

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.8
(518 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
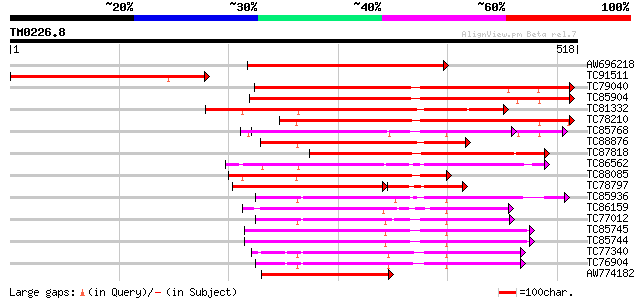
Score E
Sequences producing significant alignments: (bits) Value
AW696218 homologue to PIR|B96835|B96 CAD ATPase (AAA1) 35570-33... 365 e-101
TC91511 similar to GP|19909896|dbj|BAB87822. katanin {Arabidopsi... 280 1e-75
TC79040 homologue to PIR|F84674|F84674 probable AAA-type ATPase ... 258 4e-69
TC85904 similar to PIR|F84674|F84674 probable AAA-type ATPase [i... 258 5e-69
TC81332 similar to GP|15810167|gb|AAL06985.1 At1g64110/F22C12_22... 214 5e-56
TC78210 similar to GP|6056413|gb|AAF02877.1| Unknown protein {Ar... 202 2e-52
TC85768 homologue to SP|P54774|CC48_SOYBN Cell division cycle pr... 194 7e-50
TC88876 similar to PIR|G96537|G96537 hypothetical protein F2J10.... 185 4e-47
TC87818 similar to GP|20197082|gb|AAC26698.2 putative katanin {A... 184 5e-47
TC86562 similar to GP|21553404|gb|AAM62497.1 26S proteasome regu... 170 1e-42
TC88085 similar to GP|20259341|gb|AAM13995.1 unknown protein {Ar... 170 1e-42
TC78797 similar to GP|15810167|gb|AAL06985.1 At1g64110/F22C12_22... 126 7e-41
TC85936 homologue to GP|13537115|dbj|BAB40755. AtSUG1 {Arabidops... 162 4e-40
TC86159 homologue to GP|21387177|gb|AAM47992.1 26S proteasome AA... 159 3e-39
TC77012 homologue to SP|P54776|PRSA_LYCES 26S protease regulator... 152 2e-37
TC85745 homologue to GP|17297987|dbj|BAB78491. 26S proteasome re... 152 2e-37
TC85744 homologue to PIR|T08959|T08959 proteinase homolog F19B15... 152 2e-37
TC77340 SP|Q9BAE0|FTSH_MEDSA Cell division protein ftsH homolog ... 150 1e-36
TC76904 homologue to GP|4325041|gb|AAD17230.1| FtsH-like protein... 145 4e-35
AW774182 homologue to PIR|F84674|F84 probable AAA-type ATPase [i... 144 6e-35
>AW696218 homologue to PIR|B96835|B96 CAD ATPase (AAA1) 35570-33019
[imported] - Arabidopsis thaliana, partial (34%)
Length = 556
Score = 365 bits (937), Expect = e-101
Identities = 180/184 (97%), Positives = 184/184 (99%)
Frame = +3
Query: 218 EMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIRRPWKGVLMFGPPG 277
EMLERDVLETSPGVRW+DVAGLTEAKRLLEEAVVLPLWMPEYFQGIRRPWKGVLMFGPPG
Sbjct: 3 EMLERDVLETSPGVRWDDVAGLTEAKRLLEEAVVLPLWMPEYFQGIRRPWKGVLMFGPPG 182
Query: 278 TGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDS 337
TGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDS
Sbjct: 183 TGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDS 362
Query: 338 LCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRR 397
LCN+RGASGEHESSRRVKSELLVQVDGVSN+ATNEDGSRK+VMVLAATNFPWDIDEALRR
Sbjct: 363 LCNARGASGEHESSRRVKSELLVQVDGVSNSATNEDGSRKLVMVLAATNFPWDIDEALRR 542
Query: 398 RLEK 401
RLEK
Sbjct: 543 RLEK 554
>TC91511 similar to GP|19909896|dbj|BAB87822. katanin {Arabidopsis
thaliana}, partial (23%)
Length = 595
Score = 280 bits (716), Expect = 1e-75
Identities = 140/186 (75%), Positives = 159/186 (85%), Gaps = 4/186 (2%)
Frame = +1
Query: 1 MVGSSLGGLQEHLKLAREYALEGQYDTSIIFFDGAVAQINKHLNSLDDPLLRSKWMNVKK 60
