

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.7
(399 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
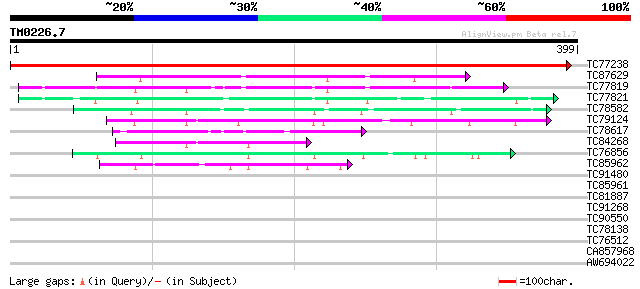
Score E
Sequences producing significant alignments: (bits) Value
TC77238 similar to GP|15289767|dbj|BAB63467. putative aspartate ... 719 0.0
TC87629 similar to GP|17381280|gb|AAL36058.1 At2g22250/T26C19.9 ... 111 6e-25
TC77819 similar to GP|21618197|gb|AAM67247.1 putative aminotrans... 83 2e-16
TC77821 similar to GP|21618197|gb|AAM67247.1 putative aminotrans... 82 4e-16
TC78582 similar to GP|10177259|dbj|BAB10727. tyrosine aminotrans... 62 4e-10
TC79124 1-aminocyclopropanecarboxylic acid synthase [Medicago tr... 62 5e-10
TC78617 similar to PIR|T03270|T03270 probable histidinol-phospha... 57 2e-08
TC84268 similar to GP|6650982|gb|AAF22112.1| 1-aminocyclopropane... 47 1e-05
TC76856 similar to PIR|H96729|H96729 probable alanine aminotrans... 46 3e-05
TC85962 similar to GP|21954071|gb|AAK64147.2 putative alanine am... 44 1e-04
TC91480 homologue to GP|13359455|dbj|BAB33423. ACC synthase {Pis... 37 0.018
TC85961 similar to PIR|B96747|B96747 probable alanine aminotrans... 34 0.087
TC81887 weakly similar to GP|6006360|dbj|BAA84790.1 Similar to 1... 33 0.19
TC91268 homologue to PIR|T06252|T06252 1-aminocyclopropane-1-car... 32 0.56
TC90550 weakly similar to GP|17065498|gb|AAL32903.1 tyrosine ami... 30 1.3
TC78138 similar to GP|2792155|emb|CAA11226.1 chalcone reductase ... 29 3.7
TC76512 similar to GP|4959932|gb|AAD34548.1| cystathionine-gamma... 29 3.7
CA857968 similar to PIR|T48188|T48 aldose reductase-like protein... 28 4.8
AW694022 similar to GP|16494|emb| DNA-directed RNA polymerase {A... 28 6.2
>TC77238 similar to GP|15289767|dbj|BAB63467. putative aspartate
transaminase {Oryza sativa (japonica cultivar-group)},
partial (98%)
Length = 1876
Score = 719 bits (1857), Expect = 0.0
Identities = 349/396 (88%), Positives = 380/396 (95%), Gaps = 1/396 (0%)
Frame = +3
Query: 1 MGSYANLARRALETEMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPS 60
MGSY NLA+RA+ETEMPIMV+MQE+LRGAKN VSLAQGVVYWQPPK ALDKVKELV+EPS
Sbjct: 450 MGSYGNLAKRAVETEMPIMVKMQELLRGAKNAVSLAQGVVYWQPPKQALDKVKELVYEPS 629
Query: 61 VSRYGADEGLPELRAALVKKLRQENNLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFA 120
+SRYG DEG+PELRAALVKKLR ENNLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVM+A
Sbjct: 630 ISRYGNDEGIPELRAALVKKLRNENNLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMYA 809
Query: 121 PYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGT 180
PYYFN+YMSFQMTGITNI VGPG+P+TLYPD DWLE++L+ESKPVPKLVTVVNPGNPTGT
Sbjct: 810 PYYFNSYMSFQMTGITNILVGPGNPETLYPDADWLEKVLSESKPVPKLVTVVNPGNPTGT 989
