

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.11
(231 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
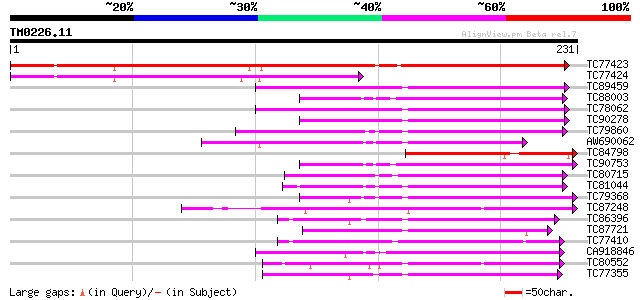
TC77423 weakly similar to PIR|D96569|D96569 probable oxidoreduct... 297 3e-81
TC77424 weakly similar to PIR|D96569|D96569 probable oxidoreduct... 152 1e-37
TC89459 weakly similar to GP|9280317|dbj|BAB01696.1 oxylase-like... 86 9e-18
TC88003 weakly similar to PIR|T04184|T04184 hypothetical protein... 76 1e-14
TC78062 weakly similar to GP|9280317|dbj|BAB01696.1 oxylase-like... 75 2e-14
TC90278 weakly similar to GP|9280318|dbj|BAB01697.1 oxidase-like... 74 5e-14
TC79860 homologue to PIR|T06533|T06533 probable gibberellin 20-o... 74 5e-14
AW690062 weakly similar to GP|7268917|emb gibberellin 20-oxidase... 73 8e-14
TC84798 weakly similar to PIR|B96569|B96569 probable oxidoreduct... 72 1e-13
TC90753 weakly similar to GP|21327037|gb|AAM48133.1 putative fla... 66 1e-11
TC80715 similar to GP|5579092|gb|AAD45424.1| gibberellin 2-oxida... 66 1e-11
TC81044 weakly similar to GP|10177853|dbj|BAB11205. flavanone 3-... 65 2e-11
TC79368 weakly similar to PIR|S44261|S44261 SRG1 protein - Arabi... 65 2e-11
TC87248 similar to GP|7594580|emb|CAB88075.1 hypothetical protei... 65 3e-11
TC86396 similar to GP|10178137|dbj|BAB11549. leucoanthocyanidin ... 64 5e-11
TC87721 weakly similar to PIR|H96594|H96594 hypothetical protein... 64 6e-11
TC77410 similar to GP|21392365|gb|AAM48289.1 flavanone 3 beta-hy... 63 8e-11
CA918846 weakly similar to GP|21537316|gb ethylene-forming-enzym... 63 8e-11
TC80552 similar to PIR|G86472|G86472 probable hyoscyamine 6-diox... 63 1e-10
TC77355 weakly similar to PIR|T05552|T05552 SRG1 protein-related... 63 1e-10
>TC77423 weakly similar to PIR|D96569|D96569 probable oxidoreductase
38288-39393 [imported] - Arabidopsis thaliana, partial
(87%)
Length = 1141
Score = 297 bits (760), Expect = 3e-81
Identities = 161/292 (55%), Positives = 191/292 (65%), Gaps = 64/292 (21%)
Frame = +1
Query: 1 MGSETTLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY------------------- 41
MG+E T KLP+IDF NLN L SPNWE VKS+V+KALVEY
Sbjct: 25 MGTEATPKLPIIDFNNLN-LEAKSPNWELVKSQVYKALVEYGCFEAVFDKVPLDLRKAIF 201
Query: 42 -------------MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN 88
LNVSKKPY GY GQ+P IPLF+SMGIDDAN+ EKV+S+T LWP+
Sbjct: 202 ASLQELFDLPLETKLLNVSKKPYHGYVGQYPMIPLFESMGIDDANILEKVDSMTNNLWPD 381
Query: 89 GNPSFRYA----SQPIT----------------------------YFLKTMKYKGPQTSD 116
GN +F S+ +T Y L+ MKYKGPQTSD
Sbjct: 382 GNQNFSKTIHSFSEELTELDQIIRKMILESLGVEKYLEEHMNSTNYLLRVMKYKGPQTSD 561
Query: 117 TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSN 176
TK+GL TH+D ++TIL+Q+QV+GLEV+TKDG+ WISYKPSS S SF+ M+ D HAWSN
Sbjct: 562 TKLGLSTHSDKNVVTILYQNQVEGLEVMTKDGQ-WISYKPSS-SGSFVAMIGDSIHAWSN 735