+ G SL GLQ+H+KLAR+YALEG YDTSIIFFDGA+AQIN+HLNS+DDPL+R+KWMNVKK
Sbjct: 31 VAGGSLAGLQDHVKLARDYALEGLYDTSIIFFDGAIAQINRHLNSVDDPLIRAKWMNVKK 210
Query: 61 ALSEETEVVKQLDAERSAFKNNPIGRRPPSPPISTKSSFVFQPLDEYPTSSSGPMDDPDV 120
ALSEETEVVKQLDA++ AFK+NPIGRRPPSPPISTKSSFVFQPLDEYPTSS+ DDPDV
Sbjct: 211 ALSEETEVVKQLDADKRAFKDNPIGRRPPSPPISTKSSFVFQPLDEYPTSSNSSFDDPDV 390
Query: 121 WRPPSRDTSRRPTRPGQMSTRKS--DGAWARGATPR-GGATGRGAKA-GATSKVNTGTRA 176
WRPPSRDTSRRP R GQ+ TRKS D WARGAT + TGRGAKA GATS+ N+G
Sbjct: 391 WRPPSRDTSRRPGRGGQVGTRKSNQDSKWARGATTKTTSTTGRGAKAGGATSRTNSGGPX 570
Query: 177 STTGKK 182
+GK+
Sbjct: 571 INSGKE 588
>TC79040 homologue to PIR|F84674|F84674 probable AAA-type ATPase [imported]
- Arabidopsis thaliana, partial (74%)
Length = 1193
Score = 258 bits (659), Expect = 4e-69
Identities = 137/318 (43%), Positives = 201/318 (63%), Gaps = 25/318 (7%)
Frame = +3
Query: 224 VLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIRRPWKGVLMFGPPGTGKTLL 283
++ P V+W DVAGL AK+ L+EAV+LP+ P++F G RRPW+ L++GPPGTGK+ L
Sbjct: 30 IIREKPNVKWNDVAGLESAKQALQEAVILPVKFPQFFTGKRRPWRAFLLYGPPGTGKSYL 209
Query: 284 AKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRG 343
AKAVATE +TFF++SS+ L SKW GESE++V LF +AR APS IF+DEIDSLC RG
Sbjct: 210 AKAVATEADSTFFSISSSDLVSKWMGESEKLVSNLFQMARESAPSIIFVDEIDSLCGQRG 389
Query: 344 ASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRRRLEKRI 403
E E+SRR+K+ELLVQ+ GV N + + V+VLAATN P+ +D+A+RRR +KRI
Sbjct: 390 EGNESEASRRIKTELLVQMQGVGN-------NDQKVLVLAATNTPYALDQAIRRRFDKRI 548
Query: 404 YIPLPNFESRKELIRINL-KTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDA-------- 454
YIPLP+ ++R+ + +++L T + + + +A RT+G+SG D++ +D
Sbjct: 549 YIPLPDLKARQHMFKVHLGDTPHNLTEKDYEYLASRTEGFSGSDISVCVKDVLFEPVRKT 728
Query: 455 --------SLNGMRRKIAGKTRDEIKNMSKDDISK--------DPVAMCDFEEALVKVQR 498
S GM K + ++ D +K P+ DFE+ L + +
Sbjct: 729 QDAMFFFKSPEGMWIPCGPKQQGAVQTTMTDLATKGLASKILPPPITRTDFEKVLARQRP 908
Query: 499 SVSQADIERHEKWFHEFG 516
+VS++D+E HE++ EFG
Sbjct: 909 TVSKSDLEVHERFTKEFG 962
>TC85904 similar to PIR|F84674|F84674 probable AAA-type ATPase [imported] -
Arabidopsis thaliana, complete
Length = 1793
Score = 258 bits (658), Expect = 5e-69
Identities = 133/322 (41%), Positives = 206/322 (63%), Gaps = 25/322 (7%)
Frame = +2
Query: 220 LERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIRRPWKGVLMFGPPGTG 279
L ++ P V+W DVAGL AK+ L+EAV+LP+ P++F G RRPW+ L++GPPGTG
Sbjct: 536 LNSAIVREKPNVKWNDVAGLESAKQSLQEAVILPVKFPQFFTGKRRPWRAFLLYGPPGTG 715
Query: 280 KTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLC 339
K+ LAKAVATE +TFF+VSS+ L SKW GESE++V LF++AR APS IF+DEIDSLC
Sbjct: 716 KSYLAKAVATEADSTFFSVSSSDLVSKWMGESEKLVSNLFEMARESAPSIIFVDEIDSLC 895
Query: 340 NSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRRRL 399
+RG E E+SRR+K+ELLVQ+ GV + + + V+VLAATN P+ +D+A+RRR
Sbjct: 896 GTRGEGNESEASRRIKTELLVQMQGVGH-------NDQKVLVLAATNTPYALDQAIRRRF 1054
Query: 400 EKRIYIPLPNFESRKELIRINL-KTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASLNG 458
+KRIYIPLP+ ++R+ + +++L T + + + +AR+T+G+SG D+ +D
