

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0225.5
(295 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
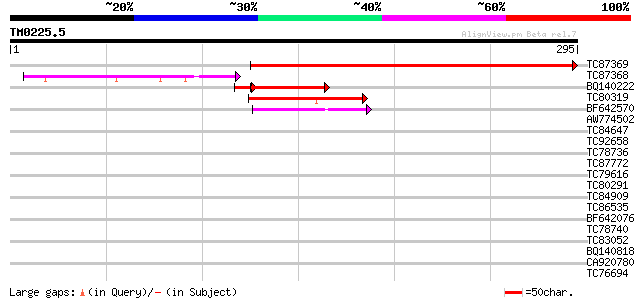
TC87369 homologue to GP|14517500|gb|AAK62640.1 K16N12.18/K16N12.... 313 7e-86
TC87368 homologue to GP|14517500|gb|AAK62640.1 K16N12.18/K16N12.... 102 2e-22
BQ140222 homologue to GP|14517500|gb K16N12.18/K16N12.18 {Arabid... 77 4e-15
TC80319 similar to GP|20259261|gb|AAM14366.1 putative HhoA prote... 70 1e-12
BF642570 similar to GP|13172275|gb DegP2 protease {Arabidopsis t... 49 3e-06
AW774502 similar to GP|17428071|emb PROBABLE PERIPLASMIC PROTEAS... 37 0.007
TC84647 similar to GP|9294054|dbj|BAB02011.1 contains similarity... 32 0.38
TC92658 similar to GP|19310429|gb|AAL84951.1 AT3g03380/T21P5_20 ... 30 0.85
TC78736 similar to GP|21553515|gb|AAM62608.1 unknown {Arabidopsi... 30 0.85
TC87772 weakly similar to GP|8574455|gb|AAF77578.1| pepper ester... 30 1.1
TC79616 similar to GP|21553745|gb|AAM62838.1 Yippee-like protein... 30 1.1
TC80291 similar to GP|19310429|gb|AAL84951.1 AT3g03380/T21P5_20 ... 30 1.1
TC84909 similar to GP|3395431|gb|AAC28763.1| unknown protein {Ar... 30 1.5
TC86535 similar to GP|9758076|dbj|BAB08520.1 contains similarity... 30 1.5
BF642076 homologue to GP|23505128|em hypothetical protein {Plasm... 29 1.9
TC78740 similar to GP|3482921|gb|AAC33206.1| Unknown protein {Ar... 29 1.9
TC83052 similar to GP|4063748|gb|AAC98456.1| unknown protein {Ar... 28 4.2
BQ140818 weakly similar to GP|6523547|emb| hydroxyproline-rich g... 28 4.2
CA920780 similar to GP|21689829|gb| unknown protein {Arabidopsis... 28 5.5
TC76694 homologue to GP|3549691|emb|CAA09228.1 thaumatin-like pr... 28 5.5
>TC87369 homologue to GP|14517500|gb|AAK62640.1 K16N12.18/K16N12.18
{Arabidopsis thaliana}, partial (42%)
Length = 1210
Score = 313 bits (801), Expect = 7e-86
Identities = 157/170 (92%), Positives = 163/170 (95%)
Frame = +2
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
++ +DAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIP+DTVNGIVDQLVKF
Sbjct: 455 VIQTDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPVDTVNGIVDQLVKF 634
Query: 186 GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT 245
GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAP GPAGKAGLQSTKRDSYGRLILGDIIT
Sbjct: 635 GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPVTGPAGKAGLQSTKRDSYGRLILGDIIT 814
Query: 246 SVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKPDES 295
SVNG KV +GSDLYRILDQCKVGDKV VEVLRGDHKEKIPVILEPK DE+
