

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.9
(223 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
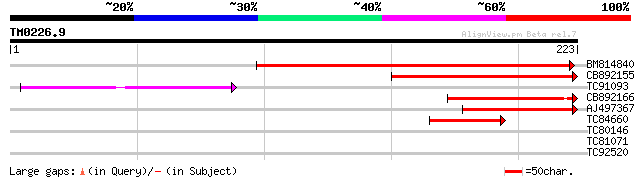
BM814840 weakly similar to GP|15128241|db helicase-like protein ... 144 2e-35
CB892155 similar to PIR|D86481|D86 hypothetical protein AAG28292... 92 2e-19
TC91093 weakly similar to PIR|T01873|T01873 hypothetical protein... 65 3e-11
CB892166 similar to GP|20197614|gb unknown protein {Arabidopsis ... 59 2e-09
AJ497367 similar to GP|14140286|gb putative helicase {Oryza sati... 55 3e-08
TC84660 similar to PIR|D86481|D86481 hypothetical protein AAG282... 41 3e-04
TC80146 37 0.006
TC81071 weakly similar to PIR|T52653|T52653 probable sigma-like ... 30 0.73
TC92520 similar to GP|15010768|gb|AAK74043.1 AT3g06130/F28L1_7 {... 27 6.2
>BM814840 weakly similar to GP|15128241|db helicase-like protein {Oryza
sativa (japonica cultivar-group)}, partial (6%)
Length = 733
Score = 144 bits (364), Expect = 2e-35
Identities = 74/125 (59%), Positives = 94/125 (75%)
Frame = +3
Query: 98 NGTRLIVDELGERFIGATVITGTNVNDKVHILRMDLVLSDSKFPIKFRIRQFPIAICFAM 157
+GTRLI+ LG+ I A VI GT+ + +I RM+L+ S + I F QFP+ + FAM
Sbjct: 27 HGTRLIIVSLGKNVICARVIGGTHAGEVSYIPRMNLIPSGANVSITFERCQFPLVLSFAM 206
Query: 158 TINKSQGQTLSQVGLFLPRPVFTHGQLYVALSRVRSRKGLKLCILDEAGKQQTSTVNVVF 217
TINKSQGQTL+ VGL+LPRPVFTHGQLYVA+SRV+SR GLK+ I DE G +STVNVV+
Sbjct: 207 TINKSQGQTLTSVGLYLPRPVFTHGQLYVAVSRVKSRSGLKILITDENGSPSSSTVNVVY 386
Query: 218 KEVFE 222
+EVF+
Sbjct: 387 QEVFQ 401
>CB892155 similar to PIR|D86481|D86 hypothetical protein AAG28292.1
[imported] - Arabidopsis thaliana, partial (4%)
Length = 572
Score = 91.7 bits (226), Expect = 2e-19
Identities = 43/73 (58%), Positives = 57/73 (77%)
Frame = -2
Query: 151 IAICFAMTINKSQGQTLSQVGLFLPRPVFTHGQLYVALSRVRSRKGLKLCILDEAGKQQT 210
+ + FAMTINKSQGQ+L +G++LP VF+HGQLYVALSRV SR+GLK+ I ++ G+
Sbjct: 370 VEVYFAMTINKSQGQSLKHIGVYLPSSVFSHGQLYVALSRVTSREGLKILISNDDGEDDC 191
Query: 211 STVNVVFKEVFEN 223
T NVV++EVF N
Sbjct: 190 VTSNVVYREVFHN 152
>TC91093 weakly similar to PIR|T01873|T01873 hypothetical protein T24M8.10 -
Arabidopsis thaliana, partial (11%)
Length = 701
Score = 64.7 bits (156), Expect = 3e-11
Identities = 37/85 (43%), Positives = 50/85 (58%)
Frame = +3
Query: 5 QFFQDRAILAPTLEAVEMLNTYMLGMIAAPEHEYLSSDSALRSDEDSEIKVEWFTTEFSN 64
QF Q RAIL T E V+ +N Y+L +I E S++ RS+ + + EF
Sbjct: 453 QFLQSRAILTSTDEVVDQINDYVLKLIPGEERVIYSAN---RSEVNDVQAFDAIPPEFLQ 623
Query: 65 DTKSSGMPNHQLLLKVGTPIMLLRN 89
K+S +PNH+L LKVGTPIMLLR+
Sbjct: 624 SLKTSDLPNHKLTLKVGTPIMLLRD 698
>CB892166 similar to GP|20197614|gb unknown protein {Arabidopsis thaliana},
partial (3%)
Length = 748
Score = 58.5 bits (140), Expect = 2e-09
Identities = 30/51 (58%), Positives = 38/51 (73%)