Sbjct: 1055DKRIYIPLPDLKARQHMFKVHLGDTPHNLTESDFEHLARKTEGFSGSDIAVCVKDVLFEP 1234
Query: 459 MRRK---------------IAGKTRDEIKNMSKDDISKD---------PVAMCDFEEALV 494
+R+ G+ + ++ D++ P++ DF++ L
Sbjct: 1235VRKTQDAMFFFKSPEGMWIPCGQKQQNAVQVTMQDLATQGLASKILPPPISRIDFDKVLA 1414
Query: 495 KVQRSVSQADIERHEKWFHEFG 516
+ + +VS++D++ HE++ EFG
Sbjct: 1415RQRPTVSKSDLDVHERFTKEFG 1480
>TC81332 similar to GP|15810167|gb|AAL06985.1 At1g64110/F22C12_22
{Arabidopsis thaliana}, partial (39%)
Length = 1514
Score = 214 bits (546), Expect = 5e-56
Identities = 115/289 (39%), Positives = 181/289 (61%), Gaps = 13/289 (4%)
Frame = +1
Query: 180 GKKSGVSGKAGKTDSVISDAEESKSKKSQYEG----------PDPELAEMLERDVLETSP 229
G+ S K K +S D+ + K K+S+ +G PD E + ++++ +
Sbjct: 643 GESSEKDNKKTKKESKRDDSRKEKPKESKKDGDIKASAKSDSPDNAFEECIRQELIPANE 822
Query: 230 -GVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQG--IRRPWKGVLMFGPPGTGKTLLAKA 286
V + D+ L + K L+EAV+LPL P+ F+G + +P KGVL+FGPPGTGKT+LAKA
Sbjct: 823 IKVTFSDIGALDDVKESLQEAVMLPLRRPDLFKGDGVLKPCKGVLLFGPPGTGKTMLAKA 1002
Query: 287 VATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRGASG 346
+A E G +F NVS +T+ S W+G+SE+ VR LF LA AP+ IFIDE+DS+ R ++
Sbjct: 1003IANEAGASFINVSPSTITSMWQGQSEKNVRALFSLAAKVAPTIIFIDEVDSMLGQRSSTR 1182
Query: 347 EHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRRRLEKRIYIP 406
EH S RRVK+E + + DG+ + + + VLAATN P+D+DEA+ RR ++RI +
Sbjct: 1183EHSSMRRVKNEFMSRWDGLLSKPDEK------ITVLAATNMPFDLDEAIIRRFQRRIMVG 1344
Query: 407 LPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDAS 455
LP+ ++R+ +++ + E + ++N +E++ T+GYSG DL N+C A+
Sbjct: 1345LPSADNRETILK-TILAKEKSENMNFEELSTMTEGYSGSDLKNLCMTAA 1488
>TC78210 similar to GP|6056413|gb|AAF02877.1| Unknown protein {Arabidopsis
thaliana}, partial (23%)
Length = 1254
Score = 202 bits (515), Expect = 2e-52
Identities = 112/282 (39%), Positives = 174/282 (60%), Gaps = 12/282 (4%)
Frame = +3
Query: 247 EEAVVLPLWMPEYF--QGIRRPWKGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLA 304
+E V+LPL PE F + +P KG+L+FGPPGTGKT+LAKAVATE G F N+S +++
Sbjct: 3 KELVMLPLKRPELFCKGQLTKPCKGILLFGPPGTGKTMLAKAVATEAGANFINISMSSIT 182
Query: 305 SKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDG 364
SKW GE E+ V+ +F LA APS IF+DE+DS+ R GEHE+ R++K+E +V DG
Sbjct: 183 SKWFGEGEKYVKAVFSLASKIAPSVIFVDEVDSMLGRRENPGEHEAMRKMKNEFMVNWDG 362
Query: 365 VSNTATNEDGSRKIVMVLAATNFPWDIDEALRRRLEKRIYIPLPNFESRKELIRINLKTV 424
+ ++ ++VLAATN P+D+DEA+ RRL +R+ + LP+ +R +++R+ L
Sbjct: 363 LRTK------EKERILVLAATNRPFDLDEAVIRRLPRRLMVDLPDAPNRGKILRVILAKE 524
Query: 425 EVAPDVNIDEVARRTDGYSGDDLTNVCRDASLNGMRRKIAGKTRDEIKNMSKDDISKD-- 482
++A DV+++ +A TDGYSG DL N+C A+ +R + + +D+ ++++ +
Sbjct: 525 DLAADVDLEAIANMTDGYSGSDLKNLCVTAAHCPIREILEKEKKDKSLALAENKPEPELC 704
Query: 483 ------PVAMCDFEEALVKVQRSVSQADIERHE--KWFHEFG 516
P+ M DF A +V SVS +E +W +G
Sbjct: 705 SSADIRPLKMEDFRYAHEQVCASVSSESTNMNELQQWNDLYG 830
>TC85768 homologue to SP|P54774|CC48_SOYBN Cell division cycle protein 48
homolog (Valosin containing protein homolog) (VCP).