Sbjct: 815 SVNGNKVANGSDLYRILDQCKVGDKVIVEVLRGDHKEKIPVILEPKADET 964
>TC87368 homologue to GP|14517500|gb|AAK62640.1 K16N12.18/K16N12.18
{Arabidopsis thaliana}, partial (30%)
Length = 700
Score = 102 bits (254), Expect = 2e-22
Identities = 71/146 (48%), Positives = 81/146 (54%), Gaps = 33/146 (22%)
Frame = +2
Query: 8 HKQRLHYHSH------YSPHFTPNSLSLLRSSTPKTCFNSILILCTSIALSFT----NAD 57
H H SH + P T LSL + + PKTCF+S+LILCTS+ALS T N D
Sbjct: 119 HHTPTHLLSHPTLFLLHPPSSTKPPLSLPKLTIPKTCFDSVLILCTSLALSLTLFISNVD 298
Query: 58 SAYAFVVTPPRKLQSDELAT----------VVYITNLAVKMRL-------------RWMC 94
SA AFVVT PRKLQ+DELAT VVYITNLAVK
Sbjct: 299 SASAFVVTAPRKLQTDELATVRLFQENTPSVVYITNLAVKQDAFTLDVLEVPQGSGSGFV 478
Query: 95 WRFLKEGNIVTNYHVIPGASDLSLDL 120
W K+G+IVTNYHVI GASDL + L
Sbjct: 479 WD--KDGHIVTNYHVIRGASDLRVTL 550
>BQ140222 homologue to GP|14517500|gb K16N12.18/K16N12.18 {Arabidopsis
thaliana}, partial (23%)
Length = 307
Score = 77.0 bits (188), Expect(2) = 4e-15
Identities = 36/41 (87%), Positives = 39/41 (94%)
Frame = +1
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGV 166
++ +DAAINPGNSGGPLLDSSGNLIGINT IYSPSGASSGV
Sbjct: 184 VIQTDAAINPGNSGGPLLDSSGNLIGINTXIYSPSGASSGV 306
Score = 21.2 bits (43), Expect(2) = 4e-15
Identities = 10/11 (90%), Positives = 10/11 (90%)
Frame = +2
Query: 118 LDLTTLLQLVS 128
LDLTTLLQL S
Sbjct: 95 LDLTTLLQLES 127
>TC80319 similar to GP|20259261|gb|AAM14366.1 putative HhoA protease
precursor {Arabidopsis thaliana}, partial (47%)
Length = 1195
Score = 69.7 bits (169), Expect = 1e-12
Identities = 33/64 (51%), Positives = 48/64 (74%), Gaps = 2/64 (3%)
Frame = +2
Query: 125 QLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYS--PSGASSGVGFSIPLDTVNGIVDQL 182
+ + +DAAIN GNSGGPL+DS G+++G+NTA ++ +GASSGV F+IP+D V V L
Sbjct: 821 EAIQTDAAINAGNSGGPLIDSHGHVVGVNTATFTRKGTGASSGVNFAIPIDAVLRSVPYL 1000
Query: 183 VKFG 186
+ +G
Sbjct: 1001IVYG 1012
>BF642570 similar to GP|13172275|gb DegP2 protease {Arabidopsis thaliana},
partial (29%)
Length = 607
Score = 48.5 bits (114), Expect = 3e-06
Identities = 24/62 (38%), Positives = 34/62 (54%)
Frame = +2
Query: 127 VSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFG 186
+ DAAINPGNSGGP + G IG+ +Y A + +G+ IP V+ + K G
Sbjct: 356 IQIDAAINPGNSGGPAFNDQGECIGVAFXVYRSEDAEN-IGYVIPSTVVSHFLTDYXKNG 532
Query: 187 KV 188
K+
Sbjct: 533 KL 538
>AW774502 similar to GP|17428071|emb PROBABLE PERIPLASMIC PROTEASE SIGNAL
PEPTIDE PROTEIN {Ralstonia solanacearum}, partial (2%)
Length = 641
Score = 37.4 bits (85), Expect = 0.007