Query: 177 GRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFD 228
GRLHSPFHRVMM+GNEARYS LF +PKGG IIKAPEELVDEEHPLL++P+D
Sbjct: 736 GRLHSPFHRVMMSGNEARYSTGLFSIPKGGYIIKAPEELVDEEHPLLFRPYD 891
>TC77424 weakly similar to PIR|D96569|D96569 probable oxidoreductase
38288-39393 [imported] - Arabidopsis thaliana, partial
(54%)
Length = 670
Score = 152 bits (383), Expect = 1e-37
Identities = 90/208 (43%), Positives = 114/208 (54%), Gaps = 64/208 (30%)
Frame = +2
Query: 1 MGSETTLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY------------------- 41
MGSE T KLP+IDF NLN +PNWE VKS+V+KALVEY
Sbjct: 50 MGSEHTQKLPLIDFNNLN-FEAKNPNWELVKSQVYKALVEYGCFEAIFDKVPLDLRKAIF 226
Query: 42 -------------MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN 88
LNVS KPY GY GQ+P +PL++SM I+DAN+ EKV+S+T ILWP+
Sbjct: 227 TSIEELFDLPLETKLLNVSNKPYDGYIGQYPVVPLYESMAIEDANILEKVKSMTNILWPD 406
Query: 89 GNPSF-----------------------------RYASQPI---TYFLKTMKYKGPQTSD 116
GN +F +Y + + Y L+ MKYKGPQTSD
Sbjct: 407 GNKNFSETIHSFSEELIELDQIIRKMILESLGVEKYLEEHMNSTNYRLRAMKYKGPQTSD 586
Query: 117 TKVGLDTHTDTAILTILFQSQVDGLEVL 144
TKVGL HTD +TIL+Q+QV+GLE++
Sbjct: 587 TKVGLTPHTDKNFVTILYQNQVEGLEMM 670
>TC89459 weakly similar to GP|9280317|dbj|BAB01696.1 oxylase-like protein
{Arabidopsis thaliana}, partial (70%)
Length = 1302
Score = 86.3 bits (212), Expect = 9e-18
Identities = 44/128 (34%), Positives = 68/128 (52%)
Frame = +3
Query: 101 TYFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSES 160
T F++ Y +G+ H D LTIL Q +V GLEV K ++W+ KP+ ++
Sbjct: 651 TSFIRFNHYPPCPYPHLALGVGRHKDAGALTILAQDEVGGLEVKRKSDQQWVLVKPTPDA 830
Query: 161 QSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEH 220
+I+ + D WSN S HRVM+ + R+S FF P ++K EEL +EE+
Sbjct: 831 --YIINVGDIIQVWSNDAYESVEHRVMVNSEKERFSIPFFFFPAHDTVVKPLEELTNEEN 1004
Query: 221 PLLYKPFD 228
P Y+P++
Sbjct: 1005PPKYRPYN 1028
>TC88003 weakly similar to PIR|T04184|T04184 hypothetical protein F7L13.70 -
Arabidopsis thaliana, partial (26%)
Length = 1209
Score = 75.9 bits (185), Expect = 1e-14
Identities = 45/109 (41%), Positives = 61/109 (55%)
Frame = +2
Query: 119 VGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGR 178
+GL+ HTD LT+L QS+V GL+V KDG KWIS +F++ + D SNGR
Sbjct: 662 MGLNEHTDLNALTVLLQSEVSGLQV-NKDG-KWISIP--CIPNAFVINLADQIEVLSNGR 829
Query: 179 LHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPF 227
S HR + + R S A+F+ P II EL+DEEHP Y+ +
Sbjct: 830 YKSVLHRAVTNNVQPRISMAMFYGPNPETIIGPIHELIDEEHPPKYRNY 976
>TC78062 weakly similar to GP|9280317|dbj|BAB01696.1 oxylase-like protein
{Arabidopsis thaliana}, partial (80%)
Length = 1322
Score = 75.1 bits (183), Expect = 2e-14
Identities = 39/128 (30%), Positives = 65/128 (50%)
Frame = +3
Query: 101 TYFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSES 160
