESITTEKRLKKLRQSQGPQSNSSS 293
LEL+P+YIRHL KFI +E +++ + Q Q S+
Sbjct: 541 LELFPEYIRHLKKFITTVEKSPSQRYSFRRSTDQFEQPRKST 666
>TC89755 similar to GP|10177798|dbj|BAB11289.
emb|CAB87778.1~gene_id:MXA21.9~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (59%)
Length = 864
Score = 251 bits (642), Expect = 3e-67
Identities = 130/225 (57%), Positives = 162/225 (71%), Gaps = 8/225 (3%)
Frame = +3
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDDGKLTVVSAAS--------PSVPIPHADDSSL 52
MG + S +AAKFAFFPP PP+Y L + T + + P VP+ ++
Sbjct: 201 MGGVTSSIAAKFAFFPPQPPSYTLTVSESTATTAAANGTSPATKLLIPEVPMKE----NV 368
Query: 53 DVLLVDTKHGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDY 112
DV+ V T+ GN+I A Y+K T+LYSHGNAADLGQ+++LFV+L LR+N+MGYDY
Sbjct: 369 DVVKVKTRRGNEIAAVYVKYHRPACTMLYSHGNAADLGQMFELFVELSNRLRINVMGYDY 548
Query: 113 SGYGASTGKPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLR 172
SGYG STGKP+E +TYADIEA Y+CL+ +YGV E +ILYGQSVGSGPTL LA+++ LR
Sbjct: 549 SGYGQSTGKPTEYNTYADIEAAYKCLKEQYGVKDEQLILYGQSVGSGPTLDLASRISELR 728
Query: 173 GVVLHSGILSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHG 217
GVVLHS ILSGLRVL VK ++ FDIYKNI+KI VKCP LVIHG
Sbjct: 729 GVVLHSPILSGLRVLYPVKRTYWFDIYKNIDKIGMVKCPGLVIHG 863
>BG457988 similar to PIR|C84635|C84 hypothetical protein At2g24320 [imported]
- Arabidopsis thaliana, partial (37%)
Length = 601
Score = 175 bits (443), Expect = 3e-44
Identities = 86/148 (58%), Positives = 108/148 (72%)
Frame = +3
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDDGKLTVVSAASPSVPIPHADDSSLDVLLVDTK 60
MG + +AAKFAFFPP PPTY++ K +D + +S + D S+DV +VDTK
Sbjct: 183 MGNVTGTVAAKFAFFPPEPPTYEVYKENDDGVLKISGTT---------DKSVDVHIVDTK 335
Query: 61 HGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGASTG 120
GNKIVA K+P+AR T+LYSHGNAADLGQ+ DLF++L+ +LRVN+M YDYSGYG STG
Sbjct: 336 GGNKIVATLWKHPFARFTILYSHGNAADLGQMRDLFIELRAHLRVNIMSYDYSGYGRSTG 515
Query: 121 KPSESSTYADIEAIYECLESEYGVGQED 148
KPSE +TY DIEA+Y L +EYG QED
Sbjct: 516 KPSEFNTYYDIEAVYNLLRNEYGFKQED 599
>AW774293 similar to GP|13881624|gb PPE family protein {Mycobacterium
tuberculosis CDC1551}, partial (1%)
Length = 597
Score = 159 bits (401), Expect = 2e-39
Identities = 73/129 (56%), Positives = 85/129 (65%), Gaps = 4/129 (3%)
Frame = -1
Query: 296 SCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLPSC 355
+C+C MKCC CS C C P+C +C CSCIN S S+ C CC PSC
Sbjct: 597 TCTCPSMKCC------CS-CXCQLPKCSSC---------CSCINFSLSINCPDCCWKPSC 466
Query: 356 IKCCHLPKFSTCFGSSCCTKCSRPSCCF-KC---CCNLRCARPSCCCMSCFCWRCCVGKH 411
IKCC LPKF+ CFGS+CCTKCS P CC KC CC +C+RPS CCMSCFCW+C +GKH
Sbjct: 465 IKCCRLPKFTDCFGSTCCTKCSMPGCCCPKCSLPCCYPKCSRPS-CCMSCFCWQCFMGKH 289
Query: 412 SGRDEKQSG 420
SGR+ KQ G
Sbjct: 288 SGRNGKQRG 262
Score = 37.0 bits (84), Expect = 0.014
Identities = 19/51 (37%), Positives = 23/51 (44%), Gaps = 2/51 (3%)
Frame = -1
Query: 292 SSCFSCSCCRMKCCSNNCTGCSNCS--CINPECCNCSCINPECFNCSCINC 340
+ CF +CC KC C C CS C P+C SC C +C C C
Sbjct: 438 TDCFGSTCC-TKCSMPGCC-CPKCSLPCCYPKCSRPSC----CMSCFCWQC 304
>TC80581 similar to GP|19683021|gb|AAL92644.1 hypothetical protein
{Dictyostelium discoideum}, partial (5%)
Length = 850
Score = 77.4 bits (189), Expect = 1e-14
Identities = 52/149 (34%), Positives = 59/149 (38%), Gaps = 35/149 (23%)
Frame = -3
Query: 294 CFSCSCCRMK--CCSNNCTGCSNC---SCINPECCNCSCIN------PECFNCS------ 336
C C CCR CCS +C CS C C CC CIN E F CS
Sbjct: 731 CCCCCCCRTSCCCCSRSCCCCSCC*SRGCPMDFCC---CINWGNWFRVEEFCCS*ADEAS 561