Identities = 41/160 (25%), Positives = 70/160 (43%), Gaps = 4/160 (2%)
Frame = +3
Query: 139 GGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFGKVTRPILGIK-- 196
GG L++ SG +IGI Y G ++ +P++ + + ++G + +P LG+
Sbjct: 96 GGALINLSGEVIGI--IFYYEFGFTTPF---LPINIAHKCWEHYKRYGGLRQPSLGVVAT 260
Query: 197 --FAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIITSVNGTKVTS 254
+A D + + + L L G + ++ + DS G L L D+I G V S
Sbjct: 261 SFYAADVFLMEKVIQKFLNL---CGGVLVEKVIKGSCADSAG-LHLNDVIVQCCGETVHS 428
Query: 255 GSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKPDE 294
I+ KVGD + + V R + PV + DE
Sbjct: 429 FLQFLEIVWDKKVGDVLQLSVARASQSQNDPVHVNMVVDE 548
>TC84647 similar to GP|9294054|dbj|BAB02011.1 contains similarity to signal
peptidase (leader peptidase)~gene_id:MOB24.17
{Arabidopsis thaliana}, partial (42%)
Length = 665
Score = 31.6 bits (70), Expect = 0.38
Identities = 28/89 (31%), Positives = 40/89 (44%), Gaps = 10/89 (11%)
Frame = +3
Query: 13 HYHSHYSPHFTPNSLSLLRSSTPKTCFNSILI-----LCTSIALSFTNADS-----AYAF 62
H+HS S HF P + ++L SS NSILI L ++ L F + A F
Sbjct: 24 HHHSLSSLHFKPQTPTMLISSIQSNSLNSILIQEDSLLHHALILPFPISTKPLQLFAVEF 203
Query: 63 VVTPPRKLQSDELATVVYITNLAVKMRLR 91
V K ++ ATVV +T ++ R
Sbjct: 204 HVVKL*KTPAEVAATVVVVTGRWIRKMSR 290
>TC92658 similar to GP|19310429|gb|AAL84951.1 AT3g03380/T21P5_20
{Arabidopsis thaliana}, partial (18%)
Length = 733
Score = 30.4 bits (67), Expect = 0.85
Identities = 25/100 (25%), Positives = 46/100 (46%), Gaps = 3/100 (3%)
Frame = +3
Query: 193 LGIKFAPDQSVEQLGVS---GVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIITSVNG 249
LGI+ +Q V + G+LV+++ P G A Y L GD++ VNG
Sbjct: 87 LGIRSETEQIVRNASPATETGMLVVESAV--PGGPA---------YEHLEPGDVLVRVNG 233
Query: 250 TKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILE 289
+T L +LD V + +++ RG + + ++++
Sbjct: 234 EVITQFLKLETLLDD-SVNSNIELQIERGGTSKSVTLLVQ 350
>TC78736 similar to GP|21553515|gb|AAM62608.1 unknown {Arabidopsis
thaliana}, partial (93%)
Length = 944
Score = 30.4 bits (67), Expect = 0.85
Identities = 16/54 (29%), Positives = 25/54 (45%)
Frame = +1
Query: 11 RLHYHSHYSPHFTPNSLSLLRSSTPKTCFNSILILCTSIALSFTNADSAYAFVV 64
+L + S S H PN + S P+TCF +L ++A +A FV+
Sbjct: 91 KLRFFSLRSSHMDPNQSIVENYSNPRTCFFHVLFKAAALAFYILSALFVDNFVI 252
>TC87772 weakly similar to GP|8574455|gb|AAF77578.1| pepper esterase {Capsicum
annuum}, partial (34%)
Length = 1313
Score = 30.0 bits (66), Expect = 1.1
Identities = 13/36 (36%), Positives = 19/36 (52%), Gaps = 5/36 (13%)
Frame = -2
Query: 6 KHHKQRLHYHSHYSP-----HFTPNSLSLLRSSTPK 36