Query: 181 YIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSCVEGNHIVNIFSFSKAYGMMGW 240
YIPESLL+RIANLC+ AGSWL+VDNTYEYFMFD LKH+CVEGNH+VNIFSFSKAYGMMGW
Sbjct: 990 YIPESLLKRIANLCEKAGSWLVVDNTYEYFMFDDLKHTCVEGNHVVNIFSFSKAYGMMGW 1169
Query: 241 RVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAKNRE 300
R+GYIAYPSEVEGL +QLLKVQDNIPICASIISQHLAL+SLE+GPEWV ERV+TLAKNR+
Sbjct: 1170RIGYIAYPSEVEGLGSQLLKVQDNIPICASIISQHLALYSLELGPEWVTERVKTLAKNRQ 1349
Query: 301 IVSEALSPLGEGSVKGGEGAIYLFAKLPKG-GYDDFDVVRWLANRHGIAVIPGSACGAAG 359
IV EALS LGEGS+KGGEGAIYL+ K+P G GYDDF+VVRWLANRHGIAVIPGSACGAAG
Sbjct: 1350IVMEALSSLGEGSIKGGEGAIYLWVKIPSGHGYDDFEVVRWLANRHGIAVIPGSACGAAG 1529
Query: 360 NLRISFGGLTESDCRAAAERLKKGLEELVANGLVQD 395
NLRISFGGLTE+DCRAAAERLKKGLEELVA+GLVQD
Sbjct: 1530NLRISFGGLTENDCRAAAERLKKGLEELVAHGLVQD 1637
>TC87629 similar to GP|17381280|gb|AAL36058.1 At2g22250/T26C19.9
{Arabidopsis thaliana}, partial (72%)
Length = 1309
Score = 111 bits (277), Expect = 6e-25
Identities = 77/274 (28%), Positives = 134/274 (48%), Gaps = 11/274 (4%)
Frame = +2
Query: 62 SRYGADEGLPELRAALVKKLRQENNLHKS--SVMVTAGANQAFVNLVLTLCDAGDSVVMF 119
+RY + G E+R A+ KL++EN L + ++VT GA Q+ V+ +C GD V++
Sbjct: 467 TRYTPNAGTLEIRQAICHKLKEENGLDYTPDQIVVTNGAKQSITQAVIAVCSPGDEVIIP 646
Query: 120 APYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTG 179
AP++ + ++ T + + + D LE +TE +L+ + +P NPTG
Sbjct: 647 APFWVSYPEMARLADATPVILPTSISNNFLLDPKLLESKITEKS---RLLILCSPSNPTG 817
Query: 180 TYIPESLLQRIANL-CKNAGSWLLVDNTYEYFMFDGLKHSCVEG-----NHIVNIFSFSK 233
+ P+ LL+ IA + K+ ++ D YE+ ++ H+ N + + FSK
Sbjct: 818 SVYPKYLLEEIAKIVAKHPRLLVISDEIYEHIIYAPATHTSFASLPRMWNRTLTVNGFSK 997
Query: 234 AYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEM---GPEWVRE 290
A+ M GWR+GYIA P A + K+Q S ISQ + +L + G E V
Sbjct: 998 AFAMTGWRLGYIAGPKH---FIAAVGKIQSQFTSGPSSISQKAGVAALGLGYAGGEAVST 1168
Query: 291 RVQTLAKNREIVSEALSPLGEGSVKGGEGAIYLF 324
V+ + ++ + ++ S + + +GA YLF
Sbjct: 1169MVKAFRERKDYLVKSFSEIDGVKISEPQGAFYLF 1270
>TC77819 similar to GP|21618197|gb|AAM67247.1 putative aminotransferase
{Arabidopsis thaliana}, partial (90%)
Length = 1235
Score = 82.8 bits (203), Expect = 2e-16
Identities = 83/356 (23%), Positives = 154/356 (42%), Gaps = 11/356 (3%)
Frame = +2
Query: 7 LARRALETEMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSVSRYGA 66
+A++ + + I QM +L ++L QG + P+ + + + + + ++Y
Sbjct: 101 VAKQLEQFKTTIFTQMS-MLAITHGAINLGQGFPNFDGPEFVKEAAIQAIRDGN-NQYAR 274
Query: 67 DEGLPELRAALVKKLRQENNL---HKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYY 123
G+P+L A+ ++ +++ L + + VT+G +A VL L + G+ V++FAP+Y
Sbjct: 275 GFGVPDLNIAIAERFKKDTGLIVDPDNEITVTSGCTEAIAATVLGLINPGEEVILFAPFY 454
