

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0228a.2
(510 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
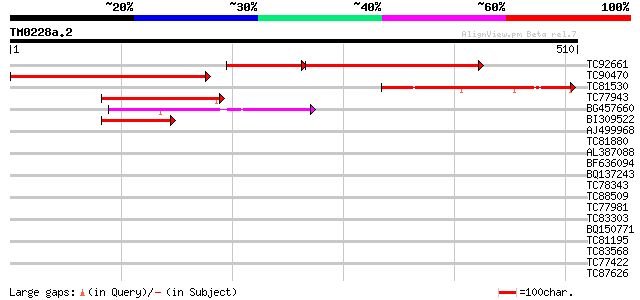
Score E
Sequences producing significant alignments: (bits) Value
TC92661 similar to GP|22093674|dbj|BAC06968. kinase-like protein... 232 3e-84
TC90470 weakly similar to GP|9294219|dbj|BAB02121.1 gb|AAF01563.... 258 3e-69
TC81530 similar to GP|10177440|dbj|BAB10736. gb|AAC55944.1~gene_... 139 3e-33
TC77943 similar to GP|10179602|gb|AAG13810.1 PSTVd RNA-binding p... 112 4e-25
BG457660 similar to PIR|T00472|T00 probable RING3 protein [impor... 85 8e-17
BI309522 similar to PIR|A86198|A86 hypothetical protein [importe... 82 6e-16
AJ499968 weakly similar to GP|23495961|gb| hypothetical protein ... 41 0.001
TC81880 similar to PIR|B85431|B85431 trichohyalin like protein [... 39 0.004
AL387088 similar to GP|18266427|gb| cytoplasmic dynein intermedi... 37 0.018
BF636094 similar to GP|15983759|g At1g05910/T20M3_16 {Arabidopsi... 37 0.024
BQ137243 similar to GP|2624970|gb|A proline-rich protein 9-1 {Mu... 35 0.069
TC78343 weakly similar to GP|15724195|gb|AAL06489.1 AT5g49210/K2... 35 0.090
TC88509 similar to PIR|T01826|T01826 microfibril-associated prot... 34 0.12
TC77981 similar to PIR|T12970|T12970 hypothetical protein T6H20.... 33 0.20
TC83303 weakly similar to GP|20503004|gb|AAM22713.1 Hypothetical... 33 0.26
BQ150771 GP|4028979|emb glutamate permease {synthetic construct}... 33 0.34
TC81195 similar to GP|15810597|gb|AAL07186.1 unknown protein {Ar... 32 0.44
TC83568 weakly similar to GP|16604681|gb|AAL24133.1 putative kin... 32 0.58
TC77422 similar to PIR|T04704|T04704 hypothetical protein F4B14.... 31 0.99
TC87626 similar to GP|20259075|gb|AAM14253.1 unknown protein {Ar... 31 0.99
>TC92661 similar to GP|22093674|dbj|BAC06968. kinase-like protein {Oryza
sativa (japonica cultivar-group)}, partial (13%)
Length = 702
Score = 232 bits (592), Expect(2) = 3e-84
Identities = 124/160 (77%), Positives = 131/160 (81%)
Frame = +2
Query: 267 DRIAPGAVAFKQKRQIKSTSPLERKSDPDSDGAVSSLDSEHVYPGSQHATPATDASSGEV 326
D GA A KQ Q K TSPL+RKSDP SDGAVSSLDSEH P S HAT ATDASS EV
Sbjct: 218 DNYXAGASALKQDCQAKCTSPLQRKSDPGSDGAVSSLDSEHACPSSPHATLATDASSVEV 397
Query: 327 WSTPDFPVQLSPKKALRAAMLKSRFADTILKAQQKTLLEHGDKGDPLKMQLEKERLERIQ 386
W+TP PVQLSPK+ALR AML+SRFA TILKAQQ TLL+HGDKGDP+KMQLEKERLERIQ
Sbjct: 398 WNTPVLPVQLSPKRALRYAMLRSRFAGTILKAQQNTLLKHGDKGDPMKMQLEKERLERIQ 577
Query: 387 REERARIEAQIKTAEAAARARAEEELRQQREKEREAARAA 426
REE+ARIEAQIK AEAA R RAEEELRQQ EKER A AA
Sbjct: 578 REEQARIEAQIKAAEAAERTRAEEELRQQIEKERXXAXAA 697
Score = 98.2 bits (243), Expect(2) = 3e-84
Identities = 49/72 (68%), Positives = 57/72 (79%)