Query: 337 --------CINCSCSLKCTQCCC----LPSCIKCCHLPKFSTCF---GSSCCTKCSRPSC 381
C NCSC + CCC P CC + K + C+ CC+ SC
Sbjct: 560 CCCNL*YCCCNCSCCCRSCCCCC*SRGCPMDFGCC-INKGNCCWLRVKEFCCS*ADEASC 384
Query: 382 CFK---CCCNLRCARPSCCCMSCFCWRCC 407
C CCC+ SCCC SC C CC
Sbjct: 383 CCSL*YCCCSC-----SCCCRSC*CCSCC 312
Score = 57.0 bits (136), Expect = 1e-08
Identities = 44/122 (36%), Positives = 51/122 (41%), Gaps = 16/122 (13%)
Frame = -3
Query: 299 CC----RMKCCSNNCTGCSNCSCINPECCNCSCINPEC---FNCSCINCS--CSLKCTQC 349
CC CC N C NCSC CC C C + C F C CIN C L+ +
Sbjct: 587 CCS*ADEASCCCNL*YCCCNCSCCCRSCC-CCC*SRGCPMDFGC-CINKGNCCWLRVKEF 414
Query: 350 CCL----PSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLRCARPSCCCM---SCF 402
CC SC CC L + C S CC C SCC C C + CC+ +C
Sbjct: 413 CCS*ADEASC--CCSL*-YCCCSCSCCCRSC*CCSCC*SCGCPMGFG----CCIN*GNC- 258
Query: 403 CW 404
CW
Sbjct: 257 CW 252
Score = 56.2 bits (134), Expect = 2e-08
Identities = 33/101 (32%), Positives = 37/101 (35%), Gaps = 18/101 (17%)
Frame = -3
Query: 325 CSCINPECFNCSCINCSCSLKCTQCCCLPSCIKC-------CHLPKFSTCFG-------- 369
C C C C C C + + CCC SC C C + F C
Sbjct: 770 CICCCRS*VRCCCCCCCCCCRTSCCCCSRSCCCCSCC*SRGCPM-DFCCCINWGNWFRVE 594
Query: 370 SSCCTKCSRPSCCFK---CCCNLRCARPSCCCMSCFCWRCC 407
CC+ SCC CCCN SCCC SC C CC
Sbjct: 593 EFCCS*ADEASCCCNL*YCCCNC-----SCCCRSCCC--CC 492
Score = 50.8 bits (120), Expect = 1e-06
Identities = 26/70 (37%), Positives = 33/70 (47%), Gaps = 1/70 (1%)
Frame = -3
Query: 336 SCINCSCS-LKCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLRCARP 394
+CI C S ++C CCC CC C +SCC CSR CC CC + C
Sbjct: 773 ACICCCRS*VRCCCCCC------CC-------CCRTSCCC-CSRSCCCCSCC*SRGCPMD 636
Query: 395 SCCCMSCFCW 404
CCC++ W
Sbjct: 635 FCCCINWGNW 606
Score = 35.4 bits (80), Expect = 0.041
Identities = 37/137 (27%), Positives = 40/137 (29%), Gaps = 27/137 (19%)
Frame = -3
Query: 294 CFSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLP 353
C SCSCC C C CSC C +C C P F C CIN
Sbjct: 362 CCSCSCC--------CRSC*CCSC----C*SCGC--PMGFGC-CIN*G------------ 264
Query: 354 SCIKCCHLPKFSTCFG----SSCC--------TKCSRPSCCFKCCCNLRCARPSC----- 396
CC C +SCC K + P C N C C
Sbjct: 263 ---NCCWFRVKELC*S*ADEASCC*L*GC*RGNKEAEPDTAGICLSN*FCIMAGCPL*GN 93
Query: 397 ----------CCMSCFC 403
CC SC C
Sbjct: 92 PTNC*LLNGACCESCSC 42
>AW586908 weakly similar to GP|2104814|emb| pleiotropic effects on cellular
differentiation and slug behaviour {Dictyostelium
discoideum}, partial (8%)
Length = 668
Score = 65.9 bits (159), Expect = 3e-11
Identities = 42/138 (30%), Positives = 49/138 (35%), Gaps = 12/138 (8%)
Frame = -3
Query: 281 LRQSQGPQSNSSSCFSCSCCRMKCCSNNCTGCSNCSCINPECCN-----CSCINPECFNC 335
LR + S C C CC CC N C C NC C + C C C P C
Sbjct: 666 LRLQKWEDVESGCCCCCCCC---CCCNFCNNC-NCCCWSEMFCGTV*DICGCCGPLC--- 508
Query: 336 SCINCSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCC-NLR-CAR 393
N C C + CC C CC +C RP+CC+ CC N C +
Sbjct: 507 ---NMLCCNNCNEDCCN*FC----------------CCCRCKRPACCWSICCWNCESCGK 385
Query: 394 -----PSCCCMSCFCWRC 406
P FCW C
Sbjct: 384 DVIINPFLPADCKFCWDC 331
Score = 33.9 bits (76), Expect = 0.12
Identities = 23/68 (33%), Positives = 24/68 (34%), Gaps = 10/68 (14%)
Frame = -3
Query: 294 CFSCSCC---------RMKCCSNNCTG-CSNCSCINPECCNCSCINPECFNCSCINCSCS 343
C+ C CC CC C C SC CC CS N N CI C C
Sbjct: 210 CWCCCCC*GCCGLC*VS*GCCCCGCGRFCDPASCCCLCCCCCSWSN-GIGNNDCIFCCC- 37
Query: 344 LKCTQCCC 351
CCC
Sbjct: 36 -----CCC 28
Score = 31.2 bits (69), Expect = 0.78
Identities = 16/40 (40%), Positives = 18/40 (45%)
Frame = -3
