

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.6
(263 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
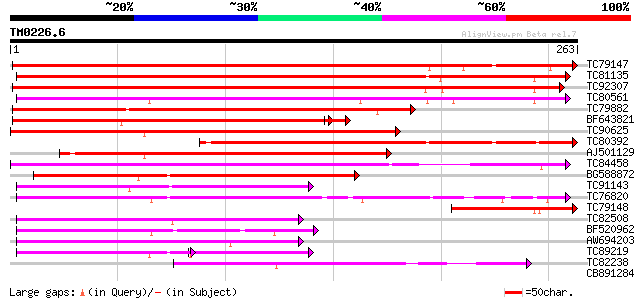
Sequences producing significant alignments: (bits) Value
TC79147 weakly similar to GP|10177856|dbj|BAB11208. phloem-speci... 305 1e-83
TC81135 weakly similar to GP|10177856|dbj|BAB11208. phloem-speci... 301 2e-82
TC92307 weakly similar to PIR|E84434|E84434 probable phloem-spec... 291 2e-79
TC80561 weakly similar to GP|10177856|dbj|BAB11208. phloem-speci... 221 2e-58
TC79882 weakly similar to PIR|E84434|E84434 probable phloem-spec... 218 2e-57
BF643821 weakly similar to PIR|F84434|F844 lectin-like protein [... 189 6e-49
TC90625 weakly similar to GP|10177856|dbj|BAB11208. phloem-speci... 179 7e-46
TC80392 weakly similar to GP|15088626|gb|AAK84134.1 nictaba {Nic... 161 3e-40
AJ501129 weakly similar to GP|10177856|dbj phloem-specific lecti... 156 9e-39
TC84458 weakly similar to PIR|B96604|B96604 hypothetical protein... 155 1e-38
BG588872 weakly similar to GP|21592660|gb phloem-specific lectin... 121 3e-28
TC91143 similar to PIR|T47557|T47557 hypothetical protein F8J2.1... 81 4e-16
TC76820 weakly similar to GP|18175688|gb|AAL59911.1 unknown prot... 79 2e-15
TC79148 77 9e-15
TC82508 similar to PIR|T50527|T50527 hypothetical protein T27I15... 72 2e-13
BF520962 weakly similar to PIR|C96656|C96 unknown protein 43836... 72 2e-13
AW694203 weakly similar to PIR|C96656|C96 unknown protein 43836... 66 2e-11
TC89219 weakly similar to PIR|C96656|C96656 unknown protein 438... 49 2e-11
TC82238 weakly similar to GP|8570046|dbj|BAA96751.1 Similar to A... 62 2e-10
CB891284 weakly similar to GP|22122920|gb unknown protein {Oryza... 40 0.001
>TC79147 weakly similar to GP|10177856|dbj|BAB11208. phloem-specific
lectin-like protein {Arabidopsis thaliana}, partial
(32%)
Length = 958
Score = 305 bits (781), Expect = 1e-83
Identities = 167/272 (61%), Positives = 197/272 (72%), Gaps = 10/272 (3%)
Frame = +2
Query: 2 ELLPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRAN 61
E L EGCI++ILSRTTPVDA RLS+VSK F SAADSDAVW+ FLPSD +SI+SQS S AN
Sbjct: 35 EDLAEGCISSILSRTTPVDAGRLSVVSKTFLSAADSDAVWNHFLPSDVNSIISQSPSLAN 214
Query: 62 ASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLP 121
A SKK YL LSD P++ID+ KKSFQLD+KSGKKC+ML AR+L I W + D Y WT +P
Sbjct: 215 APSKKALYLTLSDRPIIIDDAKKSFQLDRKSGKKCYMLAARSLHIIWGDDDRYWIWTAMP 394
Query: 122 ESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVNNIG 181
+SRFP+V LR + WLEI GMIN +LSPNTQY A+LVFK+ID F + V LSV G