T +++ Y D +G H D+ LT+L Q +V GLEV K +W+ KP +
Sbjct: 681 TSWIRLNHYPPCPNPDIVLGCGRHKDSGALTVLAQDEVSGLEVRRKSDGEWVLVKPLPNA 860
Query: 161 QSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEH 220
+I+ + D WSN S HRVM+ +AR S F P +++ +EL ++++
Sbjct: 861 --YIINVGDVIQVWSNDEYKSVEHRVMLNTEKARLSYPFFLFPSHYTMVEPLKELTNDQN 1034
Query: 221 PLLYKPFD 228
P Y+P++
Sbjct: 1035PPKYRPYN 1058
>TC90278 weakly similar to GP|9280318|dbj|BAB01697.1 oxidase-like protein
{Arabidopsis thaliana}, partial (21%)
Length = 1139
Score = 73.9 bits (180), Expect = 5e-14
Identities = 39/110 (35%), Positives = 61/110 (55%)
Frame = +2
Query: 119 VGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGR 178
+G+ H D LTIL Q +V GLEV + ++W+ KP+ + FI+ + D WSN
Sbjct: 698 LGMGRHKDAGALTILAQDEVAGLEVKHQISQEWVLVKPTPNA--FIINVGDLTQVWSNDE 871
Query: 179 LHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFD 228
S HRV++ + R+S A P +K +EL+++E+P YKPF+
Sbjct: 872 YESVEHRVIVNFEKERFSIAFGVFPAHYIEVKPLDELINDENPPKYKPFN 1021
>TC79860 homologue to PIR|T06533|T06533 probable gibberellin 20-oxidase -
garden pea, partial (93%)
Length = 1454
Score = 73.9 bits (180), Expect = 5e-14
Identities = 47/135 (34%), Positives = 69/135 (50%)
Frame = +3
Query: 93 FRYASQPITYFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWI 152
FR+ + ++ Y + D +G H D LTIL Q QV+GL+VL DG W
Sbjct: 648 FRHFFEANDSVMRLNYYPPCKNPDLALGTGPHCDPTSLTILHQDQVEGLQVLV-DGI-WH 821
Query: 153 SYKPSSESQSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAP 212
S P ++ F+V + D F A SNG S HR ++ R S A F P I+ P
Sbjct: 822 SIVPKEDA--FVVNIGDTFMALSNGIFKSCLHRAVVNDTIVRKSLAFFLCPNEEKIVTPP 995
Query: 213 EELVDEEHPLLYKPF 227
+EL+++E+P++Y F
Sbjct: 996 KELINKENPMIYPNF 1040
>AW690062 weakly similar to GP|7268917|emb gibberellin 20-oxidase-like
protein {Arabidopsis thaliana}, partial (44%)
Length = 629
Score = 73.2 bits (178), Expect = 8e-14
Identities = 50/136 (36%), Positives = 73/136 (52%), Gaps = 3/136 (2%)
Frame = +1
Query: 79 ESLTKILWPNGNPSFRYASQPI---TYFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQ 135
E+L +IL N +F Y Q T +L+ +Y +GL H DT+ +TI+ Q
Sbjct: 133 ENLVQILAQELNINFSYFQQNCSANTSYLRLNRYPPCPFPSKVIGLLPHADTSFITIVHQ 312
Query: 136 SQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARY 195
+ GL+++ KDG KWIS KP+SE+ IV + D F A SNG S HRV+ R+
Sbjct: 313 DHIGGLQLM-KDG-KWISVKPNSEA--LIVNVGDLFQALSNGLYTSVGHRVVAAEKVERF 480
Query: 196 SAALFFVPKGGCIIKA 211
S A F+ P +I++
Sbjct: 481 SLAYFYGPSIDAVIES 528
>TC84798 weakly similar to PIR|B96569|B96569 probable oxidoreductase
33116-34434 [imported] - Arabidopsis thaliana, partial
(20%)
Length = 545
Score = 72.4 bits (176), Expect = 1e-13
Identities = 39/72 (54%), Positives = 49/72 (67%), Gaps = 2/72 (2%)
Frame = +2
