

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
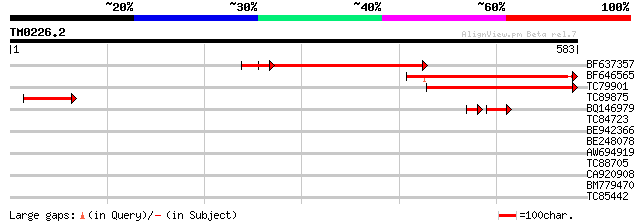
Query= TM0226.2
(583 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BF637357 similar to GP|13365578|db contains ESTs AU088719(S4716)... 263 1e-70
BF646565 weakly similar to GP|20259245|gb putative N-terminal ac... 263 2e-70
TC79901 weakly similar to GP|20259245|gb|AAM14358.1 putative N-t... 218 7e-57
TC89875 similar to GP|20259245|gb|AAM14358.1 putative N-terminal... 97 2e-20
BQ146979 weakly similar to GP|20259245|gb| putative N-terminal a... 50 3e-08
TC84723 similar to GP|16945387|emb|CAD11800. conserved hypotheti... 35 0.061
BE942366 similar to GP|7939524|dbj| contains similarity to unkno... 30 3.4
BE248078 similar to GP|14334774|gb| unknown protein {Arabidopsis... 30 3.4
AW694919 similar to GP|21554403|gb transcription factor TINY pu... 30 3.4
TC88705 similar to GP|10177834|dbj|BAB11263. gene_id:MCD7.9~unkn... 29 4.4
CA920908 homologue to PIR|T06377|T06 SAR DNA-binding protein-1 -... 29 5.7
BM779470 similar to GP|17225592|gb serine acetyltransferase {Ara... 28 7.5
TC85442 similar to GP|13543783|gb|AAH06040.1 Unknown (protein fo... 28 9.8
>BF637357 similar to GP|13365578|db contains ESTs AU088719(S4716)
C98685(E1389) AU066116(S4716)~hypothetical
protein~similar to, partial (21%)
Length = 582
Score = 263 bits (672), Expect = 1e-70
Identities = 134/173 (77%), Positives = 143/173 (82%)
Frame = +2
Query: 257 LEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSPPKSTAEEDEEMSKLLPAQKKKMRQKQ 316
L L F+ SH YFHKAAAGAIRCYIKLHD PPKST EEDE MS LLP+QKKK+RQKQ
Sbjct: 62 LTCLNFKTNCTSHPYFHKAAAGAIRCYIKLHDFPPKSTTEEDEHMSNLLPSQKKKLRQKQ 241
Query: 317 RKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPHGEKLLQVDDPLSEAIKYLKLL 376
RKAEARAKK AEEKNEEL++S VSKSGKR VKPVDPDPHGEKLLQV+DPLSEA+KYLKLL
Sbjct: 242 RKAEARAKKEAEEKNEELNSSVVSKSGKRPVKPVDPDPHGEKLLQVEDPLSEAVKYLKLL 421
Query: 377 QKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRLDAEHPDSHRCLVCQLH 429
QKNSPDSLETHLLSFELYTRK K+LL FQAV PDSHRCL+ H
Sbjct: 422 QKNSPDSLETHLLSFELYTRKXKILLAFQAVSSY*GWMLTTPDSHRCLIKFFH 580
Score = 63.9 bits (154), Expect = 2e-10
Identities = 28/34 (82%), Positives = 31/34 (90%)
Frame = +1
Query: 239 DQFDFHSYCLRKMTLRTYLEMLKFQDQLHSHSYF 272
DQFDFHSYCLRKMTLR+Y++MLKFQDQLH S F
Sbjct: 7 DQFDFHSYCLRKMTLRSYVDMLKFQDQLHLTSLF 108
>BF646565 weakly similar to GP|20259245|gb putative N-terminal
acetyltransferase {Arabidopsis thaliana}, partial (19%)
Length = 614
Score = 263 bits (671), Expect = 2e-70
Identities = 138/206 (66%), Positives = 151/206 (72%), Gaps = 31/206 (15%)
Frame = +1
Query: 409 QLLRLDAEHPDSHRCLV-------------------------------CQLHEKTLFEAN 437
QLLRLDA+HPDSHRCL+ QLHEK+LFEAN
Sbjct: 1 QLLRLDADHPDSHRCLIKFFHQLGSMSTPVTESEKLIWSVLEAERSTISQLHEKSLFEAN 180
Query: 438 NSFLEKHKDSLMHRAAFAETLYILDPNRKSEAVKLIEESTNNIVPRNGALGPIREWKLKD 497
N+F + HKDSLMHRAAFAE LYILD NRKSEAVKLIE+S NN VPRNGA+GPI EWKL+D
Sbjct: 181 NAFHDNHKDSLMHRAAFAEILYILDSNRKSEAVKLIEDSVNNTVPRNGAIGPIGEWKLED 360
Query: 498 CIAVHKLLGTVLLDQDAALRWKVRCAEYFPYSRYFEGSRSSASSNTALKQLSKNSENETL 557
CIAVHKLLGTVL+DQDAALRWKVRCAEYFPYS YFEG SSAS N+A QL KNSEN+
Sbjct: 361 CIAVHKLLGTVLVDQDAALRWKVRCAEYFPYSTYFEGRHSSASPNSAFSQLRKNSENDGP 540
Query: 558 NHSVCTQNVGSITSNGKLEAFKQLAI 583