Frame = +3
Query: 196 GTIKETARKSCNVMHPRQKDTFPKKSQVSEHKRVLKISSLAARDAKVEVPKLSQVHCKSI 255
G IKET RKS +V R KD+F K S+ SEHK + KISSLA RDAKVEVPKLSQ+ CK +
Sbjct: 3 GMIKETVRKSYDVTQSRHKDSFTKSSRASEHKGIPKISSLATRDAKVEVPKLSQIPCKPL 182
Query: 256 EKDLIKGSEERD 267
EKDLIKGSE+R+
Sbjct: 183 EKDLIKGSEDRE 218
>TC90470 weakly similar to GP|9294219|dbj|BAB02121.1
gb|AAF01563.1~gene_id:K17E12.8~similar to unknown
protein {Arabidopsis thaliana}, partial (11%)
Length = 772
Score = 258 bits (660), Expect = 3e-69
Identities = 126/180 (70%), Positives = 141/180 (78%)
Frame = +3
Query: 1 MIATETIVPTRKLKIKFSTKRIEVDPGQKCEFGQRGLQTDENGRCNSAEKLFKPDSIKRG 60
MIATE I P KLKIKFSTKRIEV G KC+FG++ Q DENGRCN +K S KR
Sbjct: 231 MIATEIIAPATKLKIKFSTKRIEVVSGPKCQFGEKVSQLDENGRCNPIKKSSLLGSNKRE 410
Query: 61 PPEGFEGQKEKRQKIDRKGSTQCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKP 120
P EG K KRQK+DRKGS +CA ILKCL+S+PYSWVF PVDPVALNIPDYF +I+ P
Sbjct: 411 PSGNIEGPKNKRQKLDRKGSHKCATILKCLISHPYSWVFKTPVDPVALNIPDYFTVISHP 590
Query: 121 MDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDM 180
MDLGTI KL KN+Y EEFAADVRLTFSNAMTYNPP NDVH+MAKELN++F+ KWKDM
Sbjct: 591 MDLGTIKFKLDKNIYYSKEEFAADVRLTFSNAMTYNPPSNDVHLMAKELNKLFERKWKDM 770
>TC81530 similar to GP|10177440|dbj|BAB10736.
gb|AAC55944.1~gene_id:K9H21.2~similar to unknown protein
{Arabidopsis thaliana}, partial (24%)
Length = 1135
Score = 139 bits (350), Expect = 3e-33
Identities = 88/188 (46%), Positives = 125/188 (65%), Gaps = 13/188 (6%)
Frame = +1
Query: 335 QLSPKKALRAAMLKSRFADTILKAQQKTLLEHGDKGDPLKMQLEKERLERIQREERARIE 394
Q+SP+K R A+L+SRFADTILKAQ+K L E +K DP K+++E+E LER Q+EE+AR++
Sbjct: 319 QVSPEKLYRQALLRSRFADTILKAQEKAL-EKDEKRDPEKLRIEREELERRQKEEKARLQ 495
Query: 395 AQIKTAEAAAR---ARAEEELRQQREKEREAARAAIEEMKRTVEIEHNLEIVKELETLSG 451
A+ K AE A R A A E +++RE EREAAR A++ M++TVEI + + +++LE LS
Sbjct: 496 AEAKAAEEARRKAEAEAAAEAKRKRELEREAARQALQNMEKTVEINESCQFLEDLEMLSA 675
Query: 452 C----TLSYKALGGRNDYKVALETLDKLQFENPLEQLGLFIKDELADEDEEVINGT---- 503
T S+K +D++ + KLQ NPLEQLGL++K + DE+EE+
Sbjct: 676 VHDEKTPSFKEEASPDDHQNGFGGI-KLQ-GNPLEQLGLYMKVDDEDEEEELPQSAAEPS 849
Query: 504 --VEEGEI 509
VEEGEI
Sbjct: 850 KDVEEGEI 873
>TC77943 similar to GP|10179602|gb|AAG13810.1 PSTVd RNA-binding protein
Virp1a {Lycopersicon esculentum}, partial (43%)
Length = 1783
Score = 112 bits (279), Expect = 4e-25
Identities = 57/114 (50%), Positives = 75/114 (65%), Gaps = 3/114 (2%)
Frame = +3
Query: 83 CAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFA 142
C IL+ LM W+F+ PVDPVALN+ DYF II PMDLGT+ SKL KN YS EFA
Sbjct: 651 CGQILQKLMKTKIGWIFSSPVDPVALNLHDYFDIIKHPMDLGTVKSKLAKNAYSTPAEFA 830
Query: 143 ADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKKW---KCEDELGKS 193