Query: 162 SFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALF-FVPKGGCIIKAPEELVDEEH 220
SF+V+ D F WSNGR+ + HRV+M + RY+AALF F K I++ PEELV+EEH
Sbjct: 17 SFVVLAGDAFTIWSNGRIRACEHRVIMNATKTRYAAALFSFASK---IMEIPEELVNEEH 187
Query: 221 PLLYKP-FDRED 231
PL YKP FD D
Sbjct: 188 PLRYKPTFDHYD 223
>TC90753 weakly similar to GP|21327037|gb|AAM48133.1 putative flavanone
3-hydroxylase {Saussurea medusa}, partial (24%)
Length = 625
Score = 66.2 bits (160), Expect = 1e-11
Identities = 36/113 (31%), Positives = 62/113 (54%)
Frame = +1
Query: 119 VGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGR 178
+G+ H+D ++TIL Q V GL+V KDG +WI+ + + +F++ + SNG+
Sbjct: 40 LGISKHSDPNLITILLQDDVFGLQVF-KDG-EWIAVE--AFPHAFVINVGYQLQIISNGK 207
Query: 179 LHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
L S HR + + AR + A F P C ++ + + DE +P ++K F +D
Sbjct: 208 LKSAEHRAVTNSSHARTTGAFFVAPSDDCYVEPAQAITDEHNPPIFKSFKYKD 366
>TC80715 similar to GP|5579092|gb|AAD45424.1| gibberellin 2-oxidase-like
protein {Pisum sativum}, partial (97%)
Length = 1272
Score = 65.9 bits (159), Expect = 1e-11
Identities = 40/115 (34%), Positives = 56/115 (47%)
Frame = +2
Query: 113 QTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFH 172
Q ++ +G H+D ILTIL + V GL++ T+ G WI P + F VM+ D
Sbjct: 632 QNNNNTIGFGEHSDPQILTILRSNNVGGLQISTQHGL-WIPVHP--DPNEFYVMVGDSLQ 802
Query: 173 AWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPF 227
+NGR S HRV+ + R S F P I ++V +P LYKPF
Sbjct: 803 VLTNGRFVSVRHRVLTNTTKPRMSMMYFAAPPLNWWISPLSKMVTAHNPSLYKPF 967
>TC81044 weakly similar to GP|10177853|dbj|BAB11205. flavanone
3-hydroxylase-like protein {Arabidopsis thaliana},
partial (27%)
Length = 652
Score = 65.5 bits (158), Expect = 2e-11
Identities = 40/116 (34%), Positives = 62/116 (52%)
Frame = +3
Query: 112 PQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCF 171
P+ S T +GL H D ++TIL Q V GL+VL KDG WI+ +P + + F++ +
Sbjct: 132 PEPSLT-LGLPKHKDAYLITILMQDDVSGLQVL-KDGN-WINVEPLTHA--FVINIGHVL 296
Query: 172 HAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPF 227
S G+L S HR + +R SAA F P C+I+ ++L D+ + + F
Sbjct: 297 EIISKGKLKSAEHRAVTNSTYSRTSAAFFIAPSDDCLIEPAQDLPDDNNQPALRSF 464
>TC79368 weakly similar to PIR|S44261|S44261 SRG1 protein - Arabidopsis
thaliana, partial (48%)
Length = 1325
Score = 65.5 bits (158), Expect = 2e-11
Identities = 42/114 (36%), Positives = 63/114 (54%), Gaps = 1/114 (0%)
Frame = +1
Query: 119 VGLDTHTDTAILTILFQSQ-VDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNG 177
+GL+ H+D LTIL Q+ ++GL++ KDG+ WIS KP + + F+V + D +NG
Sbjct: 856 IGLNPHSDADCLTILLQANDIEGLQI-RKDGQ-WISVKPLAGA--FVVNIGDMLEILTNG 1023
Query: 178 RLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
S HR + +AR S A F P+ +I ++LV E P L+K ED
Sbjct: 1024IYRSIEHRATVNSEKARISIAAFHRPQMSKVIGPTQKLVTPERPALFKTITVED 1185