Query: 124 --FNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTY 181
+ A +S I +I + P PD P +D L+ ++++ + + + P NPTG
Sbjct: 455 DSYGATLSMAGANIKSITLRP--PDFAVP-IDDLKSTISKNT---RAILINTPHNPTGKM 616
Query: 182 IPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSCVEG-----NHIVNIFSFSKAYG 236
L +A+LC + D Y FD ++H + V + S K++
Sbjct: 617 FTREELDSVASLCIENDVLVFTDEEYHKLAFD-MEHISIATLPGMFERTVTMNSLGKSFS 793
Query: 237 MMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLA 296
+ GW+VG+ P L + + I S Q A +L + E +
Sbjct: 794 LTGWKVGWAIAPPH---LMWGVRQAHTYIAFSISHPLQCGAAAALRAPDSYYVELKRDYI 964
Query: 297 KNREIVSEALSPLGEGSVKGGEGAIYLFA-KLPKGGYDDFDVVRWLANRHGIAVIP 351
R I+ E L +G V G ++ A P G +D +L G+A IP
Sbjct: 965 AKRAILVEGLKAVG-FKVFPTNGTFFVLADHTPFGHENDVAFCEYLIKEIGVAAIP 1129
>TC77821 similar to GP|21618197|gb|AAM67247.1 putative aminotransferase
{Arabidopsis thaliana}, partial (93%)
Length = 1631
Score = 82.0 bits (201), Expect = 4e-16
Identities = 93/400 (23%), Positives = 159/400 (39%), Gaps = 20/400 (5%)
Frame = +3
Query: 7 LARRALETEMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEP---SVSR 63
+A+R + + I QM +L ++L QG + P D VKE + ++
Sbjct: 279 VAKRLEKFKTTIFTQMS-MLAIKHGAINLGQGFPNFDGP----DFVKEAAIQAIRDGNNQ 443
Query: 64 YGADEGLPELRAALVKKLRQENNLH---KSSVMVTAGANQAFVNLVLTLCDAGDSVVMFA 120
Y G+P+L A+ ++ +++ L + + VT+G +A VL L + GD V++FA
Sbjct: 444 YARGYGVPDLNIAISERYKKDTGLAVDPEKEITVTSGCTEAIAATVLGLINPGDEVIVFA 623
Query: 121 PYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGT 180
P+Y + + M G + PD P +E + + + + + P NPTG
Sbjct: 624 PFYDSYEATLSMAGAKVKGITLRPPDFALP----IEELKSTISKNTRAILLNTPHNPTGK 791
Query: 181 YIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSCVEG-----NHIVNIFSFSKAY 235
L IA+LC + D Y+ FD ++H + V + S K +
Sbjct: 792 MFTPEELNTIASLCIENDVLVFSDEVYDKLAFD-MEHISIASLPGMFERTVTMNSLGKTF 968
Query: 236 GMMGWRVGYIAYPSE----VEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRER 291
+ GW++G+ P V A L N A+ ++ L + PE R+
Sbjct: 969 SLTGWKIGWAIAPPHLTWGVRQAHAFLTFATSNPMQWAAAVA--LRATRFLLPPELKRDY 1142
Query: 292 VQTLAKNREIVSEALSPLGEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIP 351
+ R I+ E L +G + P G +D +L G+ IP
Sbjct: 1143M----AKRSILVEGLKAVGFKVFPSSGTYFVVVDHTPFGHENDIAFCEYLVKEVGVVAIP 1310
Query: 352 GSAC-----GAAGNLRISFGGLTESDCRAAAERLKKGLEE 386
S +R +F E RAA +R+K+ L +
Sbjct: 1311TSVFYLNPEEGKNLVRFTF-CKDEGTLRAAVDRMKEKLRK 1427
>TC78582 similar to GP|10177259|dbj|BAB10727. tyrosine aminotransferase
{Arabidopsis thaliana}, partial (87%)
Length = 1788
Score = 62.0 bits (149), Expect = 4e-10
Identities = 79/364 (21%), Positives = 147/364 (39%), Gaps = 28/364 (7%)
Frame = +2
Query: 46 KPALDKVKELVWEPSVSRYGADEGLPELRAALVKKLRQE--NNLHKSSVMVTAGANQAFV 103