Query: 296 SCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNC 335
SC CC CC + G N CI CC C C + E C
Sbjct: 114 SC-CCLCCCCCSWSNGIGNNDCI--FCCCCCC*DHEAGGC 4
Score = 28.9 bits (63), Expect = 3.9
Identities = 11/27 (40%), Positives = 11/27 (40%)
Frame = -3
Query: 381 CCFKCCCNLRCARPSCCCMSCFCWRCC 407
CC CCC CCC C CC
Sbjct: 630 CCCCCCC-------CCCCNFCNNCNCC 571
Score = 28.5 bits (62), Expect = 5.1
Identities = 13/41 (31%), Positives = 17/41 (40%), Gaps = 9/41 (21%)
Frame = -3
Query: 294 CFSCSCCRMKCC---------SNNCTGCSNCSCINPECCNC 325
C SCC + CC +N+C C C C + E C
Sbjct: 126 CDPASCCCLCCCCCSWSNGIGNNDCIFCCCCCC*DHEAGGC 4
>TC92466 similar to GP|11141605|gb|AAG32022.1 LEUNIG {Arabidopsis thaliana},
partial (14%)
Length = 632
Score = 62.8 bits (151), Expect = 2e-10
Identities = 33/93 (35%), Positives = 35/93 (37%)
Frame = -1
Query: 297 CSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLPSCI 356
C CC CC +C GC C C CC C C C CI C C CCC C
Sbjct: 566 CCCCGCGCCGCDC-GCC*CCC----CCCCCCA------CLCIRSCCICMCCCCCCCACC- 423
Query: 357 KCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNL 389
C G CC CC CCC+L
Sbjct: 422 ---------DC*G*GCCC-----CCCCCCCCSL 366
Score = 58.9 bits (141), Expect = 3e-09
Identities = 30/86 (34%), Positives = 33/86 (37%)
Frame = -1
Query: 322 CCNCSCINPECFNCSCINCSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSC 381
CC C C +C C C C C CCC CI+ C C CC C
Sbjct: 560 CCGCGCCGCDCGCC*CCCCCC------CCCACLCIRSC-------CICMCCC-------C 441
Query: 382 CFKCCCNLRCARPSCCCMSCFCWRCC 407
C CC+ C CCC C C CC
Sbjct: 440 CCCACCD--C*G*GCCCCCCCC--CC 375
Score = 58.2 bits (139), Expect = 6e-09
Identities = 28/86 (32%), Positives = 31/86 (35%)
Frame = -1
Query: 315 CSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCCT 374
C C CC C C C C C C C+ C + CC+ C CC C
Sbjct: 566 CCCCGCGCCGCDC--GCC*CCCCCCCCCACLCIRSCCICMCCCCC----------CCACC 423
Query: 375 KCSRPSCCFKCCCNLRCARPSCCCMS 400
C CC CCC CCC S
Sbjct: 422 DC*G*GCCCCCCC--------CCCCS 369
Score = 56.2 bits (134), Expect = 2e-08
Identities = 28/80 (35%), Positives = 28/80 (35%), Gaps = 4/80 (5%)
Frame = -1
Query: 332 CFNCSCINCSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLRC 391
C C C C C C CCC C CC C C R C CCC C
Sbjct: 566 CCCCGCGCCGCDCGCC*CCC---CCCCC------------CACLCIRSCCICMCCCCCCC 432
Query: 392 ARPSC----CCMSCFCWRCC 407
A C CC C C CC
Sbjct: 431 ACCDC*G*GCCCCCCCCCCC 372
Score = 45.4 bits (106), Expect = 4e-05
Identities = 21/50 (42%), Positives = 21/50 (42%), Gaps = 3/50 (6%)
Frame = -1
Query: 361 LPKFSTCFGSSCC---TKCSRPSCCFKCCCNLRCARPSCCCMSCFCWRCC 407
L F C G CC C CC CCC C R C CM C C CC
Sbjct: 578 LSLFCCCCGCGCCGCDCGCC*CCCCCCCCCACLCIRSCCICMCCCC--CC 435
Score = 42.0 bits (97), Expect = 4e-04
Identities = 19/51 (37%), Positives = 21/51 (40%)
Frame = -1
Query: 294 CFSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSL 344
C C CC C +C C C C CC+ C C C C C CSL
Sbjct: 512 CCCCCCCCACLCIRSCCICMCCCCCCCACCD--C*G*GCCCCCCCCCCCSL 366
Score = 29.3 bits (64), Expect = 3.0
Identities = 13/36 (36%), Positives = 14/36 (38%)
Frame = -1
Query: 292 SSCFSCSCCRMKCCSNNCTGCSNCSCINPECCNCSC 327
S C CC C +C G C C CC C C
Sbjct: 470 SCCICMCCCCCCCACCDC*G*GCCCC----CCCCCC 375
>CA919710 similar to GP|10177798|dbj emb|CAB87778.1~gene_id:MXA21.9~strong
similarity to unknown protein {Arabidopsis thaliana},
partial (18%)
Length = 738
Score = 57.8 bits (138), Expect = 8e-09
Identities = 20/36 (55%), Positives = 29/36 (80%)
Frame = -1
Query: 234 MARESYDPLWIKGGGHCNLELYPDYIRHLCKFIQEM 269
+ + Y+PLW+ G GHCNLELYP++I+HL KF+Q +
