

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0227.11
(425 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
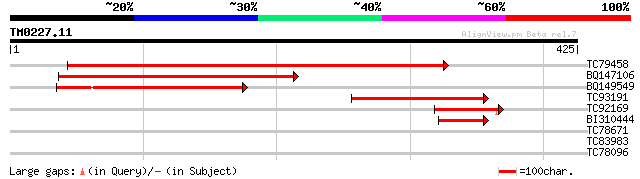
TC79458 weakly similar to PIR|D85436|D85436 MAP3K-like protein k... 424 e-119
BQ147106 similar to GP|15451188|gb Unknown protein {Arabidopsis ... 227 8e-60
BQ149549 similar to PIR|B86270|B86 hypothetical protein AAF81295... 160 7e-40
TC93191 similar to GP|15451188|gb|AAK96865.1 Unknown protein {Ar... 102 2e-22
TC92169 similar to PIR|D85436|D85436 MAP3K-like protein kinase [... 75 5e-14
BI310444 similar to GP|15451188|gb| Unknown protein {Arabidopsis... 56 2e-08
TC78671 similar to GP|21536650|gb|AAM60982.1 sodium-dicarboxylat... 30 2.3
TC83983 similar to GP|19386796|dbj|BAB86175. putative alpha-gluc... 28 5.1
TC78096 weakly similar to GP|17529096|gb|AAL38758.1 unknown prot... 28 8.8
>TC79458 weakly similar to PIR|D85436|D85436 MAP3K-like protein kinase
[imported] - Arabidopsis thaliana, partial (35%)
Length = 1088
Score = 424 bits (1091), Expect = e-119
Identities = 188/286 (65%), Positives = 235/286 (81%)
Frame = +1
Query: 44 NCNSGLHCETCVTNGNVRPRCTRTQPINPTSKVKGLPFNRYSWLTTHNSFALLGQKSMTG 103
+C++GL C+TC NGN RPRC+R NP +KVKGLPFNRYSWLTTHNSFA+ G +S TG
Sbjct: 226 SCDAGLTCQTCPANGNTRPRCSRILSSNPVNKVKGLPFNRYSWLTTHNSFAMAGARSATG 405
Query: 104 SVILSPTNQQDTITDQLNNGVRGLMLDLYDFENDVWLCHSFGGQCYNYTAFQPAINVLKE 163
S+IL+P NQ DTI DQL NGVRG MLD+YDF+NDVWLCHS GG+C+N+++F PA+N L++
Sbjct: 406 SIILAPMNQDDTIADQLKNGVRGFMLDMYDFQNDVWLCHSTGGKCFNFSSFIPAVNALRD 585
Query: 164 IQVFLEANPSEIVTIIIEDYVTSPKGLTKVFDAAGLRKYWFPVSRMPKNGGDWPKVDDMV 223
++ FL+ANPSEI+TI IEDYV +P LTKV A+G+ KY FPV R+PKNG DWP VDDM+
Sbjct: 586 MRSFLDANPSEIITIFIEDYVRAPAALTKVIQASGINKYMFPVGRLPKNGSDWPTVDDMI 765
Query: 224 QKNQRLVVFTSKASKEASEGIAYEWRYLVENQYGNGGMKAGSCPNRAESPSMNTTSRSLV 283
NQR + F+S++SKEA+EGI + W+Y+VENQYG+ GM+ GSCPNR ESP MNT SRSLV
Sbjct: 766 LNNQRFIAFSSRSSKEAAEGIPFTWKYVVENQYGDEGMQPGSCPNRNESPPMNTKSRSLV 945
Query: 284 LVNFFRDLPDVAQSCKDNSAPLLSMVSTCNQAAGKRWPNFIAVDFY 329
L+NFF P+ +Q+C DNSAPLLSM+ TC++AA RWPNFIAVD+Y
Sbjct: 946 LMNFFHSTPNRSQACGDNSAPLLSMLKTCHEAARNRWPNFIAVDYY 1083
>BQ147106 similar to GP|15451188|gb Unknown protein {Arabidopsis thaliana},
partial (36%)
Length = 662
Score = 227 bits (578), Expect = 8e-60
Identities = 108/181 (59%), Positives = 131/181 (71%), Gaps = 1/181 (0%)
Frame = +2