K A + V + + + Y GL + R A+ K L + L V +T G QA
Sbjct: 506 KVAEEAVADALCSGNFHGYAPTAGLLQARNAIAKYLSDDLPYELSSDDVFITCGCTQAID 685
Query: 104 NLVLTLCDAGDSVVMFAPYY--FNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTE 161
V L G ++++ P + + +F+ + + + P + D+D +E ++ +
Sbjct: 686 VSVALLSRPGANILLPRPGFPIYELCAAFRQVEVRHYDLLPEKGWEV--DLDAIETLVDQ 859
Query: 162 SKPVPKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNTYEYFMFD-------G 214
+ + ++NPGNP G L++IA K G+ ++ D Y + F G
Sbjct: 860 NTVA---LVIINPGNPCGNVYTYHHLEKIAETAKRLGTIVIADEVYGHLAFGDNPFVPMG 1030
Query: 215 LKHSCVEGNHIVNIFSFSKAYGMMGWRVGYIA---------YPSEVEGLSAQLLKVQDNI 265
+ S V ++ + S SK + + GWR+G+ P VE ++ K D +
Sbjct: 1031VFGSTVP---VITLGSLSKRWIVPGWRLGWFVTNDPSGTFRKPKVVE----RIKKYFDLL 1189
Query: 266 PICASIISQHLALHSLEMGPEWVRERVQTLAKNREIVSEALSPL--------GEGSVKGG 317
A+ I + + + R+ + L +I + + + +GS+
Sbjct: 1190GGPATFIQAAVPRIITQTEEVFFRKTIDKLRHTADICCQEMEDIPCISFPCKPQGSM--- 1360
Query: 318 EGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGNLRISFGGLTESDCRAAA 377
+ L L + DD D LA + ++PG+A G +RI+F S R
Sbjct: 1361AMMVELNLSLLEDISDDIDFCFKLAKEESVIILPGTAVGLKDWIRITFAA-DPSALRDGM 1537
Query: 378 ERLK 381
+R+K
Sbjct: 1538QRIK 1549
>TC79124 1-aminocyclopropanecarboxylic acid synthase [Medicago truncatula]
Length = 1862
Score = 61.6 bits (148), Expect = 5e-10
Identities = 78/344 (22%), Positives = 143/344 (40%), Gaps = 31/344 (9%)
Frame = +3
Query: 69 GLPELRAALVKKLRQENN----LHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYY- 123
GLPE R A+ K + + ++++ GA A L D GD+ ++ PYY
Sbjct: 456 GLPEFRNAVAKFMSRTRGNRVTFDPDRIVMSGGATGAHEATAFCLADPGDAFLVPTPYYP 635
Query: 124 -FNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERIL---TESKPVPKLVTVVNPGNPTG 179
F+ + ++ TG+ + V S + LE TE K + + NP NP G
Sbjct: 636 GFDRDLRWR-TGVKLVPVICESANNFKLTKQALEEAYEKATEDNIRIKGLLITNPSNPLG 812
Query: 180 TYIPESLLQRIANLCKNAGSWLLVDNTYEYFMF---------DGLKHSC---VEGNHIVN 227
T + + L+ + N L+ D Y +F + L+H + N +
Sbjct: 813 TVMDRNTLRTVVNFINEKRIHLISDEIYAATVFSHPSFISIAEILEHDTDIECDRNLVHI 992
Query: 228 IFSFSKAYGMMGWRVGYI-AYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSL---EM 283
++S SK G G+RVG I +Y V + ++ S +Q+L L E
Sbjct: 993 VYSLSKDMGFPGFRVGIIYSYNDTVVNCARKMSSFG-----LVSTQTQYLMAKMLSDDEF 1157
Query: 284 GPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAIY---LFAKLPKGGYD-DFDVVR 339
+++ E + LA+ I + L+ +G ++ G L L + ++ + ++ R
Sbjct: 1158VKKFLTESAKRLAQRYRIFTSGLTKVGINCLQSNGGLFVWMDLRGLLKEATFESELELWR 1337
Query: 340 WLANRHGIAVIPGSA--CGAAGNLRISFGGLTESDCRAAAERLK 381
+ + I V PG + C G R+ + + + D + A +R++
Sbjct: 1338VIIHEVKINVSPGVSFHCSEPGWFRVCYANMDDRDVQIALQRIR 1469
>TC78617 similar to PIR|T03270|T03270 probable histidinol-phosphate