Sbjct: 738 LCKVKYEPLWVSGAGHCNLELYPEFIKHLKKFVQTL 631
>TC80931 similar to GP|14190783|gb|AAF65166.2 putative phloem transcription
factor M1 {Apium graveolens}, partial (67%)
Length = 1193
Score = 55.1 bits (131), Expect = 5e-08
Identities = 38/107 (35%), Positives = 45/107 (41%)
Frame = -3
Query: 304 CCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLPSCIKCCHLPK 363
CC +C CS C+C + CC C CS +C C + CCC SC CC +
Sbjct: 783 CC*GSC*CCSCCNCGSC-CCCC---------CS*ESCCC*S*GSCCCCCWSCGICC*I-- 640
Query: 364 FSTCFGSSCCTKCSRPSCCFKCCCNLRCARPSCCCMSCFCWRCCVGK 410
SC CS SCC CCC CCC+ C C GK
Sbjct: 639 -----WESC*FCCSCGSCC--CCC--------CCCICC----CNCGK 556
Score = 47.0 bits (110), Expect(2) = 1e-08
Identities = 25/63 (39%), Positives = 28/63 (43%), Gaps = 4/63 (6%)
Frame = -3
Query: 293 SCFSCSCCRMKCCSNNCTGC---SNCSCINPECCNCSCINPEC-FNCSCINCSCSLKCTQ 348
+C SC CC CCS C +C C C C I C F CSC +C C C
Sbjct: 747 NCGSCCCC---CCS*ESCCC*S*GSCCCCCWSCGICC*IWESC*FCCSCGSCCCCCCCCI 577
Query: 349 CCC 351
CCC
Sbjct: 576 CCC 568
Score = 44.7 bits (104), Expect = 7e-05
Identities = 23/51 (45%), Positives = 26/51 (50%), Gaps = 6/51 (11%)
Frame = -3
Query: 362 PKFSTCFGSS-CCTKCSRPSCCFKCCCNLRCARPSCCCMS-----CFCWRC 406
P C GS CC+ C+ SCC CCC C+ SCCC S C CW C
Sbjct: 795 PTICCC*GSC*CCSCCNCGSCC--CCC---CS*ESCCC*S*GSCCCCCWSC 658
Score = 32.7 bits (73), Expect = 0.27
Identities = 8/13 (61%), Positives = 9/13 (68%)
Frame = -1
Query: 396 CCCMSCFCWRCCV 408
CCC C CW CC+
Sbjct: 554 CCCCDCICWCCCI 516
Score = 32.7 bits (73), Expect = 0.27
Identities = 17/41 (41%), Positives = 22/41 (53%)
Frame = -1
Query: 313 SNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLP 353
++ + I CC C CI C+ C CI C C + C CCC P
Sbjct: 581 ASVAAIVERCCCCDCI---CW-CCCI-CCCCICC--CCC*P 480
Score = 31.6 bits (70), Expect = 0.60
Identities = 11/25 (44%), Positives = 13/25 (52%)
Frame = -1
Query: 335 CSCINCSCSLKCTQCCCLPSCIKCC 359
C C +C C C CCC+ C CC
Sbjct: 554 CCCCDCICWCCCICCCCI--CCCCC 486
Score = 29.6 bits (65), Expect(2) = 1e-08
Identities = 12/27 (44%), Positives = 13/27 (47%)
Frame = -1
Query: 372 CCTKCSRPSCCFKCCCNLRCARPSCCC 398
CC C CC CCC + C CCC
Sbjct: 551 CCCDCICWCCCICCCC-ICC----CCC 486
Score = 28.1 bits (61), Expect = 6.6
Identities = 10/23 (43%), Positives = 11/23 (47%)
Frame = -1
Query: 381 CCFKCCCNLRCARPSCCCMSCFC 403
CC C C C CCC+ C C
Sbjct: 551 CCCDCICWCCCI--CCCCICCCC 489
>BG644345 weakly similar to GP|19697333|gb putative protein potential
transcriptional repressor Not4hp - Mus musculus, partial
(7%)
Length = 635
Score = 56.6 bits (135), Expect = 2e-08
Identities = 40/133 (30%), Positives = 47/133 (35%), Gaps = 18/133 (13%)
Frame = -1
Query: 295 FSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLPS 354
++C CC CC ++C C C N CNC C C C C +C C CCC
Sbjct: 479 YNCCCCDD*CCCSSCC----CCCCN---CNCCCC-*SCCCCCC*SCCC------CCC--- 351
Query: 355 CIKCCHLPKFSTCFGSSCCTKCSR-------PSCCF-------KCCCNLRCARPSCCCMS 400
CC C SCC C + CC+ CCC CCC
Sbjct: 350 ---CC-------C*N*SCCCCCEKGVI*ENGDGCCWPGVWNGDACCCPGVWNGDDCCCPG 201
Query: 401 CFCWR----CCVG 409
W CC G
Sbjct: 200 --VWNGDACCCPG 168
Score = 54.3 bits (129), Expect = 9e-08
Identities = 38/111 (34%), Positives = 43/111 (38%), Gaps = 6/111 (5%)
Frame = -1
Query: 294 CFSCSCCRMKCCSNNCTGC-SNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCL 352
C SC CC CC+ NC C S C C CC C C C C N SC CCC
Sbjct: 446 CSSCCCC---CCNCNCCCC*SCCCCCC*SCCCCCC-------CCC*N*SCC-----CCCE 312
Query: 353 PSCI-----KCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLRCARPSCCC 398