NHSV QNVGS TSNG+ AF+ L I
Sbjct: 541 NHSVDNQNVGSTTSNGR--AFENLTI 612
>TC79901 weakly similar to GP|20259245|gb|AAM14358.1 putative N-terminal
acetyltransferase {Arabidopsis thaliana}, partial (11%)
Length = 965
Score = 218 bits (554), Expect = 7e-57
Identities = 109/155 (70%), Positives = 126/155 (80%)
Frame = +1
Query: 429 HEKTLFEANNSFLEKHKDSLMHRAAFAETLYILDPNRKSEAVKLIEESTNNIVPRNGALG 488
H K+L EAN+ FLEKH+ S+MHRAAF E +YILDPNR++EAVKLIE STNN V NGALG
Sbjct: 34 HGKSLLEANSLFLEKHEGSMMHRAAFGEMMYILDPNRRAEAVKLIEGSTNNPVSSNGALG 213
Query: 489 PIREWKLKDCIAVHKLLGTVLLDQDAALRWKVRCAEYFPYSRYFEGSRSSASSNTALKQL 548
PIREW LKDCIAVHKLLG+VL DQDAALRWKVRCAE+FPYS YFEGS+SSAS N+AL Q+
Sbjct: 214 PIREWTLKDCIAVHKLLGSVLDDQDAALRWKVRCAEFFPYSTYFEGSQSSASPNSALNQI 393
Query: 549 SKNSENETLNHSVCTQNVGSITSNGKLEAFKQLAI 583
K + N + +HS V S+TSNGKL +FK L I
Sbjct: 394 CKTTINGSSSHSPGDNIVESVTSNGKLASFKDLTI 498
>TC89875 similar to GP|20259245|gb|AAM14358.1 putative N-terminal
acetyltransferase {Arabidopsis thaliana}, partial (38%)
Length = 1170
Score = 96.7 bits (239), Expect = 2e-20
Identities = 47/54 (87%), Positives = 49/54 (90%)
Frame = +1
Query: 15 QRIPLDFLQGDKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGKADILEQLIL 68
+RIPLDFLQGDKFREAA+NYIRPLLTKGVPSLFSDLSSLY H GKADILE L
Sbjct: 1009 KRIPLDFLQGDKFREAAENYIRPLLTKGVPSLFSDLSSLYTHSGKADILEHSFL 1170
>BQ146979 weakly similar to GP|20259245|gb| putative N-terminal
acetyltransferase {Arabidopsis thaliana}, partial (3%)
Length = 644
Score = 49.7 bits (117), Expect(2) = 3e-08
Identities = 23/26 (88%), Positives = 24/26 (91%)
Frame = +3
Query: 491 REWKLKDCIAVHKLLGTVLLDQDAAL 516
REW LKDCIAVHKLLG+VL DQDAAL
Sbjct: 66 REWTLKDCIAVHKLLGSVLDDQDAAL 143
Score = 26.2 bits (56), Expect(2) = 3e-08
Identities = 13/17 (76%), Positives = 13/17 (76%)
Frame = +1
Query: 470 VKLIEESTNNIVPRNGA 486
VKLIE STNN V NGA
Sbjct: 1 VKLIEGSTNNPVSSNGA 51
>TC84723 similar to GP|16945387|emb|CAD11800. conserved hypothetical protein
{Neurospora crassa}, partial (5%)
Length = 999
Score = 35.4 bits (80), Expect = 0.061
Identities = 30/131 (22%), Positives = 57/131 (42%), Gaps = 3/131 (2%)
Frame = +3
Query: 206 ELASGESFFRQGDLGRALKKFLGVEKHYADINEDQFDFHSYCLRKMTLRTYLEMLKFQDQ 265
EL + + + +L RA + + A I+E++ L++ +E+ K+Q++
Sbjct: 561 ELRVQKEYREEAELQRAKRDLDEIRNREARISEEKH-----------LKSQVELQKYQEE 707
Query: 266 LH---SHSYFHKAAAGAIRCYIKLHDSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEAR 322
+ K AA A+ + K +E +E K + ++ RQK+ A+
Sbjct: 708 MRLAEKAKQREKEAADAVERFQKKEQERLAKQEKEKQEREKEFQRRLQEERQKELDRIAK 887
Query: 323 AKKGAEEKNEE 333
AKK EE +E
Sbjct: 888 AKKEKEENEKE 920
>BE942366 similar to GP|7939524|dbj| contains similarity to unknown
protein~gb|AAD25764.1~gene_id:K14B15.5 {Arabidopsis
thaliana}, partial (12%)
Length = 391
Score = 29.6 bits (65), Expect = 3.4
Identities = 21/75 (28%), Positives = 33/75 (44%), Gaps = 4/75 (5%)
Frame = +3
Query: 456 ETLYILDPNRKSEAVKLIEESTNNIV----PRNGALGPIREWKLKDCIAVHKLLGTVLLD 511
+TLY D + EA+ + + N I +N LG + L+DC+ G LLD
Sbjct: 39 QTLYFADKIKTEEAICELLKGLNYICRYEQQQNALLGCASSFNLEDCMKWKLQCGDSLLD 218
Query: 512 QDAALRWKVRCAEYF 526
+ +C E+F
Sbjct: 219 *THSRYCNPQCGEHF 263
>BE248078 similar to GP|14334774|gb| unknown protein {Arabidopsis thaliana},
partial (24%)
Length = 392