[Soybean], partial (95%)
Length = 2785
Score = 194 bits (493), Expect = 7e-50
Identities = 115/300 (38%), Positives = 172/300 (57%), Gaps = 12/300 (4%)
Frame = +2
Query: 222 RDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGI-RRPWKGVLMFGPPGTGK 280
R+ + P W+D+ GL KR L+E V P+ PE F+ P KGVL +GPPG GK
Sbjct: 1499 RETVVEVPNCSWDDIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPGCGK 1678
Query: 281 TLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCN 340
TLLAKA+A EC F ++ L + W GESE VR +FD AR AP +F DE+DS+
Sbjct: 1679 TLLAKAIANECQANFISIKGPELLTMWFGESEANVREIFDKARGSAPCVLFFDELDSIAT 1858
Query: 341 SRGAS--GEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRR- 397
RG+S ++ RV ++LL ++DG+S ++K V ++ ATN P ID AL R
Sbjct: 1859 QRGSSVGDAGGAADRVLNQLLTEMDGMS--------AKKTVFIIGATNRPDIIDPALLRP 2014
Query: 398 -RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASL 456
RL++ IYIPLP+ +SR ++ + L+ ++ DV+I +A+ T G+SG D+T +C+ A
Sbjct: 2015 GRLDQLIYIPLPDEDSRHQIFKACLRKSPISKDVDIRALAKYTQGFSGADITEICQRACK 2194
Query: 457 NGMRRKI-----AGKTRDEIKNMSKDDISKD--PVAMCDFEEALVKVQRSVSQADIERHE 509
+R I + R E ++DI + + FEE++ +RSVS ADI +++
Sbjct: 2195 YAIRENIEKDIEKERKRSENPEAMEEDIEDEVAEIKAAHFEESMKYARRSVSDADIRKYQ 2374
Score = 168 bits (426), Expect = 4e-42
Identities = 103/258 (39%), Positives = 156/258 (59%), Gaps = 6/258 (2%)
Frame = +2
Query: 212 PDPEL---AEMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIR-RPW 267
PD E+ E ++R+ V ++DV G+ + + E V LPL P+ F+ I +P
Sbjct: 641 PDTEIFCEGEPIKREDENRLDEVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKPP 820
Query: 268 KGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAP 327
KG+L++GPPG+GKTL+A+AVA E G FF ++ + SK GESE +R F+ A AP
Sbjct: 821 KGILLYGPPGSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAEKNAP 1000
Query: 328 STIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNF 387
S IFIDEIDS+ R + E RR+ S+LL +DG+ SR V+V+ ATN
Sbjct: 1001SIIFIDEIDSIAPKREKT-HGEVERRIVSQLLTLMDGLK--------SRAHVIVMGATNR 1153
Query: 388 PWDIDEALRR--RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGD 445
P ID ALRR R ++ I I +P+ R E++RI+ K +++A DV+++++++ T GY G
Sbjct: 1154PNSIDPALRRFGRFDREIDIGVPDEVGRLEVLRIHTKNMKLAEDVDLEKISKETHGYVGA 1333
Query: 446 DLTNVCRDASLNGMRRKI 463
DL +C +A+L +R K+
Sbjct: 1334DLAALCTEAALQCIREKM 1387
>TC88876 similar to PIR|G96537|G96537 hypothetical protein F2J10.1 [imported]
- Arabidopsis thaliana, partial (72%)
Length = 1572
Score = 185 bits (469), Expect = 4e-47
Identities = 88/194 (45%), Positives = 135/194 (69%), Gaps = 2/194 (1%)
Frame = +2
Query: 230 GVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQ--GIRRPWKGVLMFGPPGTGKTLLAKAV 287
GV+++D+ L + K+ L+E V+LP+ PE F + RP KG+L+FGPPGTGKTLLAKA+
Sbjct: 908 GVKFDDIGALEDVKKALQELVILPMRRPELFSHGNLLRPCKGILLFGPPGTGKTLLAKAL 1087
Query: 288 ATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRGASGE 347
ATE G F +V+ +TL SKW G++E++ + LF A AP IF+DE+DSL +RG + E
Sbjct: 1088 ATEAGANFISVTGSTLTSKWFGDAEKLTKALFSFASKLAPVIIFVDEVDSLLGARGGAHE 1267
Query: 348 HESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRRRLEKRIYIPL 407
HE++RR+++E + DG+ + +++L ATN P+D+D+A+ RRL +RIY+ L
Sbjct: 1268 HEATRRMRNEFMAAWDGLRSKENQR------ILILGATNRPFDLDDAVIRRLPRRIYVDL 1429
Query: 408 PNFESRKELIRINL 421
P+ E+R +++RI L
Sbjct: 1430 PDVENRMKILRIFL 1471
Score = 34.3 bits (77), Expect = 0.12
Identities = 17/41 (41%), Positives = 25/41 (60%)
Frame = +1
Query: 411 ESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVC 451
ES ++ I+ K + PD D++A T+GYSG DL N+C
Sbjct: 1441 ESNEDSKNISCKR-NLNPDFQYDKLASLTEGYSGSDLKNLC 1560