Sbjct: 395 DSRFPEVANLRLVWWLEIRGMINNLALSPNTQYAAYLVFKLIDGYGFETLPVDLSVGVEG 574
Query: 182 GHNITKIAYLNP------DSLEGEVHDRWDRVQG---PSVRSDGWLEIEMGEFFISGLED 232
GH+ TKI L+P +S + R DRV G P VRSDGWLEIEMGE F SGLE+
Sbjct: 575 GHSSTKIVCLDPNVERRQNSRHARFYGRADRVVGLPRPGVRSDGWLEIEMGE-FNSGLEN 751
Query: 233 EEVEMSVLQIDGCFNVG-LIIEGIEVRPKNDN 263
EEV+MSV++I G +EGIEVRPK DN
Sbjct: 752 EEVQMSVIEIKAGETKGNFFLEGIEVRPKVDN 847
>TC81135 weakly similar to GP|10177856|dbj|BAB11208. phloem-specific
lectin-like protein {Arabidopsis thaliana}, partial
(32%)
Length = 1186
Score = 301 bits (770), Expect = 2e-82
Identities = 157/260 (60%), Positives = 197/260 (75%), Gaps = 3/260 (1%)
Frame = +2
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPEGCIAAI+SRTTP+DA RLS+ SK FRSA+DSDAVW++FLPSD SI+S S S N
Sbjct: 116 LPEGCIAAIVSRTTPLDAGRLSVFSKTFRSASDSDAVWNQFLPSDISSIISHSPSLTNIP 295
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPES 123
+KK YL LSD PV+ID+ KKS QL++KSGKKC+ML AR+L I W + Y W ++P+S
Sbjct: 296 TKKDLYLALSDRPVIIDHDKKSIQLERKSGKKCYMLSARSLAIVWGDDRRYWNWISMPDS 475
Query: 124 RFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVNNIGGH 183
RFP+V L + WLEIHG+INT LSPNTQY A++VFK+I+A F LSV GG
Sbjct: 476 RFPEVAKLVDVCWLEIHGVINTIVLSPNTQYAAYVVFKMINASGFQNRPADLSVGVEGGQ 655
Query: 184 NITKIAYLNPDSLEG--EVHDRWDRVQGPSVRSDGWLEIEMGEFFISGLEDEEVEMSVLQ 241
+ TKI L+P ++EG ++HDR D +Q PSVRSDGWLEIEMGEFF SG+E+EEV+M+V+Q
Sbjct: 656 SSTKIVCLDP-NVEGRPQLHDRVDGLQRPSVRSDGWLEIEMGEFFNSGIENEEVQMNVIQ 832
Query: 242 I-DGCFNVGLIIEGIEVRPK 260
G + GL++EGIEVRPK
Sbjct: 833 TKGGNWKRGLVLEGIEVRPK 892
>TC92307 weakly similar to PIR|E84434|E84434 probable phloem-specific lectin
[imported] - Arabidopsis thaliana, partial (19%)
Length = 817
Score = 291 bits (745), Expect = 2e-79
Identities = 157/265 (59%), Positives = 191/265 (71%), Gaps = 9/265 (3%)
Frame = +1
Query: 2 ELLPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRAN 61
E LPEGCIAA+LS TTPVDA RLS +SK F SA+DSDAVW+RFLPSD SI+SQ S +N
Sbjct: 22 EELPEGCIAAVLSSTTPVDACRLSFISKNFLSASDSDAVWNRFLPSDISSIISQFPSLSN 201
Query: 62 ASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLP 121
+KK Y+ LSD P++IDNGKKSFQL++KSGKKC+ML AR+L I W + D Y WT +P
Sbjct: 202 IPTKKDLYIALSDRPIIIDNGKKSFQLERKSGKKCYMLSARSLAITWGDTDQYWNWTVMP 381
Query: 122 ESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVNNIG 181
ESRFP+V L H+ WLEI G NT +LSPNT+Y +LVFK+I+A F V LSV G
Sbjct: 382 ESRFPEVAELLHVCWLEIRGK*NTLALSPNTRYVTYLVFKMINAYGFEYFPVELSVGIEG 561
Query: 182 GHNITKIAYL-NPDSLEGE-------VHDRWDRVQGPSVRSDGWLEIEMGEFFISGLEDE 233