>TC87248 similar to GP|7594580|emb|CAB88075.1 hypothetical protein
{Arabidopsis thaliana}, partial (82%)
Length = 1271
Score = 64.7 bits (156), Expect = 3e-11
Identities = 45/163 (27%), Positives = 74/163 (44%), Gaps = 2/163 (1%)
Frame = +1
Query: 71 DANVYEKVESLTKILWPNGNPSFRYASQPITYFLKTMKYKGPQTSDTKV-GLDTHTDTAI 129
D N +EK+ +L K P FL+ ++Y G + ++ G H+D +
Sbjct: 508 DENYFEKISALNK---PEA-------------FLRLLRYPGELGPNEEICGASAHSDYGM 639
Query: 130 LTILFQSQVDGLEVLTKDGRKWISYKPSSESQ-SFIVMMNDCFHAWSNGRLHSPFHRVMM 188
+T+L + V GL++ + ++ ++ S + + IV + D W+N S HRVM
Sbjct: 640 ITLLLTNGVPGLQICKEKLKQPQVWEDVSHVEGAIIVNIGDMMERWTNCLYRSTLHRVMP 819
Query: 189 TGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
TG E RYS A F P C+++ E E P + P D
Sbjct: 820 TGKE-RYSVAFFMDPPSDCVVECFESCCSESSPPRFPPIRSGD 945
>TC86396 similar to GP|10178137|dbj|BAB11549. leucoanthocyanidin
dioxygenase-like protein {Arabidopsis thaliana}, partial
(82%)
Length = 1326
Score = 63.9 bits (154), Expect = 5e-11
Identities = 43/116 (37%), Positives = 61/116 (52%), Gaps = 1/116 (0%)
Frame = +2
Query: 110 KGPQTSDTKVGLDTHTDTAILTILFQSQ-VDGLEVLTKDGRKWISYKPSSESQSFIVMMN 168
K PQ D +GL H+D +TIL V GL+V + G WI+ KP + FI+ +
Sbjct: 764 KCPQP-DLTLGLSPHSDPGGMTILLPDDFVSGLQV--RKGNDWITVKPVPNA--FIINIG 928
Query: 169 DCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLY 224
D SN S HRV++ + R S A+F+ PK +I+ +ELV +E P LY
Sbjct: 929 DQIQVLSNAIYKSVEHRVIVNSIKDRVSLAMFYNPKSDLLIEPAKELVTKERPALY 1096
>TC87721 weakly similar to PIR|H96594|H96594 hypothetical protein F7A10.24
[imported] - Arabidopsis thaliana, partial (50%)
Length = 1365
Score = 63.5 bits (153), Expect = 6e-11
Identities = 36/103 (34%), Positives = 54/103 (51%), Gaps = 1/103 (0%)
Frame = +1
Query: 120 GLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRL 179
G+ H+D + +T+L Q + GL V KDG WI+ P + + ++ + D SNGR
Sbjct: 868 GVGPHSDISSITVLLQDDIGGLYVRGKDGDSWINVPPVNGA--LVINIGDVLQIMSNGRY 1041
Query: 180 HSPFHRVMMTGNEARYSAALFFVPKGGCII-KAPEELVDEEHP 221
S HRV++ GN+ R S +F P +I PE L + E P
Sbjct: 1042KSIEHRVVVDGNKTRISMPIFVNPAPDAVIGTLPEVLENGEEP 1170
>TC77410 similar to GP|21392365|gb|AAM48289.1 flavanone 3 beta-hydroxylase
{Solanum tuberosum}, partial (98%)
Length = 1471
Score = 63.2 bits (152), Expect = 8e-11
Identities = 39/117 (33%), Positives = 61/117 (51%)
Frame = +1
Query: 110 KGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMND 169
K PQ D +GL HTD +T+L Q QV GL+ +G+ WI+ +P +F+V + D
Sbjct: 670 KCPQP-DLTLGLKRHTDPGTITLLLQDQVGGLQATKDNGKTWITVQP--VEGAFVVNLGD 840
Query: 170 CFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKP 226
H SNGR + H+ ++ N +R S A F P + P ++ D E ++ +P
Sbjct: 841 HGHYLSNGRFKNADHQAVVNSNYSRLSIATFQNPAPDATV-YPLKIRDGEKSVMEEP 1008
>CA918846 weakly similar to GP|21537316|gb ethylene-forming-enzyme-like