Query: 37 QICVANKNCNSGLHCETCVTNGN-VRPRCTRTQPINPTSKVKGLPFNRYSWLTTHNSFAL 95
+ C A +C +G +C C G R CTR Q TS V GLPFN+YSW+ THNSF++
Sbjct: 119 EACSAATDCGTGYYCGHCPGLGRKTRSVCTRGQATLVTSIVNGLPFNKYSWIMTHNSFSI 298
Query: 96 LGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFENDVWLCHSFGGQCYNYTAFQ 155
+ S+ G L+ NQ+DT+T+QL NGVRGLMLD+YDF+ND+WLCHSF GQCYN+TAFQ
Sbjct: 299 MDAPSLNGVQRLTFYNQEDTVTNQLRNGVRGLMLDMYDFQNDIWLCHSFQGQCYNFTAFQ 478
Query: 156 PAINVLKEIQVFLEANPSEIVTIIIEDYVTSPKGLTKVFDAAGLRKYWFPVSRMPKNGGD 215
PAIN LKE++ FL NP EIVTI+IEDYV +PK L +F AGL KY FPVS MPKNG D
Sbjct: 479 PAINTLKEVEAFLTENPMEIVTIVIEDYVRTPKALINLFINAGLDKYLFPVSDMPKNGED 658
Query: 216 W 216
W
Sbjct: 659 W 661
>BQ149549 similar to PIR|B86270|B86 hypothetical protein AAF81295.1
[imported] - Arabidopsis thaliana, partial (29%)
Length = 635
Score = 160 bits (406), Expect = 7e-40
Identities = 76/144 (52%), Positives = 106/144 (72%), Gaps = 1/144 (0%)
Frame = +3
Query: 36 GQICVANKNCNSGLHCETCVTNGNVRPRCTRTQPINPTSKVKG-LPFNRYSWLTTHNSFA 94
G C ++++C +GL+C C + + RC R+ + + LPFN+Y++LTTHN+FA
Sbjct: 207 GDECNSDEDCGTGLYCFDCSLEFDGK-RCVRSSVTDQFKLINNSLPFNKYAFLTTHNAFA 383
Query: 95 LLGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFENDVWLCHSFGGQCYNYTAF 154
+ G+ S TG ++ TNQ+DTIT QLNNGVR LMLD YDF++DVWLCHSF G C+++TAF
Sbjct: 384 IEGEPSHTGVPRITITNQEDTITQQLNNGVRALMLDTYDFDDDVWLCHSFQGHCHDFTAF 563
Query: 155 QPAINVLKEIQVFLEANPSEIVTI 178
+PAI+ LKEI+ FL ANP+EIVT+
Sbjct: 564 EPAIDTLKEIEAFLSANPNEIVTL 635
>TC93191 similar to GP|15451188|gb|AAK96865.1 Unknown protein {Arabidopsis
thaliana}, partial (23%)
Length = 688
Score = 102 bits (255), Expect = 2e-22
Identities = 50/103 (48%), Positives = 65/103 (62%)
Frame = +2
Query: 257 GNGGMKAGSCPNRAESPSMNTTSRSLVLVNFFRDLPDVAQSCKDNSAPLLSMVSTCNQAA 316
G+ G+ A ES +N+ + SL L N+F P A+SCK+NSAPL MV+TC + A
Sbjct: 8 GDPGVHAVLAHIAKESKPLNSKTASLFLQNYFPTTPVEAESCKENSAPLTDMVNTCYKTA 187
Query: 317 GKRWPNFIAVDFYKRSDGGGAPEAVDVANGHLVCGCGNIATCK 359
G PNFIAV+FY RSDGGG + VD NGH +CGC + C+
Sbjct: 188 GNVLPNFIAVNFYMRSDGGGVFDIVDRINGHALCGCSTVTACQ 316
>TC92169 similar to PIR|D85436|D85436 MAP3K-like protein kinase [imported] -
Arabidopsis thaliana, partial (6%)
Length = 505
Score = 75.1 bits (183), Expect = 5e-14
Identities = 36/59 (61%), Positives = 40/59 (67%), Gaps = 7/59 (11%)
Frame = +2
Query: 319 RWPNFIAVDFYKRSDGGGAPEAVDVANGHLVCGCGNIATCKVRLT-------PITFIPP 370