GH+ TKI L +P++ +R R+Q P+VRSDGWLEIEMGEFFISGLEDE
Sbjct: 562 GHSSTKIVCLADPNARRRHRSRIIVTEPNRVLRLQRPNVRSDGWLEIEMGEFFISGLEDE 741
Query: 234 EVEMSVLQI-DGCFNVGLIIEGIEV 257
EV+MSV+ I G + G +EGIEV
Sbjct: 742 EVQMSVIDIKGGHWKRGFFVEGIEV 816
>TC80561 weakly similar to GP|10177856|dbj|BAB11208. phloem-specific
lectin-like protein {Arabidopsis thaliana}, partial
(52%)
Length = 1436
Score = 221 bits (563), Expect = 2e-58
Identities = 128/291 (43%), Positives = 173/291 (58%), Gaps = 34/291 (11%)
Frame = +3
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPEGCI +LS TTP D ARLSLVS FRSAA+SD+VWD+FLPSD HSI+S+S + + S
Sbjct: 171 LPEGCIENVLSFTTPRDVARLSLVSSTFRSAAESDSVWDKFLPSDYHSIISKSDAEIDNS 350
Query: 64 ----SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTT 119
KK Y++L+ P++I+NGKKSFQLDK GKKC+ML AR+LFI W + Y W
Sbjct: 351 LSVLPKKDLYVNLTQKPLLIENGKKSFQLDKVDGKKCYMLSARSLFIVWGDTPRYWRWIP 530
Query: 120 LPESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFK--IIDAPNFHEVLVVLSV 177
P+SRFP+ L + W EI G I+T LSP T YGA+LVFK A F S+
Sbjct: 531 DPDSRFPEAAELVSVCWFEIRGWISTIMLSPKTLYGAYLVFKSSAAGAFGFEYQPCEASI 710
Query: 178 NNIGGHNITKIAYLN------------PDSLEGEVHDRW---------------DRVQGP 210
+ GG + + +L+ P S + R D + P
Sbjct: 711 DIAGGDTVERNVFLDAGRGRRLRYQIVPRSRTTGILTRLRSPVEAPVEPTESVADLRKYP 890
Query: 211 SVRSDGWLEIEMGEFFISGLEDEEVEMSVLQI-DGCFNVGLIIEGIEVRPK 260
R+DGWLE+E+GEFF G +D++V++ V ++ G + GL+++GIE+RPK
Sbjct: 891 KERADGWLEMELGEFFNEGGDDKQVDIGVCEVKGGGWKGGLVVQGIEIRPK 1043
>TC79882 weakly similar to PIR|E84434|E84434 probable phloem-specific lectin
[imported] - Arabidopsis thaliana, partial (14%)
Length = 636
Score = 218 bits (555), Expect = 2e-57
Identities = 116/189 (61%), Positives = 136/189 (71%), Gaps = 2/189 (1%)
Frame = +1
Query: 2 ELLPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRAN 61
E LPEGCI+AILSRTTP DA RLS+VSK SAADSDAVW++FLPSDSH I S +S AN
Sbjct: 73 EELPEGCISAILSRTTPADAGRLSVVSKTILSAADSDAVWNQFLPSDSHFIDS-IISLAN 249
Query: 62 ASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLP 121
+KK YL LSD P++IDN +KS QLD+KSGKKC+ML AR+L IAW N D Y W +P
Sbjct: 250 VPTKKSLYLALSDSPIIIDNDQKSIQLDRKSGKKCYMLAARSLSIAWGNDDRYWNWIAMP 429
Query: 122 ESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFH--EVLVVLSVNN 179
+SRFP+V L + WL I GMIN +LSPNTQY A+LVFK+I F VVLS+
Sbjct: 430 DSRFPEVAELLVVCWLHISGMINMLALSPNTQYAAYLVFKMIGGFGFRNPNCPVVLSICV 609
Query: 180 IGGHNITKI 188
GGH TKI
Sbjct: 610 EGGHKSTKI 636
>BF643821 weakly similar to PIR|F84434|F844 lectin-like protein [imported] -
Arabidopsis thaliana, partial (14%)
Length = 584
Score = 189 bits (479), Expect(2) = 6e-49