transaminase (EC 2.6.1.9) - common tobacco, partial
(86%)
Length = 1707
Score = 56.6 bits (135), Expect = 2e-08
Identities = 50/180 (27%), Positives = 87/180 (47%), Gaps = 1/180 (0%)
Frame = +3
Query: 73 LRAALVKKLRQENNLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQM 132
LRAAL Q++ L ++V GA++ ++ + D GD +V P + +
Sbjct: 438 LRAALA----QDSGLESEYILVGCGADELIDLIMRCVLDPGDKIVDCPPTFTMYEFDAAV 605
Query: 133 TGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIAN 192
G I V P PD +V+ + ++ + KP K + + +P NP G+ I + L +I
Sbjct: 606 NGALVIKV-PRRPDFSL-NVEQIIEVVKQEKP--KCIFLTSPNNPDGSIIDDDDLLKILE 773
Query: 193 LCKNAGSWLLVDNTYEYFMFDGLKHSCVEGN-HIVNIFSFSKAYGMMGWRVGYIAYPSEV 251
L +++D Y F K S V+ + +++ + +FSK G+ G RVGY A+P +
Sbjct: 774 L----PILVVLDEAYIEFSTIESKMSWVKKHDNLIVLRTFSKRAGLAGLRVGYGAFPLSI 941
>TC84268 similar to GP|6650982|gb|AAF22112.1|
1-aminocyclopropane-1-carboxylate synthase 5 {Lupinus
albus}, partial (59%)
Length = 856
Score = 47.4 bits (111), Expect = 1e-05
Identities = 38/144 (26%), Positives = 68/144 (46%), Gaps = 6/144 (4%)
Frame = +1
Query: 75 AALVKKLR-QENNLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYY--FNAYMSFQ 131
A+ ++K+R + +++TAGA A L L + GD++++ PYY F+ + ++
Sbjct: 340 ASFMEKIRGNKAKFDYERIVLTAGATAANELLTFILANPGDALLVPTPYYPGFDRDLRWR 519
Query: 132 MTGITNIHVGPGSPDTLYPDVDWLERILTESKPVP---KLVTVVNPGNPTGTYIPESLLQ 188
TG+ I + S + ++ LE ++ + K V + NP NP G I S+L+
Sbjct: 520 -TGVNIIPIHCDSSNNFQITLEALETAYKNAESMNMKVKAVLITNPSNPLGISIQRSVLE 696
Query: 189 RIANLCKNAGSWLLVDNTYEYFMF 212
I N L+ D Y +F
Sbjct: 697 DILNFVTRKNIHLVSDEIYSGSVF 768
>TC76856 similar to PIR|H96729|H96729 probable alanine aminotransferase
F5A18.24 - Arabidopsis thaliana, complete
Length = 2043
Score = 45.8 bits (107), Expect = 3e-05
Identities = 76/357 (21%), Positives = 140/357 (38%), Gaps = 45/357 (12%)
Frame = +2
Query: 45 PKPALDKVKELVWEPS--VSRYGADEGLPELRAALVKKLRQENNLHKSS--VMVTAGANQ 100
P A+ + K+ + S + Y G P +R + + +++ + + +T GA++
Sbjct: 641 PADAIARAKQYLSFTSGGLGAYSDSRGTPAVRKEVAEFIQRRDGYPSDPEFIYLTDGASK 820
Query: 101 AFVNLVLTLCDAG-DSVVMFAPYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERIL 159
A + ++ T+ D +++ P Y + + G T + D + L R +
Sbjct: 821 AVMQILNTIIRGEVDGIMVPVPQYPLYSATIALLGGTLVPYYLEETANWGLDTNELRRSV 1000
Query: 160 TESKPVP---KLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNTYEYFMFD--- 213
E++ K + ++NPGNPTG + E L+ + C LL D Y+ ++
Sbjct: 1001REARYKGLHVKAMVIINPGNPTGQCLSEENLREVLQFCYEENLVLLGDEVYQTNIYQDER 1180
Query: 214 ----------GLKHSCVEGNHIVNIFSFSKAY-GMMGWRVGYIAY----PSEVEGL-SAQ 257
+ + +V+ S SK Y G G R GY P V+ +
Sbjct: 1181PFISAKKVLMDIGPPLSKEVQLVSFHSVSKGYFGECGQRGGYFEMTNIPPETVDEIYKVA 1360