>TC87818 similar to GP|20197082|gb|AAC26698.2 putative katanin {Arabidopsis
thaliana}, partial (62%)
Length = 984
Score = 184 bits (468), Expect = 5e-47
Identities = 98/221 (44%), Positives = 149/221 (67%), Gaps = 2/221 (0%)
Frame = +2
Query: 275 PPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDE 334
P G + +LAKAVATEC TTFFN+S++++ SKWRG+SE++V+ LF+LAR +AP+TIF+DE
Sbjct: 5 PQGLERPMLAKAVATECNTTFFNISASSMFSKWRGDSEKLVKVLFELARHHAPATIFLDE 184
Query: 335 IDSLCNSRG-ASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDE 393
ID++ + RG EHE+SRR+K+ELL+Q+DG++ T ++V VLAATN PW++D
Sbjct: 185 IDAIISQRGEGRSEHEASRRLKTELLIQMDGLART-------DELVFVLAATNLPWELDA 343
Query: 394 ALRRRLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRD 453
A+ RRLEKRI +PLP E+R+ + L + D + RT+GYSG D+ +C++
Sbjct: 344 AMLRRLEKRILVPLPEPEARRAMFEELLPLQPDEEPMPYDLLVDRTEGYSGSDIRLLCKE 523
Query: 454 ASLNGMRRKIAGKTRDEIKNMSKDDISK-DPVAMCDFEEAL 493
++ +RR + + E + ++++ K PV D E AL
Sbjct: 524 TAMQPLRR-LMTQLEQEPDVVPEEELPKVGPVVPEDVEAAL 643
>TC86562 similar to GP|21553404|gb|AAM62497.1 26S proteasome regulatory
particle chain RPT6-like protein {Arabidopsis thaliana},
partial (77%)
Length = 1564
Score = 170 bits (431), Expect = 1e-42
Identities = 108/313 (34%), Positives = 169/313 (53%), Gaps = 17/313 (5%)
Frame = +1
Query: 198 DAEESKSKKSQYEGPDPELAEMLERDVLETSP---------------GVRWEDVAGLTEA 242
D SKK+ + E+A+ L R +++T+ V ++ + GL
Sbjct: 259 DPNRESSKKALEQ--KKEIAKRLGRPLIQTNSYEDVIACDVINPDHIDVEFDSIGGLETI 432
Query: 243 KRLLEEAVVLPLWMPEYFQG--IRRPWKGVLMFGPPGTGKTLLAKAVATECGTTFFNVSS 300
K+ L E V+LPL P+ F + P KGVL++GPPGTGKT+LAKA+A E G F NV
Sbjct: 433 KQTLFELVILPLQRPDLFSHGKLLGPQKGVLLYGPPGTGKTMLAKAIAKESGAVFINVRI 612
Query: 301 ATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRGASGEHESSRRVKSELLV 360
+ L SKW G+++++V +F LA PS IFIDE+DS R S +HE+ +K+E +
Sbjct: 613 SNLMSKWFGDAQKLVAAVFSLAHKLQPSIIFIDEVDSFLGQR-RSSDHEAVLNMKTEFMA 789
Query: 361 QVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRRRLEKRIYIPLPNFESRKELIRIN 420
DG AT++ VMVLAATN P ++DEA+ RRL + I P+ + R +++++
Sbjct: 790 LWDGF---ATDQSAR---VMVLAATNRPSELDEAILRRLPQAFEIGYPDRKERADILKVI 951
Query: 421 LKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASLNGMRRKIAGKTRDEIKNMSKDDIS 480
LK +V +++ +A GY+G DL ++C+ A+ +R + + E
Sbjct: 952 LKGEKVEDNIDFSYIAGLCKGYTGSDLFDLCKKAAYFPIREILHNEKNGE------QSCE 1113
Query: 481 KDPVAMCDFEEAL 493
P++ D E+AL
Sbjct: 1114PRPLSQADLEKAL 1152
>TC88085 similar to GP|20259341|gb|AAM13995.1 unknown protein {Arabidopsis
thaliana}, partial (29%)
Length = 1261
Score = 170 bits (431), Expect = 1e-42
Identities = 95/208 (45%), Positives = 132/208 (62%), Gaps = 5/208 (2%)
Frame = +1
Query: 201 ESKSKKSQYEG--PDPELAEMLERDVLE-TSPGVRWEDVAGLTEAKRLLEEAVVLPLWMP 257
E+KS K + + E + L DV+ T GV + D+ L K L+E V+LPL P
Sbjct: 655 ENKSVKKSLKDVVTENEFEKKLLGDVIPPTDIGVSFNDIGALENVKDTLKELVMLPLQRP 834
Query: 258 EYF--QGIRRPWKGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMV 315
E F + +P KG+L+FGPPGTGKT+LAKAVATE G F N+S +++ SKW GE E+ V
Sbjct: 835 ELFCKGQLTKPCKGILLFGPPGTGKTMLAKAVATEAGANFINISMSSITSKWFGEGEKYV 1014
Query: 316 RCLFDLARAYAPSTIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGS 375
+ +F LA APS IF+DE+DS+ R GEHE+ R++K+E +V DG+
Sbjct: 1015KAVFSLASKIAPSVIFVDEVDSMLGRRENPGEHEAMRKMKNEFMVNWDGLRTK------D 1176
Query: 376 RKIVMVLAATNFPWDIDEALRRRLEKRI 403
R+ V+VLAATN P+D+DEA+ RRL +R+
Sbjct: 1177RERVLVLAATNRPFDLDEAVIRRLPRRL 1260
>TC78797 similar to GP|15810167|gb|AAL06985.1 At1g64110/F22C12_22 {Arabidopsis
thaliana}, partial (74%)
Length = 2270
Score = 126 bits (316), Expect(2) = 7e-41
Identities = 64/144 (44%), Positives = 92/144 (63%), Gaps = 2/144 (1%)