Identities = 96/152 (63%), Positives = 114/152 (74%), Gaps = 3/152 (1%)
Frame = +2
Query: 2 ELLPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSH---SIVSQSLS 58
E L EGCI+AI+SRTTP DA RLS+VSK F SAADSDAVW++FLPSDSH SI+S S S
Sbjct: 86 EELSEGCISAIISRTTPADAGRLSVVSKTFLSAADSDAVWNQFLPSDSHFIDSIISHSPS 265
Query: 59 RANASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWT 118
AN SKK YL LSD P++IDN +KSFQLD+KSGKKC+ML AR+L IAW + Y W
Sbjct: 266 LANVPSKKALYLALSDLPIIIDNDQKSFQLDRKSGKKCYMLAARSLSIAWGDDRRYWNWI 445
Query: 119 TLPESRFPDVVVLRHLLWLEIHGMINTFSLSP 150
++P SRFP+V L + WL+I GMINT L P
Sbjct: 446 SMPNSRFPEVAELLEVWWLDIRGMINTLGLVP 541
Score = 22.7 bits (47), Expect(2) = 6e-49
Identities = 8/12 (66%), Positives = 10/12 (82%)
Frame = +3
Query: 147 SLSPNTQYGAFL 158
+LSPNTQY F+
Sbjct: 531 ALSPNTQYAGFI 566
>TC90625 weakly similar to GP|10177856|dbj|BAB11208. phloem-specific
lectin-like protein {Arabidopsis thaliana}, partial (9%)
Length = 892
Score = 179 bits (455), Expect = 7e-46
Identities = 100/185 (54%), Positives = 123/185 (66%), Gaps = 4/185 (2%)
Frame = +2
Query: 1 MELLPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRA 60
+E+LPEGCIA ILS T PVD+ RLSLV K F SAA SD VWDRFLPSD SI+S S S +
Sbjct: 224 IEVLPEGCIAKILSHTAPVDSCRLSLVCKGFCSAAKSDTVWDRFLPSDLISIISDSPSAS 403
Query: 61 N----ASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRC 116
+ + SKK YL LSDHP+VI+NGKKSFQL+K+SG+K +ML AR + IA + +
Sbjct: 404 SLFSTSPSKKSLYLTLSDHPIVIENGKKSFQLEKQSGRKIYMLSARDISIALGDTPQFWD 583
Query: 117 WTTLPESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLS 176
W LPESRF +V LR + W G IN LS NTQY AFLV + F E+ + L+
Sbjct: 584 WPILPESRFREVARLRIVCWFAFEGTINKHVLSSNTQYAAFLVSR*SMPFGFDEIPIELT 763
Query: 177 VNNIG 181
V +G
Sbjct: 764 VGVLG 778
>TC80392 weakly similar to GP|15088626|gb|AAK84134.1 nictaba {Nicotiana
tabacum}, partial (29%)
Length = 1161
Score = 161 bits (407), Expect = 3e-40
Identities = 93/175 (53%), Positives = 119/175 (67%)
Frame = +2
Query: 89 DKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPESRFPDVVVLRHLLWLEIHGMINTFSL 148
DKK K+ ML AR+L IAW + Y W +P+SRFP+V L + WL+I G INT SL
Sbjct: 74 DKK--KESCMLSARSLSIAWGDDKRYWNWINMPDSRFPEVAQLLKVWWLDISGTINTLSL 247
Query: 149 SPNTQYGAFLVFKIIDAPNFHEVLVVLSVNNIGGHNITKIAYLNPDSLEGEVHDRWDRVQ 208
SPNTQY A+LVFK+I+A F + +SV GG + TKI L+P ++EG H+R +Q
Sbjct: 248 SPNTQYAAYLVFKMIEAEGFQNCPIEISVGVEGGQSGTKIDCLDP-NVEGGQHNRAVGLQ 424
Query: 209 GPSVRSDGWLEIEMGEFFISGLEDEEVEMSVLQIDGCFNVGLIIEGIEVRPKNDN 263
PSVRSDGWLEIEMGE F SGLED +V M+V + + + GL +EGIEVRPK D+