DV+LTF NA+TYNP +DV+ A +L + F+ ++ + +K+ +DEL S
Sbjct: 831 DDVKLTFKNALTYNPKGHDVNTAAMQLLEKFEELYRPIQEKFDEKSFDDELQAS 992
>BG457660 similar to PIR|T00472|T00 probable RING3 protein [imported] -
Arabidopsis thaliana, partial (34%)
Length = 654
Score = 84.7 bits (208), Expect = 8e-17
Identities = 64/191 (33%), Positives = 99/191 (51%), Gaps = 5/191 (2%)
Frame = +3
Query: 90 LMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNV---YSCVEEFAADVR 146
+ + ++W F +PVD L + DY+ II KPMD GTI +K++ Y V E ADVR
Sbjct: 96 ITQHKWAWPFLEPVDVEGLGLHDYYEIIDKPMDFGTIKNKMEAKDGTGYKNVREIYADVR 275
Query: 147 LTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKKWKCEDELGKSVTGTIKETARKSC 206
L F NAM YN +DVH+MAK L + F+ KW + K E++ I+E A+
Sbjct: 276 LIFKNAMKYNNEKHDVHVMAKTLMEKFEDKWLLLLPKVAEEEK------RQIEEEAQVQM 437
Query: 207 NVMHPRQKDTFPKKSQ-VSEHKRVLKISSLAARDAKVE-VPKLSQVHCKSIEKDLIKGSE 264
++ H Q+ T+ ++ +S + I + R+ ++ KLS K++ K + K S
Sbjct: 438 DI-HLAQETTYADMAKDLSNELNEVGIRLMEFREKVIQNCRKLSTGEKKALGKAIAKLSP 614
Query: 265 ERDRIAPGAVA 275
E + A VA
Sbjct: 615 ENLQRALDLVA 647
>BI309522 similar to PIR|A86198|A86 hypothetical protein [imported] -
Arabidopsis thaliana, partial (17%)
Length = 729
Score = 81.6 bits (200), Expect = 6e-16
Identities = 37/67 (55%), Positives = 49/67 (72%)
Frame = +1
Query: 83 CAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFA 142
C+++L+ LM + + WVFN PVD L + DYF II++PMDLGT+ S+L KN Y +EFA
Sbjct: 529 CSSLLEKLMKHKHGWVFNTPVDVERLGLHDYFIIISQPMDLGTVKSRLSKNWYKSPKEFA 708
Query: 143 ADVRLTF 149
DVRLTF
Sbjct: 709 EDVRLTF 729
>AJ499968 weakly similar to GP|23495961|gb| hypothetical protein {Plasmodium
falciparum 3D7}, partial (19%)
Length = 342
Score = 41.2 bits (95), Expect = 0.001
Identities = 27/99 (27%), Positives = 49/99 (49%), Gaps = 10/99 (10%)
Frame = -3
Query: 359 QQKTLLEHGDKGDPLKMQLEKERLERIQREERARIEAQIKTAEAAARARAEEELRQQREK 418
+++ E ++ + ++ EKE ER +REER R+E + E R R E+E R++RE+
Sbjct: 334 EERERKEREEREEQERIAKEKEEQERKEREERERLELERIAKEKEERERKEKEEREERER 155
Query: 419 ----------EREAARAAIEEMKRTVEIEHNLEIVKELE 447
+EA A++E + ++ + KE E
Sbjct: 154 LEAVVRIEKERKEAEEKALKEQEEKALLKEGVIDKKEEE 38
Score = 34.3 bits (77), Expect = 0.12
Identities = 39/130 (30%), Positives = 59/130 (45%), Gaps = 1/130 (0%)
Frame = -3
Query: 378 EKERLERIQREERARIEAQIKTAEAAARARAEEELRQQREKER-EAARAAIEEMKRTVEI 436
EKE ER +REER + E A+ + E+E +++ E+ER E R A E+ +R +
Sbjct: 340 EKEERERKEREER-------EEQERIAKEKEEQERKEREERERLELERIAKEKEERERKE 182
Query: 437 EHNLEIVKELETLSGCTLSYKALGGRNDYKVALETLDKLQFENPLEQLGLFIKDELADED 496
+ E + LE + + + K A E K Q E L + G+ K E E+
Sbjct: 181 KEEREERERLEAV---------VRIEKERKEAEEKALKEQEEKALLKEGVIDKKE---EE 38
Query: 497 EEVINGTVEE 506
E T EE