Frame = -1
Query: 173 FLPRPVFTHGQLYVALSRVRSRKGLKLCILDEAGKQQTSTVNVVFKEVFEN 223
F R VF+HGQLYVA+SRV SR GLK+ ++DE G +T NVV+K VF+N
Sbjct: 289 FYDREVFSHGQLYVAISRVSSRNGLKILMIDENGDCIDNTTNVVYK-VFQN 140
>AJ497367 similar to GP|14140286|gb putative helicase {Oryza sativa (japonica
cultivar-group)}, partial (1%)
Length = 543
Score = 54.7 bits (130), Expect = 3e-08
Identities = 26/45 (57%), Positives = 34/45 (74%)
Frame = +1
Query: 179 FTHGQLYVALSRVRSRKGLKLCILDEAGKQQTSTVNVVFKEVFEN 223
F++G+LYVA+SRV SRKGLK+ + E G +T NVV+KEVF N
Sbjct: 19 FSNGKLYVAVSRVTSRKGLKILLAHEDGNCMNTTSNVVYKEVFRN 153
>TC84660 similar to PIR|D86481|D86481 hypothetical protein AAG28292.1
[imported] - Arabidopsis thaliana, partial (1%)
Length = 1009
Score = 41.2 bits (95), Expect = 3e-04
Identities = 18/30 (60%), Positives = 25/30 (83%)
Frame = +2
Query: 166 TLSQVGLFLPRPVFTHGQLYVALSRVRSRK 195
+++ G++LP+P+F HG LYVALSRV SRK
Sbjct: 68 SMATKGMYLPQPIF*HG*LYVALSRVTSRK 157
>TC80146
Length = 476
Score = 37.0 bits (84), Expect = 0.006
Identities = 16/35 (45%), Positives = 26/35 (73%)
Frame = -3
Query: 151 IAICFAMTINKSQGQTLSQVGLFLPRPVFTHGQLY 185
I I + TINKS+ Q+LS + ++L RP+F+H ++Y
Sbjct: 471 IQIPYFKTINKSR*QSLSYMKIYLSRPIFSHEEMY 367
>TC81071 weakly similar to PIR|T52653|T52653 probable sigma-like transcription
initiation factor [imported] - Arabidopsis thaliana
(fragment), partial (64%)
Length = 1856
Score = 30.0 bits (66), Expect = 0.73
Identities = 17/63 (26%), Positives = 35/63 (54%), Gaps = 5/63 (7%)
Frame = -1
Query: 126 VHILRMDLVLSDSKF---PIKFRIRQ--FPIAICFAMTINKSQGQTLSQVGLFLPRPVFT 180
+H L+ D+ +SD+++ + + I Q + + IC + NK+ + + ++ L L P+ T
Sbjct: 1793 IHNLKYDIYISDNQYNCSTLVYAIDQLYY*VYICIKLLTNKALLELVQRLALQLSDPLLT 1614
Query: 181 HGQ 183
H Q
Sbjct: 1613 HTQ 1605
>TC92520 similar to GP|15010768|gb|AAK74043.1 AT3g06130/F28L1_7 {Arabidopsis
thaliana}, partial (13%)
Length = 756
Score = 26.9 bits (58), Expect = 6.2
Identities = 13/37 (35%), Positives = 20/37 (53%)
Frame = -1
Query: 74 HQLLLKVGTPIMLLRNIDQAASLCNGTRLIVDELGER 110
H +L + TP + I QAA LC R + ++G+R
Sbjct: 270 HHNILMIHTPTFSMMKILQAAVLCEEMRC*IRDVGKR 160
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.137 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,929,907
Number of Sequences: 36976
Number of extensions: 71216
Number of successful extensions: 264
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 264
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 264
length of query: 223
length of database: 9,014,727
effective HSP length: 93
effective length of query: 130
effective length of database: 5,575,959
effective search space: 724874670
effective search space used: 724874670
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0226.9