Sbjct: 425 RPSVRSDGWLEIEMGE-FNSGLEDGDVLMNVKETNN-WKSGLFVEGIEVRPK*DH 583
>AJ501129 weakly similar to GP|10177856|dbj phloem-specific lectin-like
protein {Arabidopsis thaliana}, partial (33%)
Length = 558
Score = 156 bits (394), Expect = 9e-39
Identities = 89/158 (56%), Positives = 110/158 (69%), Gaps = 4/158 (2%)
Frame = +2
Query: 24 LSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRAN----ASSKKRFYLDLSDHPVVI 79
LS+VS F SAA+SD VWDRFLPSD SIVS S S ++ + SKK YL LSDHP++I
Sbjct: 86 LSVVS--FCSAAESDIVWDRFLPSDLISIVSDSQSASSLFTTSPSKKSLYLTLSDHPILI 259
Query: 80 DNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPESRFPDVVVLRHLLWLEI 139
+NGK SFQL+K+SGKK +MLGAR L IAW + Y W LPESRF +V LR + W EI
Sbjct: 260 NNGKTSFQLEKQSGKKVYMLGARDLSIAWGDTPCYWDWIILPESRFQEVARLRTVCWFEI 439
Query: 140 HGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSV 177
G IN LS ++QY AFLVFK+I+A F ++ L+V
Sbjct: 440 SGTINKRVLSSDSQYVAFLVFKMINAYRFEDLPTKLTV 553
>TC84458 weakly similar to PIR|B96604|B96604 hypothetical protein F14G9.14
[imported] - Arabidopsis thaliana, partial (24%)
Length = 766
Score = 155 bits (393), Expect = 1e-38
Identities = 94/261 (36%), Positives = 140/261 (53%), Gaps = 1/261 (0%)
Frame = +1
Query: 1 MELLPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRA 60
ME PE C+A ILS T+P+D R+SL+S I +S ADSD VW++FLP + IVS+ +
Sbjct: 16 MEFSPEDCVAHILSFTSPIDVCRVSLISSIVKSMADSDNVWEKFLPHNYQGIVSRLVDPP 195
Query: 61 NASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTL 120
+ S K+ L P +ID+G K F ++K +GK C+ L AR L I + N + W +
Sbjct: 196 LSCSSKKELLARLCKPQLIDDGNKMFSIEKTTGKICYSLSARQLSITFGNNPLHWSWRQV 375
Query: 121 PESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVNNI 180
SRF +V LR + +LE G IN LSP T YGA+L K ++ ++L + +N
Sbjct: 376 QGSRFAEVAELRTICYLETIGSINNEMLSPKTMYGAYLKVKNVECAYGLDLLPCIR-SNC 552
Query: 181 GGHNITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISGLEDEEVEMSVL 240
+ I + D WLEIE+G F+ ++ +EV M +
Sbjct: 553 KRNGICNCEH-----------------------KDEWLEIELGSFYTEKVQVQEVRMCLK 663
Query: 241 QIDGC-FNVGLIIEGIEVRPK 260
+++G F LI++GIE+RPK
Sbjct: 664 EVNGVDFKGRLIVDGIELRPK 726
>BG588872 weakly similar to GP|21592660|gb phloem-specific lectin PP2-like
protein {Arabidopsis thaliana}, partial (55%)
Length = 655
Score = 121 bits (303), Expect = 3e-28
Identities = 66/152 (43%), Positives = 92/152 (60%), Gaps = 1/152 (0%)
Frame = +1
Query: 12 ILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLS-RANASSKKRFYL 70
ILS T+P+D R+SL+S + +S ADSD +W++FLP I+S+ + SSKK +
Sbjct: 19 ILSFTSPLDVCRVSLLSSVVQSMADSDFLWEKFLPHKYQEIISRLVDPPLPCSSKKELFA 198