Frame = +1
Query: 204 SKKSQYEGPDPELAEMLERDVLETSP-GVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQG 262
+ K+ PD E + + +V+ + V + D+ L E K L+E V+LPL P+ F G
Sbjct: 1621 ASKASEVPPDNEFEKRIRPEVIPANEIDVTFSDIGALEETKESLQELVMLPLRRPDLFTG 1800
Query: 263 -IRRPWKGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDL 321
+ +P +G+L+FGPPGTGKT+LAKA+A E G +F NVS +T+ SKW GE E+ VR LF L
Sbjct: 1801 GLLKPCRGILLFGPPGTGKTMLAKAIAKEAGASFINVSMSTITSKWFGEDEKNVRALFTL 1980
Query: 322 ARAYAPSTIFIDEIDSLCNSRGAS 345
A +P+ IF+DE+DS+ R S
Sbjct: 1981 ASKVSPTIIFVDEVDSMLGQRTKS 2052
Score = 59.7 bits (143), Expect(2) = 7e-41
Identities = 30/73 (41%), Positives = 51/73 (69%)
Frame = +3
Query: 346 GEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRRRLEKRIYI 405
G ++ R++K+E + DG+ T + G R ++VLAATN P+D+DEA+ RR E+RI +
Sbjct: 2055 GSMKAMRKIKNEFMSHWDGL----TTKQGER--ILVLAATNKPFDLDEAIIRRFERRIMV 2216
Query: 406 PLPNFESRKELIR 418
LP+ E+R++++R
Sbjct: 2217 GLPSVENREKILR 2255
>TC85936 homologue to GP|13537115|dbj|BAB40755. AtSUG1 {Arabidopsis
thaliana}, partial (98%)
Length = 1633
Score = 162 bits (409), Expect = 4e-40
Identities = 104/294 (35%), Positives = 168/294 (56%), Gaps = 7/294 (2%)
Frame = +1
Query: 225 LETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQ--GIRRPWKGVLMFGPPGTGKTL 282
+E P ++ + GL + + ++E + LP+ PE F+ GI +P KGVL++GPPGTGKTL
Sbjct: 595 VEKVPDSTYDMIGGLDQQIKEIKEVIELPIKHPELFESLGIAQP-KGVLLYGPPGTGKTL 771
Query: 283 LAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSR 342
LA+AVA TF VS + L K+ GE RMVR LF +AR +APS IF+DEIDS+ ++R
Sbjct: 772 LARAVAHHTDCTFIRVSGSELVQKYIGEGSRMVRELFVMAREHAPSIIFMDEIDSIGSAR 951
Query: 343 GASGEHESS---RRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRR-- 397
SG +R ELL Q+DG A+N+ + VL ATN +D+AL R
Sbjct: 952 MESGSGNGDSEVQRTMLELLNQLDGFE--ASNK------IKVLMATNRIDILDQALLRPG 1107
Query: 398 RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASLN 457
R++++I P PN ESR+++++I+ + + + +++ ++A + +G SG +L VC +A +
Sbjct: 1108RIDRKIEFPNPNEESRRDILKIHSRRMNLMRGIDLKKIAEKMNGASGAELKAVCTEAGMF 1287
Query: 458 GMRRKIAGKTRDEIKNMSKDDISKDPVAMCDFEEALVKVQRSVSQADIERHEKW 511
+R + T++ DFE A+ KV + ++ ++ + W
Sbjct: 1288ALRERRVHVTQE------------------DFEMAVAKVMKKETEKNMSLRKLW 1395
>TC86159 homologue to GP|21387177|gb|AAM47992.1 26S proteasome AAA-ATPase
subunit RPT4a-like protein {Arabidopsis thaliana},
complete
Length = 1640
Score = 159 bits (401), Expect = 3e-39
Identities = 100/255 (39%), Positives = 153/255 (59%), Gaps = 7/255 (2%)
Frame = +2
Query: 213 DPELAEMLERDVLETSPG-VRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIR-RPWKGV 270
DP + ML D PG + + V GL++ R L E++ LPL PE F + +P KGV
Sbjct: 434 DPVVYNMLHED-----PGNISYSAVGGLSDQIRELRESIELPLMNPELFLRVGIKPPKGV 598
Query: 271 LMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTI 330
L++GPPGTGKTLLA+A+A+ F V S+ + K+ GES R++R +F AR + P I
Sbjct: 599 LLYGPPGTGKTLLARAIASNIDANFLKVVSSAIIDKYIGESARLIREMFGYARDHQPCII 778
Query: 331 FIDEIDSLCN---SRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNF 387
F+DEID++ S G S + E R + ELL Q+DG ++ G K++M ATN
Sbjct: 779 FMDEIDAIGGRRFSEGTSADREIQRTL-MELLNQLDGF-----DQLGKVKMIM---ATNR 931
Query: 388 PWDIDEALRR--RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGD 445
P +D AL R RL+++I IPLPN +SR E+++I+ + +++ + V + +G++G
Sbjct: 932 PDVLDPALLRPGRLDRKIEIPLPNEQSRMEILKIHAAGIAKHGEIDYEAVVKLAEGFNGA 1111
Query: 446 DLTNVCRDASLNGMR 460
DL NVC +A ++ +R
Sbjct: 1112DLRNVCTEAGMSAIR 1156
>TC77012 homologue to SP|P54776|PRSA_LYCES 26S protease regulatory subunit
6A homolog (TAT-binding protein homolog 1) (TBP-1)