Query: 258 LLKVQDNIPICASIISQHLALHSLEMG----PEWVRER---VQTLAKNREIVSEALSPLG 310
+ + N+P + I L ++ + G +VRE +++L + I+++ +
Sbjct: 1361SISLSPNVP---AQIFMGLMINPPKPGDISYDRFVRESKGVLESLRRRARIMTDGFNSCR 1531
Query: 311 EGSVKGGEGAIYLF--AKLP---------KGGYDDFDVVRWLANRHGIAVIPGSACG 356
EGA+Y F KLP G D L GI+ +PGS G
Sbjct: 1532NVVCNFTEGAMYSFPQIKLPPKALETAKQAGKAADVYYCLKLLEATGISTVPGSGFG 1702
>TC85962 similar to GP|21954071|gb|AAK64147.2 putative alanine
aminotransferase {Arabidopsis thaliana}, partial (71%)
Length = 1411
Score = 43.9 bits (102), Expect = 1e-04
Identities = 47/201 (23%), Positives = 83/201 (40%), Gaps = 23/201 (11%)
Frame = +3
Query: 64 YGADEGLPELRAALVKKLRQENNL--HKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAP 121
Y +G+ LR + + + + + + + +T GA+ A V++++ L + + P
Sbjct: 588 YSHSQGIQGLRDTIAAGIEERDGFPCNANDIFLTDGASPA-VHMMMQLLIRSEKDGILCP 764
Query: 122 YYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDW------LERILTESKPVP---KLVTVV 172
S +T +H G P L W L++ L ++K + + V+
Sbjct: 765 IPQYPLYSASIT----LHGGHLVPYYLDEATGWGLEISELKKQLEDAKSKGISVRALAVI 932
Query: 173 NPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNTYE---------YFMFDGLKHSCVEGN 223
NPGNPTG + E + I CK G LL D Y+ + F + S G+
Sbjct: 933 NPGNPTGQVLAEDNQRAIVEFCKQEGLVLLADEVYQENVYVPEKKFHSFKKVSRSMGYGD 1112
Query: 224 HIVNIFSF---SKAYGMMGWR 241
+ + + SF SK Y W+
Sbjct: 1113NDICLVSFQSVSKGYHGGVWK 1175
>TC91480 homologue to GP|13359455|dbj|BAB33423. ACC synthase {Pisum
sativum}, partial (38%)
Length = 666
Score = 36.6 bits (83), Expect = 0.018
Identities = 36/145 (24%), Positives = 63/145 (42%), Gaps = 9/145 (6%)
Frame = +2
Query: 69 GLPELRAALVKKLRQENN----LHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYY- 123
GLPE R A+ + + ++++ GA A ++ L D GD+ ++ +PYY
Sbjct: 233 GLPEFRNAVANFMSKVRGGRVRFDPDRILMSGGATGANELIMFCLADPGDAFLVPSPYYP 412
Query: 124 -FNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLE---RILTESKPVPKLVTVVNPGNPTG 179
F + ++ TG+ I V S + + LE + E K + + NP NP G
Sbjct: 413 AFVRDLCWR-TGVQLIPVQCHSSNNFKITREALEEAHKKAQEENINVKGLIITNPSNPLG 589
Query: 180 TYIPESLLQRIANLCKNAGSWLLVD 204
T + + L+ I + L+ D
Sbjct: 590 TTLEKDTLKSIVSFINENNIHLVCD 664
>TC85961 similar to PIR|B96747|B96747 probable alanine aminotransferase
T10D10.20 [imported] - Arabidopsis thaliana, partial
(29%)
Length = 744
Score = 34.3 bits (77), Expect = 0.087
Identities = 37/124 (29%), Positives = 54/124 (42%), Gaps = 24/124 (19%)
Frame = +1
Query: 257 QLLKVQDNIPICASIISQHLALHSLEMGPEWVRER------------VQTLAKNREIVSE 304
Q+ KV ++ +C++I Q LA S+ M P V + + +L + + E
Sbjct: 1 QIYKVA-SVNLCSNITGQILA--SVIMSPPKVGDESYESFMAERGAILSSLTTRAKALEE 171
Query: 305 ALSPLGEGSVKGGEGAIYLF------------AKLPKGGYDDFDVVRWLANRHGIAVIPG 352