Query: 71 DLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPESRFPDVVV 130
L P +ID+G K F ++K +GK C++L AR L I + N Y W + SRF +
Sbjct: 199 RLC-KPSLIDDGNKMFSIEKTTGKICYLLSARQLSITFGNTSLYWSWKQVQGSRFAEAAE 375
Query: 131 LRHLLWLEIHGMINTFSLSPNTQYGAFLVFKI 162
LR + WLEI G IN+ LSP T YGA+L KI
Sbjct: 376 LRTICWLEIKGSINSEMLSPKTMYGAYLKVKI 471
>TC91143 similar to PIR|T47557|T47557 hypothetical protein F8J2.170 -
Arabidopsis thaliana, partial (60%)
Length = 669
Score = 81.3 bits (199), Expect = 4e-16
Identities = 44/140 (31%), Positives = 70/140 (49%), Gaps = 2/140 (1%)
Frame = +1
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVS--QSLSRAN 61
+PE C+A + TP + L+ +++ FR AA +D+VW+ LPS+ ++ R
Sbjct: 157 IPENCVARVFLHLTPPEICNLARLNRAFRGAASADSVWESKLPSNYQHLLRLMPEEQRCR 336
Query: 62 ASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLP 121
SKK + S P+ D+G K LD+ +G+ C + A+ L I + Y W
Sbjct: 337 NLSKKDIFAVFSK-PLPFDDGNKQLWLDRVTGRVCMSISAKGLSITGIDDRRYWTWVPTE 513
Query: 122 ESRFPDVVVLRHLLWLEIHG 141
ESRF V L+ + W E+ G
Sbjct: 514 ESRFNIVAYLQQIWWFEVDG 573
>TC76820 weakly similar to GP|18175688|gb|AAL59911.1 unknown protein
{Arabidopsis thaliana}, partial (67%)
Length = 1388
Score = 78.6 bits (192), Expect = 2e-15
Identities = 72/265 (27%), Positives = 119/265 (44%), Gaps = 8/265 (3%)
Frame = +2
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
+PE CI++I+ R P + +L+ V+K F A+ D VW+ LP +V + L
Sbjct: 197 IPESCISSIMMRLDPQEICKLARVNKTFHRASSDDYVWESKLPPSYKFLVKKILGEEKLC 376
Query: 64 S--KKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLP 121
S KK Y L P D G K LD+ SG+ C + +++ I + Y + +
Sbjct: 377 SMTKKEIYAKLC-QPNFFDGGTKEILLDRCSGQVCLFISSKSFKITGIDDRRYWNYISTE 553
Query: 122 ESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKI-IDAPNFHEVLVVLSVNNI 180
ESRF V L+ + W+E+ G + P Y F FK+ + V ++ +
Sbjct: 554 ESRFKSVAYLQQMWWVEVLGELEL--EFPKGNYSIF--FKLQLGKTTKRLGRRVCNLEQV 721
Query: 181 GGHNITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFI---SGLEDEEVEM 237
G ++ + + S +G+ + GP W +G+F I +GL ++
Sbjct: 722 HGWDLKPVRFQLSTS-DGQNSLSQCYMNGPG----EWAYYHVGDFVIEKPNGL--ISIKF 880
Query: 238 SVLQIDGCFNV--GLIIEGIEVRPK 260
S+ QID C + GL I+G + PK
Sbjct: 881 SLAQID-CTHTKGGLCIDGAVICPK 952
>TC79148
Length = 553
Score = 76.6 bits (187), Expect = 9e-15
Identities = 40/60 (66%), Positives = 46/60 (76%), Gaps = 2/60 (3%)
Frame = +1
Query: 206 RVQGPSVRSDGWLEIEMGEFFISGLEDEEVEMSVLQI-DG-CFNVGLIIEGIEVRPKNDN 263
R+Q P +RSDGWLE EMGEFF SGLEDEEV+MS+ I DG + G +EGIEVRPK DN
Sbjct: 118 RLQQPRMRSDGWLETEMGEFFNSGLEDEEVQMSITGIKDGYTWKRGFFVEGIEVRPKEDN 297