I CC ++ G +CC C CCC +CCC
Sbjct: 311 KGVI*ENGDGCCWPGVWN---GDACC--CPGVWNGDDCCCPGVWNGDACCC 174
Score = 44.7 bits (104), Expect = 7e-05
Identities = 35/111 (31%), Positives = 39/111 (34%), Gaps = 4/111 (3%)
Frame = -1
Query: 291 SSSCFSCSCCRMKCC-SNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQC 349
SS C C C CC S C C +C C CC C C N C C C +
Sbjct: 443 SSCCCCCCNCNCCCC*SCCCCCC*SCCC----CCCCCC*N*SC--CCCCEKGVI*ENGDG 282
Query: 350 CCLPSCIK---CCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLRCARPSCC 397
CC P CC ++ G CC C CCC R CC
Sbjct: 281 CCWPGVWNGDACCCPGVWN---GDDCC--CPGVWNGDACCCPGVWNRDGCC 144
>TC85575 similar to PIR|D86410|D86410 protein F3M18.16 [imported] -
Arabidopsis thaliana, partial (15%)
Length = 1225
Score = 48.9 bits (115), Expect = 4e-06
Identities = 37/144 (25%), Positives = 52/144 (35%), Gaps = 36/144 (25%)
Frame = -2
Query: 300 CRMKCCSNNCTGCS---NCSCINPECCNCSCINPEC----FNCSCIN-----------CS 341
C S++C C +C C + C CS + C F SC C
Sbjct: 891 CHSFLASHHCCNCCYF*SCICH*QQLCCCSYTSRSCCCSDFALSCNLLLGTLCC*KNLCC 712
Query: 342 CSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCC--NLRCA------- 392
C + T CCC C L + C + C C R + C++CCC + R +
Sbjct: 711 CCCRNTCCCCCSGFPWSCILLL*TLCC*KNPCCCCCR-NTCYRCCCCSDFRSSYILLLGT 535
Query: 393 ---------RPSCCCMSCFCWRCC 407
+ +CCC S C CC
Sbjct: 534 FCC*SHQN*KNTCCCRSYCCCCCC 463
Score = 48.1 bits (113), Expect = 6e-06
Identities = 42/138 (30%), Positives = 51/138 (36%), Gaps = 32/138 (23%)
Frame = -2
Query: 304 CCSNNCTGCSNCSCINPECCNCS---------------CINPECFN---------CSCIN 339
C S+NC+ C C NP CC S I+P+ + SC +
Sbjct: 1056 CYSHNCSHC--CYQSNPFCCTNSQSSQLL*MNLH*MLGYISPQS*HHKNIFHLP*LSCHS 883
Query: 340 CSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCC--FKCCCNLR----CAR 393
S C CC SCI CH + CC+ SR CC F CNL C
Sbjct: 882 FLASHHCCNCCYF*SCI--CH*QQLC------CCSYTSRSCCCSDFALSCNLLLGTLCC* 727
Query: 394 PSCCCMSC--FCWRCCVG 409
+ CC C C CC G
Sbjct: 726 KNLCCCCCRNTCCCCCSG 673
Score = 43.1 bits (100), Expect = 2e-04
Identities = 33/125 (26%), Positives = 36/125 (28%), Gaps = 34/125 (27%)
Frame = -2
Query: 297 CSCCRMKCCSNNCTGCSNCSCI----------NPECCNCSCINPECFNCSCIN------- 339
C CCR CC C SCI NP CC C C+ C C +
Sbjct: 714 CCCCRNTCCC--CCSGFPWSCILLL*TLCC*KNPCCC---CCRNTCYRCCCCSDFRSSYI 550
Query: 340 -----------------CSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCC 382
C C C CCC + CH TC C S
Sbjct: 549 LLLGTFCC*SHQN*KNTCCCRSYCCCCCCKMLVLVLCH*LHSGTC----CRNLAGS*SNY 382
Query: 383 FKCCC 387
CCC
Sbjct: 381 *LCCC 367
>TC91905 similar to GP|19347881|gb|AAL85997.1 unknown protein {Arabidopsis
thaliana}, partial (67%)
Length = 867
Score = 47.4 bits (111), Expect = 1e-05
Identities = 27/106 (25%), Positives = 50/106 (46%), Gaps = 1/106 (0%)
Frame = +1
Query: 78 TLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGASTGKPSESSTYADIEAIYEC 137
T+L+ NA ++ ++ + L+ N+ Y GYGAS G PS+ D +A +
Sbjct: 481 TILFFQENAGNIAHRLEMVRIMLQQLQCNVFMLSYRGYGASEGSPSQKGITKDAQAALDH 660
Query: 138 LESEYGVGQEDIILYGQSVGSGPTLHLAAKLP-RLRGVVLHSGILS 182
L + I+++G+S+G L P ++ G++L + S
Sbjct: 661 LSQRSDIDTSRIVVFGRSLGGAVGAVLTRNNPEKVAGLILENTFTS 798
>TC93379 similar to GP|19569994|gb|AAL92295.1 hypothetical protein
{Dictyostelium discoideum}, partial (1%)
Length = 535
Score = 45.4 bits (106), Expect = 4e-05
Identities = 22/59 (37%), Positives = 23/59 (38%)
Frame = -2
Query: 349 CCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLRCARPSCCCMSCFCWRCC 407
C P I CC F CC C CC CCC + A CCC C C CC