AL+ L + EGA+YLF A+ K D F R L N GI V+PG
Sbjct: 172 ALNKLEGVTCNKAEGAMYLFPRIRLPEKAIKAAEAEKSAPDAFYCKR-LLNATGIVVVPG 348
Query: 353 SACG 356
S G
Sbjct: 349 SGFG 360
>TC81887 weakly similar to GP|6006360|dbj|BAA84790.1 Similar to
1-aminocyclopropane-1-carboxylate synthase (U35779),
partial (46%)
Length = 1182
Score = 33.1 bits (74), Expect = 0.19
Identities = 53/275 (19%), Positives = 116/275 (41%), Gaps = 23/275 (8%)
Frame = +2
Query: 134 GITNIHVGPGSPDTLYPDVDWLERILTESKP---VPKLVTVVNPGNPTGTYIPESLLQRI 190
GI I V S D ++ LE+ ++++ + + + NP NP G + +L +
Sbjct: 35 GIDLIPVHCRSTDNFNLNITALEQAFSQARKRGVKVRGILISNPSNPVGNILTPDMLFSL 214
Query: 191 ANLCKNAGSWLLVDNTY-------EYF--MFDGLKHSCVEGNHIVNIFSFSKAYGMMGWR 241
+ ++ ++ D + E F + + L ++ + + ++ SK + G+R
Sbjct: 215 LDFAEDKNIHIIADEVFAGSTYGSEEFVSIAEVLDSEYIDKSRVHIVYGLSKDLSIAGFR 394
Query: 242 VGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQT----LAK 297
VG I ++ +A+ + I S +Q L + S+ +++E ++T + +
Sbjct: 395 VGVIHSHNKTVLDAAKKMSRFSAI----SAPTQRL-VTSMLSDKRFIQEYIETNRNRIRR 559
Query: 298 NREIVSEALSPLGEGSVKGGEGAIYLFAKL-----PKGGYDDFDVVRWLANRHGIAVIPG 352
+ + L+ LG K G ++ +A + P + ++ + I + PG
Sbjct: 560 VHDEFVDCLTKLGIKCAKSSAG-MFCWADMSGLIRPYSEKGELELWEKFMSVAKINITPG 736
Query: 353 SACGA--AGNLRISFGGLTESDCRAAAERLKKGLE 385
SAC G RI F ++ + +R++K +E
Sbjct: 737 SACHCIEPGWFRICFTTISFEEIPVVIDRIRKVVE 841
>TC91268 homologue to PIR|T06252|T06252 1-aminocyclopropane-1-carboxylate
synthase (EC 4.4.1.14) 1 - garden pea, partial (38%)
Length = 627
Score = 31.6 bits (70), Expect = 0.56
Identities = 17/59 (28%), Positives = 31/59 (51%), Gaps = 4/59 (6%)
Frame = +1
Query: 69 GLPELRAALVKKLRQ----ENNLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYY 123
GLP + ALV + + + + +++TAGA A L+ L + G++ ++ PYY
Sbjct: 337 GLPSFKKALVDFMAEIRGNKVTFDPNHIVLTAGATSANETLMFCLAEQGEAFLLPTPYY 513
>TC90550 weakly similar to GP|17065498|gb|AAL32903.1 tyrosine
aminotransferase-like protein {Arabidopsis thaliana},
partial (44%)
Length = 925
Score = 30.4 bits (67), Expect = 1.3
Identities = 39/157 (24%), Positives = 60/157 (37%), Gaps = 14/157 (8%)
Frame = +1
Query: 239 GWRVGYIAY--PSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLA 296
GWR G+IA P ++ Q+ + NI I + L+ + + + +
Sbjct: 16 GWRTGWIATCDPHKI----FQITGIVKNIISYLEITADPLSFMQAAIPQILEKTKPEFHL 183
Query: 297 KNREIVSEALSPLGEGSVK--------GGEGAIYLFAKLP----KGGYDDFDVVRWLANR 344
KN I+ EA + + EGA+ ++ KG DD D LA
Sbjct: 184 KNMNIMKEAADIFYDVCKEIPCLTCPHKPEGAMSAMVEINFAHVKGIIDDVDFCDKLAKE 363
Query: 345 HGIAVIPGSACGAAGNLRISFGGLTESDCRAAAERLK 381
+ ++PG G LRIS + SD RLK
Sbjct: 364 ESVILLPGVTVGLKNWLRISL-AVDPSDLEEGLSRLK 471
>TC78138 similar to GP|2792155|emb|CAA11226.1 chalcone reductase {Sesbania
rostrata}, partial (83%)