KHH+Q+ H + HY HF P + S + S P+
Sbjct: 1078 KHHRQKYHVYHHYKTVQPH*HFYPQATSPIPSDDPQ 971
>TC79616 similar to GP|21553745|gb|AAM62838.1 Yippee-like protein
{Arabidopsis thaliana}, complete
Length = 641
Score = 30.0 bits (66), Expect = 1.1
Identities = 17/43 (39%), Positives = 23/43 (52%)
Frame = -3
Query: 6 KHHKQRLHYHSHYSPHFTPNSLSLLRSSTPKTCFNSILILCTS 48
KHH LH+ H+ H +PN+ + TP T I +LCTS
Sbjct: 441 KHH---LHHFQHHVLHGSPNNCR-HQDQTP*TVREQIFLLCTS 325
>TC80291 similar to GP|19310429|gb|AAL84951.1 AT3g03380/T21P5_20
{Arabidopsis thaliana}, partial (25%)
Length = 1009
Score = 30.0 bits (66), Expect = 1.1
Identities = 32/136 (23%), Positives = 52/136 (37%), Gaps = 18/136 (13%)
Frame = +3
Query: 136 GNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTV-----------NGIVDQLVK 184
G+SG P++D G +G+N S +S F +PL+ V VD+ +
Sbjct: 507 GSSGSPVIDWQGRAVGLNAG----SKTTSASAFFLPLERVVRALRFLQTGSETYVDKWMP 674
Query: 185 FGKVTRPILGIKFAPD--QSVEQLGVSGVLVLDAPANGPAGKAGLQSTKR-----DSYGR 237
+ R L + F +LG+ PA + G+ + Y
Sbjct: 675 -ASIPRGTLQLTFLHKGFDETRRLGLRSETEQVVRNASPASETGMLVVESVVPGGXXYKH 851
Query: 238 LILGDIITSVNGTKVT 253
L GD++ VNG +T
Sbjct: 852 LKPGDVLVRVNGEVIT 899
>TC84909 similar to GP|3395431|gb|AAC28763.1| unknown protein {Arabidopsis
thaliana}, partial (17%)
Length = 737
Score = 29.6 bits (65), Expect = 1.5
Identities = 16/34 (47%), Positives = 21/34 (61%)
Frame = -3
Query: 9 KQRLHYHSHYSPHFTPNSLSLLRSSTPKTCFNSI 42
K +++ S +SP F PN LS L S+T FNSI
Sbjct: 354 KMKMNQPSSFSPFFNPNPLSQLSSNT----FNSI 265
>TC86535 similar to GP|9758076|dbj|BAB08520.1 contains similarity to RNA
binding protein~gene_id:MNF13.1 {Arabidopsis thaliana},
partial (48%)
Length = 1555
Score = 29.6 bits (65), Expect = 1.5
Identities = 10/21 (47%), Positives = 16/21 (75%)
Frame = -1
Query: 1 MIQHYKHHKQRLHYHSHYSPH 21
++QH HH Q+ H+HSH++ H
Sbjct: 988 LLQHI-HHTQQTHFHSHHTDH 929
>BF642076 homologue to GP|23505128|em hypothetical protein {Plasmodium
falciparum 3D7}, partial (0%)
Length = 503
Score = 29.3 bits (64), Expect = 1.9
Identities = 13/30 (43%), Positives = 16/30 (53%)
Frame = -3
Query: 3 QHYKHHKQRLHYHSHYSPHFTPNSLSLLRS 32
QH HH Q H + H +PN L+LL S
Sbjct: 246 QHLHHHHQDHLQHHSFQLHLSPNPLTLLPS 157
>TC78740 similar to GP|3482921|gb|AAC33206.1| Unknown protein {Arabidopsis
thaliana}, partial (45%)
Length = 1624
Score = 29.3 bits (64), Expect = 1.9
Identities = 43/173 (24%), Positives = 67/173 (37%), Gaps = 4/173 (2%)
Frame = +2
Query: 4 HYKHHKQRLHYHSHYSPHFTPNSLSLLRSSTPKTC--FNSILILCTSIALSFTNADSAYA 61