dioxygenase-like {Arabidopsis thaliana}, partial (44%)
Length = 721
Score = 63.2 bits (152), Expect = 8e-11
Identities = 41/127 (32%), Positives = 68/127 (53%), Gaps = 1/127 (0%)
Frame = -3
Query: 101 TYFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQ-SQVDGLEVLTKDGRKWISYKPSSE 159
T ++T Y +D +GL H+D++ +T L Q ++V+GL+VL KD + W +K
Sbjct: 578 TMSMRTNYYPPCPMADHALGLKPHSDSSSITFLLQDNKVEGLQVL-KDNQ-W--FKVPII 411
Query: 160 SQSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEE 219
+ ++ + D SNG SP HR ++ +AR S A+ P I+ ++LV+E
Sbjct: 410 HDAIVINVGDLMEIMSNGIFQSPIHRAVVNEEKARLSVAMLCRPDSEKEIQPIDKLVNES 231
Query: 220 HPLLYKP 226
PL Y+P
Sbjct: 230 RPLSYRP 210
>TC80552 similar to PIR|G86472|G86472 probable hyoscyamine 6-dioxygenase
hydroxylase [imported] - Arabidopsis thaliana, partial
(92%)
Length = 1307
Score = 62.8 bits (151), Expect = 1e-10
Identities = 43/129 (33%), Positives = 61/129 (46%), Gaps = 6/129 (4%)
Frame = +3
Query: 104 LKTMKYKGPQTSDTKVGL---DTHTDTAILTILFQSQVDGLEVLT-KDGR--KWISYKPS 157
L+ + Y G Q SD GL HTD ++T+L V GL++ +D + KW P
Sbjct: 519 LRLLHYGG-QVSDPLKGLFGAGAHTDYGLITLLATDHVSGLQICKDRDAKPQKWEDVAPL 695
Query: 158 SESQSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVD 217
+ F+V + D WSNG S HRV+ G E RYS A F P C+++
Sbjct: 696 KGA--FVVNLGDMLERWSNGVFKSTLHRVLGNGQE-RYSIAYFLEPSHDCLVECLPTCKS 866
Query: 218 EEHPLLYKP 226
+ +P Y P
Sbjct: 867 DTNPPKYPP 893
>TC77355 weakly similar to PIR|T05552|T05552 SRG1 protein-related protein
F24A6.150 - Arabidopsis thaliana, partial (44%)
Length = 1359
Score = 62.8 bits (151), Expect = 1e-10
Identities = 37/123 (30%), Positives = 62/123 (50%), Gaps = 1/123 (0%)
Frame = +1
Query: 104 LKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQ-VDGLEVLTKDGRKWISYKPSSESQS 162
L+ Y + +GL H+DT +T+L Q V GLE+ K W+ KP S++
Sbjct: 667 LRVNYYPPCNNPEQVLGLSPHSDTTTITLLIQDDDVLGLEIRNKGN--WVPVKPISDA-- 834
Query: 163 FIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPL 222
++ + D SNG+ S HRVM N+ R S A F P+ ++ + ++D ++P
Sbjct: 835 LVINVGDVIEILSNGKYKSVEHRVMTNQNKRRISYASFLFPRDDVEVEPFDHMIDAQNPK 1014
Query: 223 LYK 225
+Y+
Sbjct: 1015MYQ 1023
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.135 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,289,296
Number of Sequences: 36976
Number of extensions: 100210
Number of successful extensions: 610
Number of sequences better than 10.0: 102
Number of HSP's better than 10.0 without gapping: 552
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 558
length of query: 231
length of database: 9,014,727
effective HSP length: 93
effective length of query: 138
effective length of database: 5,575,959
effective search space: 769482342
effective search space used: 769482342
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0226.11