Length = 1289
Score = 28.9 bits (63), Expect = 3.7
Identities = 13/45 (28%), Positives = 23/45 (50%)
Frame = +2
Query: 277 ALHSLEMGPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAI 321
A++ +EM P W + +++ K + I A SPLG + G +
Sbjct: 584 AVNQVEMNPSWQQGKLREFCKQKGIHVSAWSPLGGYKLSWGSPTV 718
>TC76512 similar to GP|4959932|gb|AAD34548.1| cystathionine-gamma-synthase
precursor {Glycine max}, partial (90%)
Length = 2103
Score = 28.9 bits (63), Expect = 3.7
Identities = 28/123 (22%), Positives = 56/123 (44%), Gaps = 4/123 (3%)
Frame = +3
Query: 89 KSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMT----GITNIHVGPGS 144
+S++++ +G + V L++ L AG +V Y + + GIT V P
Sbjct: 720 ESTILMASGMCASIV-LLMALVPAGGHLVTTTDCYRKTRIFIETVLPKMGITTSVVDPA- 893
Query: 145 PDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLLVD 204
DV L+ L ++K V++ +PT ++ ++ ++ LC G+ L++D
Sbjct: 894 ------DVGALQSALEQNK-----VSLFFTESPTNPFLRCVDIKLVSELCHKNGALLVID 1040
Query: 205 NTY 207
T+
Sbjct: 1041GTF 1049
>CA857968 similar to PIR|T48188|T48 aldose reductase-like protein -
Arabidopsis thaliana, partial (66%)
Length = 775
Score = 28.5 bits (62), Expect = 4.8
Identities = 27/79 (34%), Positives = 36/79 (45%), Gaps = 6/79 (7%)
Frame = +2
Query: 281 LEMGPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAIY------LFAKLPKGGYDD 334
+EM P W +++ K I A SPL GS GG I+ + KL K
Sbjct: 245 MEMHPGWRNDKMLEACKKNGIHVTAYSPL--GSQDGGRDLIHDQTVDRIAKKLNKS--PG 412
Query: 335 FDVVRWLANRHGIAVIPGS 353
+V+W A R G +VIP S
Sbjct: 413 QVLVKW-AMRRGTSVIPKS 466
>AW694022 similar to GP|16494|emb| DNA-directed RNA polymerase {Arabidopsis
thaliana}, partial (8%)
Length = 495
Score = 28.1 bits (61), Expect = 6.2
Identities = 23/89 (25%), Positives = 38/89 (41%), Gaps = 18/89 (20%)
Frame = +2
Query: 48 ALDKVKELVWEPSVSRYGADEGLPELRA------------------ALVKKLRQENNLHK 89
A DKVK+L+ E + A+ G + + + K L + NNL
Sbjct: 203 AKDKVKQLIREAQEKKLEAEPGRTMMDSFENRVNQTLNRARDDAGNSAQKSLSESNNL-- 376
Query: 90 SSVMVTAGANQAFVNLVLTLCDAGDSVVM 118
MVTAG+ +F+N+ C G + ++
Sbjct: 377 -KAMVTAGSKGSFINISQDDCLCGPTELL 460
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.136 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,204,164
Number of Sequences: 36976
Number of extensions: 171431
Number of successful extensions: 781
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 770
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 775
length of query: 399
length of database: 9,014,727
effective HSP length: 98
effective length of query: 301
effective length of database: 5,391,079
effective search space: 1622714779
effective search space used: 1622714779
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0226.7