Sbjct: 282 CGLPPPYIICCGYMLFMFPCCICCCCSCCCICCC--CCCCIIIAMYCCCCCCC*CCCCC 112
Score = 40.0 bits (92), Expect = 0.002
Identities = 26/76 (34%), Positives = 29/76 (37%)
Frame = -2
Query: 333 FNCSCINCSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLRCA 392
F C CI C CS CCC+ C CC + C CC CC CCC
Sbjct: 231 FPC-CICCCCS-----CCCICCCCCCCIIIAMYCC----CC-------CCC*CCC----- 118
Query: 393 RPSCCCMSCFCWRCCV 408
CC F CC+
Sbjct: 117 ---CC***GFGCICCI 79
Score = 39.3 bits (90), Expect = 0.003
Identities = 19/53 (35%), Positives = 20/53 (36%)
Frame = -2
Query: 299 CCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCC 351
CC C C CSC CC C CI + C C C C CCC
Sbjct: 255 CCGYMLFMFPCCICCCCSCCCICCCCCCCIIIAMYCCCCCCC*C------CCC 115
Score = 35.4 bits (80), Expect = 0.041
Identities = 22/63 (34%), Positives = 24/63 (37%), Gaps = 3/63 (4%)
Frame = -2
Query: 294 CFSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSC---SLKCTQCC 350
C CSCC + CC C C I CC C C C C C C C C
Sbjct: 216 CCCCSCCCICCC------CCCCIIIAMYCCCCCC-------C*CC-CCC***GFGC--IC 85
Query: 351 CLP 353
C+P
Sbjct: 84 CIP 76
Score = 27.7 bits (60), Expect = 8.6
Identities = 16/47 (34%), Positives = 16/47 (34%)
Frame = -2
Query: 291 SSSCFSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSC 337
S C C CC CC C C C C C C F C C
Sbjct: 204 SCCCICCCCC---CCIIIAMYCCCCCC----C*CCCCC***GFGCIC 85
>TC79220 similar to GP|2149570|gb|AAB58577.1| MAP kinase kinase protein
DdMEK1 {Dictyostelium discoideum}, partial (6%)
Length = 1162
Score = 43.5 bits (101), Expect = 2e-04
Identities = 27/91 (29%), Positives = 38/91 (41%), Gaps = 2/91 (2%)
Frame = -3
Query: 324 NCSCINPECFNCSCINCSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCF 383
+C+C + E +N C CCC + + CC + +C C S F
Sbjct: 569 HCTCYS-EHYNYYNYCCGMIFHLQHCCC*SNTLGCCFVFPSWSCNLLHCILCHSTHFGYF 393
Query: 384 -KC-CCNLRCARPSCCCMSCFCWRCCVGKHS 412
KC CCN C CC +C+CW C+ S
Sbjct: 392 QKCLCCNTLCYMHHCCS-TCYCWGNCLSLQS 303
>TC88336 similar to PIR|T51477|T51477 glutamine-rich protein - Arabidopsis
thaliana, partial (67%)
Length = 1297
Score = 43.1 bits (100), Expect = 2e-04
Identities = 25/66 (37%), Positives = 28/66 (41%)
Frame = -2
Query: 322 CCNCSCINPECFNCSCINCSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSC 381
CC C C C C NC S+KC CCC C +CC + S CC C
Sbjct: 321 CCCCCC-------CFC-NCCKSIKC--CCC---CCRCC------S*INSCCC-------C 220
Query: 382 CFKCCC 387
C CCC
Sbjct: 219 CCCCCC 202
Score = 42.4 bits (98), Expect = 3e-04
Identities = 21/56 (37%), Positives = 24/56 (42%)
Frame = -2
Query: 296 SCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCC 351
+ +CC CC NC C + C CC C C CS IN C C CCC
Sbjct: 330 AAACCCCCCCFCNC--CKSIKCC---CCCCRC-------CS*INSCCCCCCCCCCC 199
Score = 40.4 bits (93), Expect = 0.001
Identities = 21/59 (35%), Positives = 23/59 (38%)
Frame = -2
Query: 349 CCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLRCARPSCCCMSCFCWRCC 407
CCC CC CF C C CC CCC C+ + CC C C CC
Sbjct: 318 CCC------CC-------CF----CNCCKSIKCC--CCCCRCCS*INSCCCCCCCCCCC 199
Score = 39.7 bits (91), Expect = 0.002
Identities = 16/45 (35%), Positives = 17/45 (37%)
Frame = -2
Query: 315 CSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLPSCIKCC 359
C C CC C C + C C C C CCC C CC
Sbjct: 321 CCC----CCCCFCNCCKSIKCCCCCCRCCS*INSCCCCCCCCCCC 199
Score = 39.7 bits (91), Expect = 0.002
Identities = 20/59 (33%), Positives = 23/59 (38%)
Frame = -2
Query: 349 CCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLRCARPSCCCMSCFCWRCC 407
CCC CC C +CC CC +CC + SCCC C C CC
Sbjct: 321 CCC------CC-------CCFCNCCKSIKCCCCCCRCCS*IN----SCCCCCCCC--CC 202
Score = 39.7 bits (91), Expect = 0.002
Identities = 17/45 (37%), Positives = 22/45 (48%), Gaps = 3/45 (6%)
Frame = -2
Query: 286 GPQSNSSSCFSCSCCRMKCCSNN---CTGCSNCSCINPECCNCSC 327
G + +++C C CC CC + C C CS IN CC C C
Sbjct: 345 GNREIAAACCCCCCCFCNCCKSIKCCCCCCRCCS*INSCCCCCCC 211
Score = 37.0 bits (84), Expect = 0.014
Identities = 13/28 (46%), Positives = 15/28 (53%)
Frame = -2
Query: 380 SCCFKCCCNLRCARPSCCCMSCFCWRCC 407
+CC CCC C + CC C C RCC
Sbjct: 324 ACCCCCCCFCNCCKSIKCC--CCCCRCC 247
Score = 35.4 bits (80), Expect = 0.041
Identities = 23/80 (28%), Positives = 28/80 (34%), Gaps = 1/80 (1%)
Frame = -2
Query: 322 CCNCSCINPECFNCSCINCSCSLKCTQCCC-LPSCIKCCHLPKFSTCFGSSCCTKCSRPS 380
CC C C C C I C C C +CC + SC CC CC C S
Sbjct: 318 CCCCCCF---CNCCKSIKCCCC--CCRCCS*INSCCCCC------------CCCCCC**S 190
Query: 381 CCFKCCCNLRCARPSCCCMS 400
L + C C++
Sbjct: 189 AFASSIFFLNLEKIPCVCVA 130
>TC77589 similar to GP|1087015|gb|AAB35283.1| arabinogalactan-protein; AGP
{Pyrus communis}, partial (10%)
Length = 1064
Score = 43.1 bits (100), Expect = 2e-04
Identities = 19/36 (52%), Positives = 20/36 (54%)
Frame = -1
Query: 324 NCSCINPECFNCSCINCSCSLKCTQCCCLPSCIKCC 359
NC C P F+CSC NCSCSL C C C CC
Sbjct: 542 NCCCKTP--FHCSCCNCSCSLHCCSFLCTLHC--CC 447
Score = 34.7 bits (78), Expect = 0.071
Identities = 18/42 (42%), Positives = 20/42 (46%), Gaps = 4/42 (9%)
Frame = -1
Query: 314 NCSCINP---ECCNCSCINPECFNCSCINCSCSLKCT-QCCC 351
NC C P CCNCSC ++C CS CT CCC
Sbjct: 542 NCCCKTPFHCSCCNCSC---------SLHC-CSFLCTLHCCC 447
Score = 32.0 bits (71), Expect = 0.46
Identities = 14/42 (33%), Positives = 15/42 (35%)
Frame = -1
Query: 346 CTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCC 387
C L S CC P +C SC C C CCC
Sbjct: 572 CHSFLLLSSHNCCCKTPFHCSCCNCSCSLHCCSFLCTLHCCC 447
Score = 30.8 bits (68), Expect = 1.0
Identities = 16/48 (33%), Positives = 18/48 (37%), Gaps = 4/48 (8%)
Frame = -1
Query: 359 CHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLRC----ARPSCCCMSCF 402
CH F +CC K C C C+L C CCC S F
Sbjct: 572 CH--SFLLLSSHNCCCKTPFHCSCCNCSCSLHCCSFLCTLHCCCGSLF 435
>TC92665 homologue to GP|15291583|gb|AAK93060.1 GH28569p {Drosophila
melanogaster}, partial (4%)
Length = 668
Score = 38.1 bits (87), Expect(2) = 2e-04
Identities = 26/74 (35%), Positives = 29/74 (39%), Gaps = 6/74 (8%)
Frame = -1
Query: 339 NCSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNL--RCARPS- 395
NC C C C P C C + F CC +C R F C CNL C P+
Sbjct: 467 NCCCC--CNPCTLFPGC*-CIFVRLF**FINCCCCCRCCRFMRLFCCGCNLWDFCTSPTL 297
Query: 396 -CCCMSCFCW--RC 406
CC CW RC
Sbjct: 296 FCCPSI*ACWPLRC 255
Score = 31.6 bits (70), Expect(2) = 0.031
Identities = 23/70 (32%), Positives = 24/70 (33%), Gaps = 9/70 (12%)
Frame = -1
Query: 298 SCCRMKCCSNNCTGCSNCSCINPE--------CCNCSCIN-PECFNCSCINCSCSLKCTQ 348
+CC CC N CT C CI CC C C F C C T
Sbjct: 467 NCC---CCCNPCTLFPGC*CIFVRLF**FINCCCCCRCCRFMRLFCCGCNLWDFCTSPTL 297
Query: 349 CCCLPSCIKC 358
CC PS C
Sbjct: 296 FCC-PSI*AC 270
Score = 23.9 bits (50), Expect(2) = 2e-04
Identities = 11/32 (34%), Positives = 15/32 (46%), Gaps = 3/32 (9%)
Frame = -3
Query: 300 CRMKCCSNNCTG--CS-NCSCINPECCNCSCI 328
C + CC C C + + + CCNC CI
Sbjct: 639 CDINCC*EACVWDCCD*DGNIFDGLCCNCCCI 544
Score = 23.1 bits (48), Expect(2) = 0.031
Identities = 11/34 (32%), Positives = 13/34 (37%), Gaps = 5/34 (14%)
Frame = -2
Query: 379 PSCCFKCC-----CNLRCARPSCCCMSCFCWRCC 407
P+ C CC C CCC+ W CC