Sbjct: 37 EPKSKATPEE 8
>TC81880 similar to PIR|B85431|B85431 trichohyalin like protein [imported] -
Arabidopsis thaliana, partial (4%)
Length = 1043
Score = 39.3 bits (90), Expect = 0.004
Identities = 28/70 (40%), Positives = 39/70 (55%), Gaps = 1/70 (1%)
Frame = +2
Query: 364 LEHGDKGDPLKMQLEKERLERIQREERARIEAQIKTAEAAARARAEEELRQQREKEREAA 423
++ D+ ++ +LEKERL +I+ ER R E Q K A RA E E ++REK+R A
Sbjct: 431 VDKDDEKQKIERELEKERLRKIEELERER-ERQ-KDRMAVDRAMLEAEREREREKDRAAV 604
Query: 424 -RAAIEEMKR 432
RA E R
Sbjct: 605 DRATFEARDR 634
Score = 31.6 bits (70), Expect = 0.76
Identities = 70/290 (24%), Positives = 119/290 (40%), Gaps = 21/290 (7%)
Frame = +2
Query: 166 AKELNQI-FDSKWKDMDKKWKCEDELGKSVTGTIKETARKSCNVMHPRQKDTFPKKSQVS 224
AK + Q+ FDS+ ++ K ELG+S TI+ + N R + + +
Sbjct: 131 AKAVPQMEFDSRSQER----KIGHELGES--RTIQHHVNVALNQEGSRNLKSSSQVNTCD 292
Query: 225 EHKRVLKISSLAARDAKVEVPKLSQVHC-----KSIEKDLIK--GSEERDRIAPGAVAFK 277
++ + + A +V V SQ C KS EK + K S ++D
Sbjct: 293 DYIKNTTVGEPTAVQEEVNVHNTSQRSCAAHSTKSKEKCVSKTPASVDKDDEKQKIEREL 472
Query: 278 QKRQIKSTSPLERKSDPDSDGAV---SSLDSEHVYPGSQHATPATDASSGEVWSTPDFPV 334
+K +++ LER+ + D + L++E + A+ + D
Sbjct: 473 EKERLRKIEELERERERQKDRMAVDRAMLEAEREREREKDRAAVDRAT----FEARDRAY 640
Query: 335 QLSPKKALRAAMLKSRFADTILKAQQKTLLEHGDKGDPLKMQLEKERLERIQRE------ 388
++A RAA F +A+Q+ L E KERLE+ E
Sbjct: 641 AEGRERAERAA-----FDRATAEARQRALAE------------AKERLEKACAEARDKSY 769
Query: 389 -ERARIEAQIKTAEAAARARAEEELRQQREKEREAARAAI---EEMKRTV 434
++A EA++K AE AA RA E R++ ++ + RAA E ++R+V
Sbjct: 770 TDKATAEARLK-AERAAVERATAEARERAMEKVKVERAAFGSRERLERSV 916
>AL387088 similar to GP|18266427|gb| cytoplasmic dynein intermediate chain
{Aspergillus nidulans}, partial (2%)
Length = 495
Score = 37.0 bits (84), Expect = 0.018
Identities = 43/159 (27%), Positives = 68/159 (42%), Gaps = 5/159 (3%)
Frame = +1
Query: 356 LKAQQKTLLEHGDKGDPLKMQLEKERLERIQREE--RARIEAQIKTAEAAARARAEEELR 413
+K+ + EH D+ + + +RIQ EE R R E + AE R R EE +
Sbjct: 25 MKSLEMMRQEHRDQQERAETMFN----QRIQDEERRRERDEEARREAENTYRRRIEESEK 192
Query: 414 QQREKEREAARAAIEEMKRTVEIEHNLEIVKELETL-SGCTLSYKALGGRNDYKVALETL 472
+ +E ++EM+ E + NL+ K++E L TLS + L K
Sbjct: 193 NLKNRENNETMRLMKEMQDR-EKKANLDFAKQIEDLKKKNTLSQQELEALKKKKGCFSLD 369
Query: 473 DKLQFENP--LEQLGLFIKDELADEDEEVINGTVEEGEI 509
K+Q N +E L + D + V NG +E E+
Sbjct: 370 TKVQLANGKFVEMAELQVGDRIC---SNVRNGELEFSEV 477
>BF636094 similar to GP|15983759|g At1g05910/T20M3_16 {Arabidopsis thaliana},
partial (12%)
Length = 503
Score = 36.6 bits (83), Expect = 0.024
Identities = 18/46 (39%), Positives = 23/46 (49%)
Frame = +1
Query: 111 PDYFAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTYN 156
P+Y +II PMD+ TI + Y F D+ L SNA YN