>TC82508 similar to PIR|T50527|T50527 hypothetical protein T27I15_150 -
Arabidopsis thaliana, partial (71%)
Length = 1761
Score = 72.0 bits (175), Expect = 2e-13
Identities = 43/139 (30%), Positives = 67/139 (47%), Gaps = 6/139 (4%)
Frame = +2
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
+PE C+A +L P D +L+ +++ FR A+ +D VW+ LP + I+ ++L S
Sbjct: 719 IPESCVALVLMYLDPPDICKLARLNRAFRDASFADFVWESKLPLNYEFIMGKALEDDTTS 898
Query: 64 SKKRFYLDLSD------HPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCW 117
S L D P + DNG K LDK++G C + ++AL I + Y
Sbjct: 899 SSSGAELGKRDIYARLCKPNLFDNGTKEIWLDKRTGGVCLAISSKALRITGIDDRRYWNH 1078
Query: 118 TTLPESRFPDVVVLRHLLW 136
+ ESRF V L + W
Sbjct: 1079ISTEESRFHTVAYLHQIWW 1135
>BF520962 weakly similar to PIR|C96656|C96 unknown protein 43836-42510
[imported] - Arabidopsis thaliana, partial (58%)
Length = 681
Score = 72.0 bits (175), Expect = 2e-13
Identities = 46/145 (31%), Positives = 70/145 (47%), Gaps = 5/145 (3%)
Frame = +3
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LP+ CIA I+ P +L+ +++ F A+ +D VW+ LPS+ H I+ + ++
Sbjct: 180 LPQSCIAGIMEYMDPPQICKLASLNRAFHGASLADFVWESKLPSNYHVIIQKIFDGSSFP 359
Query: 64 S---KKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTL 120
S K+ Y L D G K LD+ GK C + A+ L I D R W +
Sbjct: 360 SHLGKRGIYSRLCSLN-TFDEGTKKVWLDRSMGKVCLSVSAKGLSIT--GIDDRRYWNHV 530
Query: 121 P--ESRFPDVVVLRHLLWLEIHGMI 143
P ESRF V L+ + W E+ G +
Sbjct: 531 PTEESRFSSVAYLQQIWWFEVDGEV 605
>AW694203 weakly similar to PIR|C96656|C96 unknown protein 43836-42510
[imported] - Arabidopsis thaliana, partial (52%)
Length = 621
Score = 65.9 bits (159), Expect = 2e-11
Identities = 38/134 (28%), Positives = 66/134 (48%), Gaps = 1/134 (0%)
Frame = +1
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPE C+A+I+ P +L+ +++ FR+A+ +D VW+ LP + H ++++ +
Sbjct: 172 LPESCVASIIGYMDPPQICQLATLNRAFRAASSADFVWESKLPPNYHLLLAKIFHHFPIN 351
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGA-RALFIAWDNGDHYRCWTTLPE 122
KR ID+G K +D+ +GK C + A + L + + Y + E
Sbjct: 352 LGKRDIYASLCRLNTIDDGTKKVWIDRATGKLCLAISANKGLSVLGVDDRRYWNYIITEE 531
Query: 123 SRFPDVVVLRHLLW 136
SRF V L+H W
Sbjct: 532 SRFNTVAYLQHTWW 573
>TC89219 weakly similar to PIR|C96656|C96656 unknown protein 43836-42510
[imported] - Arabidopsis thaliana, partial (46%)
Length = 764
Score = 48.9 bits (115), Expect(2) = 2e-11
Identities = 32/86 (37%), Positives = 46/86 (53%), Gaps = 3/86 (3%)
Frame = +3
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
+PE CI++IL P D +L+ V++ F A+ +D VW+ LPS + ++ L N S