(Mg(2+), complete
Length = 1793
Score = 152 bits (385), Expect = 2e-37
Identities = 97/244 (39%), Positives = 143/244 (57%), Gaps = 7/244 (2%)
Frame = +1
Query: 225 LETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQ--GIRRPWKGVLMFGPPGTGKTL 282
++ P + D+ GL + + L EA+VLP+ E FQ GIR P KGVL++GPPGTGKTL
Sbjct: 613 VDEKPTEDYNDIGGLEKQIQELVEAIVLPMTHKERFQKLGIRPP-KGVLLYGPPGTGKTL 789
Query: 283 LAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSR 342
+A+A A + TF ++ L + G+ ++VR F LA+ +P IFIDEID++ R
Sbjct: 790 MARACAAQTNATFLKLAGPQLVQMFIGDGAKLVRDAFQLAKEKSPCIIFIDEIDAIGTKR 969
Query: 343 ---GASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRR-- 397
SG+ E +R ELL Q+DG S S + V+AATN +D AL R
Sbjct: 970 FDSEVSGDRE-VQRTMLELLNQLDGFS--------SDDRIKVIAATNRADILDPALMRSG 1122
Query: 398 RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASLN 457
RL+++I P P E+R +++I+ + + V PDVN +E+AR TD ++G L VC +A +
Sbjct: 1123RLDRKIEFPHPTEEARARILQIHSRKMNVHPDVNFEELARSTDDFNGAQLKAVCVEAGML 1302
Query: 458 GMRR 461
+RR
Sbjct: 1303ALRR 1314
>TC85745 homologue to GP|17297987|dbj|BAB78491. 26S proteasome regulatory
particle triple-A ATPase subunit2b {Oryza sativa
(japonica cultivar-group), partial (98%)
Length = 1696
Score = 152 bits (385), Expect = 2e-37
Identities = 97/270 (35%), Positives = 151/270 (55%), Gaps = 5/270 (1%)
Frame = +3
Query: 215 ELAEMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIR-RPWKGVLMF 273
E+ M+ +E +P + D+ GL + ++EAV LPL PE ++ I +P KGV+++
Sbjct: 594 EVDPMVSVMKVEKAPLESYADIGGLDAQIQEIKEAVELPLTHPELYEDIGIKPPKGVILY 773
Query: 274 GPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFID 333
G PGTGKTLLAKAVA TF V + L K+ G+ ++VR LF +A +P+ +FID
Sbjct: 774 GEPGTGKTLLAKAVANSTSATFLRVVGSELIQKYLGDGPKLVRELFRVADDLSPAIVFID 953
Query: 334 EIDSLCNSR--GASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDI 391
EID++ R SG +R ELL Q+DG SR V V+ ATN +
Sbjct: 954 EIDAVGTKRYDAHSGGEREIQRTMLELLNQLDGFD--------SRGDVKVILATNRIESL 1109
Query: 392 DEALRR--RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTN 449
D AL R R++++I PLP+ ++R+ + +I+ + +A DVN++E D +SG D+
Sbjct: 1110DPALLRPGRIDRKIEFPLPDIKTRRRIFQIHTSRMTLADDVNLEEFVMTKDEFSGADIKA 1289
Query: 450 VCRDASLNGMRRKIAGKTRDEIKNMSKDDI 479
+C +A L +R + T + K +KD +
Sbjct: 1290ICTEAGLLALRERRMKVTHPDFKK-AKDKV 1376
>TC85744 homologue to PIR|T08959|T08959 proteinase homolog F19B15.70 -
Arabidopsis thaliana, complete
Length = 1629
Score = 152 bits (385), Expect = 2e-37
Identities = 98/270 (36%), Positives = 150/270 (55%), Gaps = 5/270 (1%)
Frame = +2
Query: 215 ELAEMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIR-RPWKGVLMF 273
E+ M+ +E +P + D+ GL + ++EAV LPL PE ++ I +P KGV+++
Sbjct: 596 EVDPMVSVMKVEKAPLESYADIGGLDAQIQEIKEAVELPLTHPELYEDIGIKPPKGVILY 775
Query: 274 GPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFID 333
G PGTGKTLLAKAVA TF V + L K+ G+ ++VR LF +A +PS +FID
Sbjct: 776 GEPGTGKTLLAKAVANSTSATFLRVVGSELIQKYLGDGPKLVRELFRVADDLSPSIVFID 955
Query: 334 EIDSLCNSR--GASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDI 391
EID++ R SG +R ELL Q+DG SR V V+ ATN +
Sbjct: 956 EIDAVGTKRYDAHSGGEREIQRTMLELLNQLDGFD--------SRGDVKVILATNRIESL 1111
Query: 392 DEALRR--RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTN 449
D AL R R++++I PLP+ ++R+ + I+ + +A DVN++E D +SG D+
Sbjct: 1112DPALLRPGRIDRKIEFPLPDIKTRRRIFTIHTSRMTLADDVNLEEFVMTKDEFSGADIKA 1291
Query: 450 VCRDASLNGMRRKIAGKTRDEIKNMSKDDI 479
+C +A L +R + T + K +KD +
Sbjct: 1292ICTEAGLLALRERRMKVTHADFKK-AKDKV 1378
>TC77340 SP|Q9BAE0|FTSH_MEDSA Cell division protein ftsH homolog chloroplast
precursor (EC 3.4.24.-). [Alfalfa] {Medicago sativa},