Sbjct: 241 PASC*CCC*IIGVCC*LTIEFCCCCL--IIWECC 146
>TC90558 similar to GP|9758584|dbj|BAB09197.1
gb|AAC80623.1~gene_id:MBD2.15~similar to unknown protein
{Arabidopsis thaliana}, partial (5%)
Length = 1047
Score = 42.7 bits (99), Expect = 3e-04
Identities = 21/76 (27%), Positives = 31/76 (40%)
Frame = -2
Query: 331 ECFNCSCINCSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLR 390
+C++C+ SC C + C SC CC + + + GS C + +C CN
Sbjct: 959 DCWSCNIFESSCCCCCRKSSCCSSCFCCCFINRSLSNRGS-----CGAEAAA*ECNCNKY 795
Query: 391 CARPSCCCMSCFCWRC 406
C CC C C
Sbjct: 794 C----CCNRFIICGSC 759
Score = 33.5 bits (75), Expect = 0.16
Identities = 24/77 (31%), Positives = 28/77 (36%), Gaps = 2/77 (2%)
Frame = -2
Query: 292 SSCFSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCC 351
SSC C CCR C C+ C C IN N E C C + CC
Sbjct: 932 SSC--CCCCRKSSC---CSSCFCCCFINRSLSNRGSCGAEAAA*EC-------NCNKYCC 789
Query: 352 LPSCIKC--CHLPKFST 366
I C C P+ S+
Sbjct: 788 CNRFIICGSCDKPEESS 738
Score = 30.8 bits (68), Expect = 1.0
Identities = 23/78 (29%), Positives = 30/78 (37%)
Frame = -2
Query: 302 MKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLPSCIKCCHL 361
+ C S N S C C C CS CF C IN S S + + C + C+
Sbjct: 962 LDCWSCNIFESSCCCCCRKSSC-CS----SCFCCCFINRSLSNRGS--CGAEAAA*ECNC 804
Query: 362 PKFSTCFGSSCCTKCSRP 379
K+ C C C +P
Sbjct: 803 NKYCCCNRFIICGSCDKP 750
>TC79570 similar to PIR|F86427|F86427 auxin response factor 6 (ARF6)
[imported] - Arabidopsis thaliana, partial (15%)
Length = 1143
Score = 42.7 bits (99), Expect = 3e-04
Identities = 25/73 (34%), Positives = 26/73 (35%), Gaps = 11/73 (15%)
Frame = -1
Query: 347 TQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLRCARPSCCCMSCF---- 402
T CCC C CC CC C CC CCC C CCC+ F
Sbjct: 672 TTCCC---CCDCC-----------CCCC*CC---CC--CCCCCCCCCCCCCCI**F*LVN 550
Query: 403 -------CWRCCV 408
CW CCV
Sbjct: 549 EC*C*RSCWNCCV 511
Score = 40.8 bits (94), Expect = 0.001
Identities = 24/67 (35%), Positives = 26/67 (37%), Gaps = 7/67 (10%)
Frame = -1
Query: 291 SSSCFSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCI-------NCSCS 343
S++C C CC CC C C C C CC C C C C CI C C
Sbjct: 675 STTC--CCCCDCCCC---CC*CCCCCC----CCCCCC----CCCCCCI**F*LVNEC*C* 535
Query: 344 LKCTQCC 350
C CC
Sbjct: 534 RSCWNCC 514
Score = 37.7 bits (86), Expect = 0.008
Identities = 17/50 (34%), Positives = 18/50 (36%), Gaps = 12/50 (24%)
Frame = -1
Query: 322 CCNCSCINPECFNCSCINCSCSLKCTQCCCL------------PSCIKCC 359
CC C C C C C C C C CCC+ SC CC
Sbjct: 663 CCCCDCCCCCC*CCCCCCCCCCCCCCCCCCI**F*LVNEC*C*RSCWNCC 514
Score = 36.6 bits (83), Expect = 0.019
Identities = 17/44 (38%), Positives = 19/44 (42%)
Frame = -1
Query: 313 SNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLPSCI 356
+ C C CC+C C C C C C C C CCC CI
Sbjct: 672 TTCCC----CCDCCCC---CC*CCCCCCCC---CCCCCCCCCCI 571
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.139 0.477
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,746,943
Number of Sequences: 36976
Number of extensions: 653850
Number of successful extensions: 12328
Number of sequences better than 10.0: 408
Number of HSP's better than 10.0 without gapping: 7699
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 10514
length of query: 420
length of database: 9,014,727
effective HSP length: 99
effective length of query: 321
effective length of database: 5,354,103
effective search space: 1718667063
effective search space used: 1718667063
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0230.1