Sbjct: 282 IPESCISSILMHLDPHDICKLARVNRAFHRASSADFVWESKLPSCYKFLANKFLGEENLS 461
Query: 64 --SKKRFYLDLSDHPVVIDNG-KKSF 86
SKK Y L P D G K+SF
Sbjct: 462 TMSKKDVYTKLC-QPNRFDGGTKRSF 536
Score = 36.6 bits (83), Expect(2) = 2e-11
Identities = 18/58 (31%), Positives = 30/58 (51%)
Frame = +1
Query: 84 KSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPESRFPDVVVLRHLLWLEIHG 141
K LDK SG+ C + +++L I + Y + ESR +V L+ + W+E+ G
Sbjct: 526 KEVLLDKYSGQVCLFMSSKSLKITGIDDRRYWIYIPTEESRIKNVAYLQQMWWVEVIG 699
>TC82238 weakly similar to GP|8570046|dbj|BAA96751.1 Similar to Arabidopsis
thaliana chromosome4 BAC clone T16H5; lectin like
protein (AL024486), partial (24%)
Length = 819
Score = 62.0 bits (149), Expect = 2e-10
Identities = 44/168 (26%), Positives = 75/168 (44%), Gaps = 2/168 (1%)
Frame = +3
Query: 77 VVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPE--SRFPDVVVLRHL 134
V + + K + ++K S CFML ARAL I W +Y W + +V L+ +
Sbjct: 369 VFLHHKTKKYWVEKNSKANCFMLYARALSITWAEDPNYWKWIQQKDVSEGTTEVAELKRV 548
Query: 135 LWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVNNIGGHNITKIAYLNPD 194
WLE+HG +T LSP Y + + D E+ V + + GG + +
Sbjct: 549 CWLEVHGKFDTRKLSPGILYQVSFIIMLKDPAQGWELPVNVRLVLPGGKK-----QQHKE 713
Query: 195 SLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISGLEDEEVEMSVLQI 242
+L ++ R W+E+ +GEF +S + E+E+S+ +
Sbjct: 714 NLMEKLRAR-------------WIEVPVGEFVVSEKDGGEMEISMFDM 818
>CB891284 weakly similar to GP|22122920|gb unknown protein {Oryza sativa
(japonica cultivar-group)}, partial (37%)
Length = 588
Score = 40.0 bits (92), Expect = 0.001
Identities = 19/62 (30%), Positives = 32/62 (50%)
Frame = +2
Query: 80 DNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPESRFPDVVVLRHLLWLEI 139
D G K LD+ SG+ C + +++ I + Y + + ESRF V L+ + W+E+
Sbjct: 236 DGGTKEILLDRCSGQVCLFISSKSFKITGIDDRRYWNYISTEESRFKSVAYLQQMWWVEV 415
Query: 140 HG 141
G
Sbjct: 416 LG 421
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.138 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,520,507
Number of Sequences: 36976
Number of extensions: 118620
Number of successful extensions: 548
Number of sequences better than 10.0: 52
Number of HSP's better than 10.0 without gapping: 521
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 523
length of query: 263
length of database: 9,014,727
effective HSP length: 94
effective length of query: 169
effective length of database: 5,538,983
effective search space: 936088127
effective search space used: 936088127
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0226.6