Sbjct: 193 PNYRSIIQNPMDIATILQHVDNGNYITCAAFLQDIDLIVSNARAYN 330
>BQ137243 similar to GP|2624970|gb|A proline-rich protein 9-1 {Mus musculus},
partial (13%)
Length = 1145
Score = 35.0 bits (79), Expect = 0.069
Identities = 19/58 (32%), Positives = 31/58 (52%)
Frame = +1
Query: 367 GDKGDPLKMQLEKERLERIQREERARIEAQIKTAEAAARARAEEELRQQREKEREAAR 424
G KG + Q KER ++ +REE AR +A T + A R E+ + R ++ ++ R
Sbjct: 277 GTKGQIGEKQTRKERKKKKEREENARADATTTTNQHAQRQTHEDSGERARRRQTKSTR 450
>TC78343 weakly similar to GP|15724195|gb|AAL06489.1 AT5g49210/K21P3_8
{Arabidopsis thaliana}, partial (78%)
Length = 903
Score = 34.7 bits (78), Expect = 0.090
Identities = 26/91 (28%), Positives = 48/91 (52%), Gaps = 8/91 (8%)
Frame = +3
Query: 338 PKKA-LRAAMLKSRFADTILKAQQKTLLEHGDKGDPLKMQLEKERLE-------RIQREE 389
PK+A + A L+ +A + + +++ ++E M+LEK+R + R+ EE
Sbjct: 204 PKEAEAQLAQLRREYAKQVKEVRKEYIVEVE------AMRLEKQRKDEARKEALRVANEE 365
Query: 390 RARIEAQIKTAEAAARARAEEELRQQREKER 420
R +++AQ K A R A+++ R+ KER
Sbjct: 366 RKKLKAQEKELRAQERNIAQQQFRETLLKER 458
>TC88509 similar to PIR|T01826|T01826 microfibril-associated protein homolog
T15F16.8 - Arabidopsis thaliana, partial (83%)
Length = 1306
Score = 34.3 bits (77), Expect = 0.12
Identities = 19/65 (29%), Positives = 34/65 (52%)
Frame = +1
Query: 383 ERIQREERARIEAQIKTAEAAARARAEEELRQQREKEREAARAAIEEMKRTVEIEHNLEI 442
ER ER R+EA+ + E A + R EE + E + +EE+++ EI+ N+E+
Sbjct: 622 ERDTIAERERLEAEEEALEEARKRRMEE-------RRNETRQIVVEEIRKDQEIQKNIEL 780
Query: 443 VKELE 447
++
Sbjct: 781 EANID 795
>TC77981 similar to PIR|T12970|T12970 hypothetical protein T6H20.190 -
Arabidopsis thaliana, partial (39%)
Length = 1191
Score = 33.5 bits (75), Expect = 0.20
Identities = 27/107 (25%), Positives = 51/107 (47%), Gaps = 14/107 (13%)
Frame = +2
Query: 340 KALRAAMLKSRFADTILKAQQKTLLEHGDKGDPLKMQLE--------------KERLERI 385
K+ A+++ F++T + A+ K + + D PL+ E E LE+
Sbjct: 350 KSRIASLVADVFSNTQV-AENKVVQVYSDPNAPLRPVDELFSTIPEDGRRKAYAEILEKA 526
Query: 386 QREERARIEAQIKTAEAAARARAEEELRQQREKEREAARAAIEEMKR 432
+ EE AR+EA+ AA + EEE + ++E +A+ E+ ++
Sbjct: 527 KAEEEARVEAEKAREAAATTKKLEEEALKLSKQEAQASNLVKEDQEK 667
>TC83303 weakly similar to GP|20503004|gb|AAM22713.1 Hypothetical protein
{Oryza sativa (japonica cultivar-group)}, partial (14%)
Length = 988
Score = 33.1 bits (74), Expect = 0.26
Identities = 19/56 (33%), Positives = 29/56 (50%), Gaps = 4/56 (7%)
Frame = +3
Query: 138 VEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQI----FDSKWKDMDKKWKCEDE 189
+E+F DV L SNAM YN P H A+ + +I F++ +D D +D+
Sbjct: 9 LEQFENDVFLVCSNAMLYNSPDTIYHRQARAMQEIARKDFENLRQDSDDDDDXDDD 176
>BQ150771 GP|4028979|emb glutamate permease {synthetic construct}, partial
(3%)
Length = 1234
Score = 32.7 bits (73), Expect = 0.34