RWPNFIAVD+Y RSDGGG P+AVD ANG L CGC +IA CK T PI+ PP
Sbjct: 2 RWPNFIAVDYYLRSDGGGVPQAVDAANGRLTCGCDSIAYCKANGTFGSCDVPPISPPPP 178
>BI310444 similar to GP|15451188|gb| Unknown protein {Arabidopsis thaliana},
partial (11%)
Length = 620
Score = 56.2 bits (134), Expect = 2e-08
Identities = 23/38 (60%), Positives = 28/38 (73%)
Frame = +3
Query: 322 NFIAVDFYKRSDGGGAPEAVDVANGHLVCGCGNIATCK 359
NFIAV+FY RSDGGG + VD NGH +CGC +A C+
Sbjct: 3 NFIAVNFYMRSDGGGVFDIVDKMNGHSLCGCSTVAACQ 116
>TC78671 similar to GP|21536650|gb|AAM60982.1 sodium-dicarboxylate
cotransporter-like {Arabidopsis thaliana}, partial
(50%)
Length = 1083
Score = 29.6 bits (65), Expect = 2.3
Identities = 18/54 (33%), Positives = 28/54 (51%), Gaps = 3/54 (5%)
Frame = +1
Query: 47 SGLHCETCVT---NGNVRPRCTRTQPINPTSKVKGLPFNRYSWLTTHNSFALLG 97
+G H ET V+ N N+ + P+ P + + PFN S+ T +N + LLG
Sbjct: 28 TGEHSETSVSDESNNNLNLKA----PLLPLHRTQSNPFNLKSFFTLNNFYVLLG 177
>TC83983 similar to GP|19386796|dbj|BAB86175. putative alpha-glucosidase 1
{Oryza sativa (japonica cultivar-group)}, partial (11%)
Length = 656
Score = 28.5 bits (62), Expect = 5.1
Identities = 17/39 (43%), Positives = 19/39 (48%)
Frame = -3
Query: 309 VSTCNQAAGKRWPNFIAVDFYKRSDGGGAPEAVDVANGH 347
V C AA K P+F V F RSD AP A+GH
Sbjct: 216 VFACGSAAVKFRPSFAGVGFRGRSD--SAPSGCSSASGH 106
>TC78096 weakly similar to GP|17529096|gb|AAL38758.1 unknown protein
{Arabidopsis thaliana}, partial (78%)
Length = 1383
Score = 27.7 bits (60), Expect = 8.8
Identities = 17/54 (31%), Positives = 26/54 (47%)
Frame = -1
Query: 70 INPTSKVKGLPFNRYSWLTTHNSFALLGQKSMTGSVILSPTNQQDTITDQLNNG 123
+ P++ K P SW TH F + + S+TG++I S Q+ T NG
Sbjct: 1182 LTPSA*GKRWPEPESSW**THGRFCAMYRCSVTGTIISSSPCQR*TEPSLFGNG 1021
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.136 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,725,086
Number of Sequences: 36976
Number of extensions: 223854
Number of successful extensions: 1127
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 1124
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1127
length of query: 425
length of database: 9,014,727
effective HSP length: 99
effective length of query: 326
effective length of database: 5,354,103
effective search space: 1745437578
effective search space used: 1745437578
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0227.11