Score = 29.6 bits (65), Expect = 3.4
Identities = 22/72 (30%), Positives = 34/72 (46%), Gaps = 6/72 (8%)
Frame = +3
Query: 285 KLHDSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEA------RAKKGAEEKNEELSASG 338
K+HD K E++ K+ QKK++R+ +R + +AK A K +ELS S
Sbjct: 120 KIHDLNSKL-----EDVQKINEEQKKQIRKTERALKVAEEEMLKAKLEATTKAKELSESV 284
Query: 339 VSKSGKRHVKPV 350
H KP+
Sbjct: 285 AESHWNEHGKPL 320
>AW694919 similar to GP|21554403|gb transcription factor TINY putative
{Arabidopsis thaliana}, partial (53%)
Length = 570
Score = 29.6 bits (65), Expect = 3.4
Identities = 23/96 (23%), Positives = 37/96 (37%)
Frame = -2
Query: 267 HSHSYFHKAAAGAIRCYIKLHDSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKG 326
H H ++H+ CY P + + L P Q + Q + AE G
Sbjct: 347 HHHHHYHQN-----HCYY------PPAKQQP*PSYELL*PPQLEYP*QYTKTAEEAPADG 201
Query: 327 AEEKNEELSASGVSKSGKRHVKPVDPDPHGEKLLQV 362
A + +S + ++H KP+ P P EK L +
Sbjct: 200 ALHQENSAVLYNLSNNMQQHHKPLQPSPAPEKTLAI 93
>TC88705 similar to GP|10177834|dbj|BAB11263. gene_id:MCD7.9~unknown protein
{Arabidopsis thaliana}, partial (32%)
Length = 1685
Score = 29.3 bits (64), Expect = 4.4
Identities = 23/87 (26%), Positives = 39/87 (44%), Gaps = 19/87 (21%)
Frame = +1
Query: 292 KSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHV---- 347
K ++ EE +L QK+K +++++AE +A + + NEE +G+ ++H
Sbjct: 34 KEQIDKAEEKERL---QKEKEEKQKKEAEEKANEKQVKTNEE--DTGIENEAEKHSDIED 198
Query: 348 ---------------KPVDPDPHGEKL 359
PVD D GEKL
Sbjct: 199 NFAASIHEKIEVKEDSPVDQDEAGEKL 279
>CA920908 homologue to PIR|T06377|T06 SAR DNA-binding protein-1 - garden pea,
partial (37%)
Length = 748
Score = 28.9 bits (63), Expect = 5.7
Identities = 12/40 (30%), Positives = 25/40 (62%)
Frame = -2
Query: 292 KSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKN 331
KS + DE+ +L A KKK +++++K E + ++ ++ N
Sbjct: 198 KSDSAMDEDTEELSAADKKKEKKEKKKKEKKEEEDTQKSN 79
>BM779470 similar to GP|17225592|gb serine acetyltransferase {Arabidopsis
thaliana}, partial (45%)
Length = 810
Score = 28.5 bits (62), Expect = 7.5
Identities = 19/60 (31%), Positives = 32/60 (52%)
Frame = +1
Query: 352 PDPHGEKLLQVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLL 411
PDP E +L V DP+ EA+K Q+ ++ + +LS LY +L + ++Q+L
Sbjct: 286 PDPDPESILDVSDPIWEAVK-----QEAKHEAEKEPVLSSFLYAS----VLAHECLEQVL 438
>TC85442 similar to GP|13543783|gb|AAH06040.1 Unknown (protein for MGC:7642)
{Mus musculus}, partial (62%)
Length = 585
Score = 28.1 bits (61), Expect = 9.8
Identities = 11/25 (44%), Positives = 18/25 (72%)
Frame = +1
Query: 308 QKKKMRQKQRKAEARAKKGAEEKNE 332
+KKK R+K+++ + R KKG KN+
Sbjct: 100 RKKKKRKKKQRRKKRRKKGKRRKNQ 174
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.133 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,101,244
Number of Sequences: 36976
Number of extensions: 233493
Number of successful extensions: 1265
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 1247
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1263
length of query: 583
length of database: 9,014,727
effective HSP length: 101
effective length of query: 482
effective length of database: 5,280,151
effective search space: 2545032782
effective search space used: 2545032782
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0226.2