complete
Length = 2473
Score = 150 bits (378), Expect = 1e-36
Identities = 97/256 (37%), Positives = 154/256 (59%), Gaps = 6/256 (2%)
Frame = +3
Query: 222 RDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQ--GIRRPWKGVLMFGPPGTG 279
++V ET GV + DVAG +AK L+E V L P+ + G + P KG L+ GPPGTG
Sbjct: 978 QEVPET--GVTFADVAGADQAKLELQEVVDF-LKNPDKYTALGAKIP-KGCLLVGPPGTG 1145
Query: 280 KTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLC 339
KTLLA+AVA E GT FF+ +++ + G VR LF+ A++ AP +FIDEID++
Sbjct: 1146 KTLLARAVAGEAGTPFFSCAASEFVELFVGVGASRVRDLFEKAKSKAPCIVFIDEIDAVG 1325
Query: 340 NSRGA--SGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRR 397
RGA G ++ + ++LL ++DG S + V+VLAATN P +D AL R
Sbjct: 1326 RQRGAGLGGGNDEREQTINQLLTEMDGFSGNSG--------VIVLAATNRPDVLDSALLR 1481
Query: 398 --RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDAS 455
R ++++ + P+ R ++++++ + +A DV+ D++ARRT G++G DL N+ +A+
Sbjct: 1482 PGRFDRQVTVDRPDVAGRVKILQVHSRGKALAKDVDFDKIARRTPGFTGADLQNLMNEAA 1661
Query: 456 LNGMRRKIAGKTRDEI 471
+ RR + ++DEI
Sbjct: 1662 ILAARRDLKEISKDEI 1709
>TC76904 homologue to GP|4325041|gb|AAD17230.1| FtsH-like protein Pftf
precursor {Nicotiana tabacum}, partial (91%)
Length = 2667
Score = 145 bits (366), Expect = 4e-35
Identities = 95/254 (37%), Positives = 145/254 (56%), Gaps = 7/254 (2%)
Frame = +1
Query: 225 LETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQ--GIRRPWKGVLMFGPPGTGKTL 282
+E + GV ++DVAG+ EAK+ E V L PE F G R P KGVL+ GPPGTGKTL
Sbjct: 802 MEPNTGVTFDDVAGVDEAKQDFMEVVEF-LKKPERFTSVGARIP-KGVLLVGPPGTGKTL 975
Query: 283 LAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSR 342
LAKA+A E G FF++S + + G VR LF A+ AP +F+DEID++ R
Sbjct: 976 LAKAIAGEAGVPFFSISGSEFVEMFVGIGASRVRDLFKKAKENAPCIVFVDEIDAVGRQR 1155
Query: 343 GA--SGEHESSRRVKSELLVQVDGV-SNTATNEDGSRKIVMVLAATNFPWDIDEALRR-- 397
G G ++ + ++LL ++DG NT V+V+AATN +D AL R
Sbjct: 1156 GTGIGGGNDEREQTLNQLLTEMDGFEGNTG---------VIVVAATNRADILDSALLRPG 1308
Query: 398 RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASLN 457
R ++++ + +P+ R E+++++ + DV+++ +A RT G+SG DL N+ +A++
Sbjct: 1309 RFDRQVSVDVPDVRGRTEILKVHANNKKFDNDVSLEVIAMRTPGFSGADLANLLNEAAIL 1488
Query: 458 GMRRKIAGKTRDEI 471
RR G + EI
Sbjct: 1489 AGRRGRTGISPKEI 1530
>AW774182 homologue to PIR|F84674|F84 probable AAA-type ATPase [imported] -
Arabidopsis thaliana, partial (27%)
Length = 487
Score = 144 bits (364), Expect = 6e-35
Identities = 68/120 (56%), Positives = 91/120 (75%)
Frame = -1
Query: 231 VRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIRRPWKGVLMFGPPGTGKTLLAKAVATE 290
V+W DVAGL AK+ L+ AV+LP+ ++ G RRPW+ L++GPPGTGK+ LAKAVAT+
Sbjct: 361 VKWNDVAGLKSAKQSLQ*AVILPVKYRQFLTGKRRPWRAFLLYGPPGTGKSYLAKAVATK 182
Query: 291 CGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRGASGEHES 350
+TFF+VSS+ L SKW GESE++V LF++AR APS IF+DEIDSLC +RG E ++
Sbjct: 181 ADSTFFSVSSSDLVSKWMGESEKLVSNLFEMARESAPSIIFVDEIDSLCGTRGEGNESKA 2
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.313 0.131 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,940,425
Number of Sequences: 36976
Number of extensions: 181967
Number of successful extensions: 1367
Number of sequences better than 10.0: 128
Number of HSP's better than 10.0 without gapping: 1072
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1298
length of query: 518
length of database: 9,014,727
effective HSP length: 100
effective length of query: 418
effective length of database: 5,317,127
effective search space: 2222559086
effective search space used: 2222559086
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0226.8