Identities = 25/89 (28%), Positives = 45/89 (50%), Gaps = 1/89 (1%)
Frame = +2
Query: 352 ADTILKAQQKTLLEHGDKGDPLKMQLEKERLERIQREERARIEAQIKTAEAAARARAEEE 411
A T+ +Q+++ E DP + + E+ R EE+ R+ + T + A +++AE
Sbjct: 812 APTMTTSQRESAQETAR--DPTRERREESARSR---EEKKRMRKREDTEDTAKQSQAETR 976
Query: 412 LRQQREKEREAARA-AIEEMKRTVEIEHN 439
R+ RE ERE A +++ E+ HN
Sbjct: 977 RRR*RETERERRHARGKRALRQLKEVRHN 1063
>TC81195 similar to GP|15810597|gb|AAL07186.1 unknown protein {Arabidopsis
thaliana}, partial (50%)
Length = 788
Score = 32.3 bits (72), Expect = 0.44
Identities = 21/84 (25%), Positives = 39/84 (46%)
Frame = +3
Query: 353 DTILKAQQKTLLEHGDKGDPLKMQLEKERLERIQREERARIEAQIKTAEAAARARAEEEL 412
D + + ++ G+K DPLK+ LER+QR+ ++E + + EE+
Sbjct: 363 DVVTDCKSHKVIVKGEKADPLKV------LERVQRKSHRQVELLSPIPKPPSE---EEKQ 515
Query: 413 RQQREKEREAARAAIEEMKRTVEI 436
++EK + +EE K + I
Sbjct: 516 IDEKEKPKPEEEKKVEEPKVIIVI 587
>TC83568 weakly similar to GP|16604681|gb|AAL24133.1 putative kinase
{Arabidopsis thaliana}, partial (6%)
Length = 417
Score = 32.0 bits (71), Expect = 0.58
Identities = 23/38 (60%), Positives = 25/38 (65%), Gaps = 7/38 (18%)
Frame = +3
Query: 479 NPLEQLGLFIKDELADEDEE-----VIN--GTVEEGEI 509
NPLEQLGL+IK E DE+EE V N VEEGEI
Sbjct: 57 NPLEQLGLYIKVE--DEEEEGDPLCVPNPVNDVEEGEI 164
>TC77422 similar to PIR|T04704|T04704 hypothetical protein F4B14.210 -
Arabidopsis thaliana, partial (9%)
Length = 1448
Score = 31.2 bits (69), Expect = 0.99
Identities = 15/54 (27%), Positives = 33/54 (60%)
Frame = +3
Query: 368 DKGDPLKMQLEKERLERIQREERARIEAQIKTAEAAARARAEEELRQQREKERE 421
D +++ LEK + ER +++++A +A K + R + E++ +++REKE +
Sbjct: 174 DDSIKIRLDLEKLQKEREEKDKKAADKAAKKERKERRREKKEKKEKRRREKEEK 335
>TC87626 similar to GP|20259075|gb|AAM14253.1 unknown protein {Arabidopsis
thaliana}, partial (98%)
Length = 1021
Score = 31.2 bits (69), Expect = 0.99
Identities = 15/37 (40%), Positives = 24/37 (64%)
Frame = +3
Query: 388 EERARIEAQIKTAEAAARARAEEELRQQREKEREAAR 424
EERAR++ K E AA A+ E++ +++ +K AAR
Sbjct: 510 EERARLDELKKQREIAAAAKEEDKKKREEQKAAAAAR 620
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.313 0.130 0.364
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,943,163
Number of Sequences: 36976
Number of extensions: 157076
Number of successful extensions: 706
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 685
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 699
length of query: 510
length of database: 9,014,727
effective HSP length: 100
effective length of query: 410
effective length of database: 5,317,127
effective search space: 2180022070
effective search space used: 2180022070
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0228a.2