H+ HHK H HS+ S + N++S + ++ TC F S LI S +F+ A
Sbjct: 44 HHLHHKS--H*HSNSSILNSRNTISFIMANEASTCLLFLSFLIFFCSGTNAFSTFSGARF 217
Query: 62 FVVTPPRKLQSDELATVVYITNLAVKMRLRWMCWRFLKEGNIVTNYHVIPGASDLSLDLT 121
+ K Q L T I + T PGA+ +++ T
Sbjct: 218 LQKSKISKSQIQFLVTNHKIGRKLQDTNPAPTIITVPSTNPVTTVSPTNPGATPVTVPST 397
Query: 122 TLLQLVSSDAAINPGNSGGPL--LDSSGNLIGINTAIYSPSGASSGVGFSIPL 172
T + S NP NS P+ + G +N+ Y P SG ++P+
Sbjct: 398 TPPSVPLSPT--NPANSPVPVTPITVPGGTTPVNS--YPPPSPLSGGTGTVPV 544
>TC83052 similar to GP|4063748|gb|AAC98456.1| unknown protein {Arabidopsis
thaliana}, partial (86%)
Length = 877
Score = 28.1 bits (61), Expect = 4.2
Identities = 12/26 (46%), Positives = 20/26 (76%), Gaps = 1/26 (3%)
Frame = -2
Query: 2 IQHYKHHKQRLHYHSHYSPHFT-PNS 26
+Q + H+Q+ H+HSH+S H+T PN+
Sbjct: 492 LQQQQQHQQQ-HHHSHFSNHWTEPNN 418
>BQ140818 weakly similar to GP|6523547|emb| hydroxyproline-rich
glycoprotein DZ-HRGP {Volvox carteri f. nagariensis},
partial (10%)
Length = 359
Score = 28.1 bits (61), Expect = 4.2
Identities = 8/17 (47%), Positives = 12/17 (70%)
Frame = +1
Query: 4 HYKHHKQRLHYHSHYSP 20
H+ HH+ R HYH ++ P
Sbjct: 46 HHHHHRSRNHYHRYHHP 96
>CA920780 similar to GP|21689829|gb| unknown protein {Arabidopsis thaliana},
partial (27%)
Length = 780
Score = 27.7 bits (60), Expect = 5.5
Identities = 12/28 (42%), Positives = 17/28 (59%)
Frame = +3
Query: 2 IQHYKHHKQRLHYHSHYSPHFTPNSLSL 29
+Q + HH +L H HYSP P++L L
Sbjct: 516 VQLFHHHSLQLRRHRHYSP---PDTLQL 590
>TC76694 homologue to GP|3549691|emb|CAA09228.1 thaumatin-like protein PR-5b
{Cicer arietinum}, complete
Length = 947
Score = 27.7 bits (60), Expect = 5.5
Identities = 17/56 (30%), Positives = 26/56 (46%), Gaps = 3/56 (5%)
Frame = +2
Query: 15 HSHYSPHFTPNSL---SLLRSSTPKTCFNSILILCTSIALSFTNADSAYAFVVTPP 67
HSHY+P+++P+ L LRSST L+ +A S ++ PP
Sbjct: 56 HSHYAPYYSPSLLHKQPTLRSSTIALTPFGQLLAQVVVAASTAVKHGTSGLILVPP 223
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.136 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,303,628
Number of Sequences: 36976
Number of extensions: 135181
Number of successful extensions: 1536
Number of sequences better than 10.0: 54
Number of HSP's better than 10.0 without gapping: 1413
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1495
length of query: 295
length of database: 9,014,727
effective HSP length: 95
effective length of query: 200
effective length of database: 5,502,007
effective search space: 1100401400
effective search space used: 1100401400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0225.5


