

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0218.20
(631 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
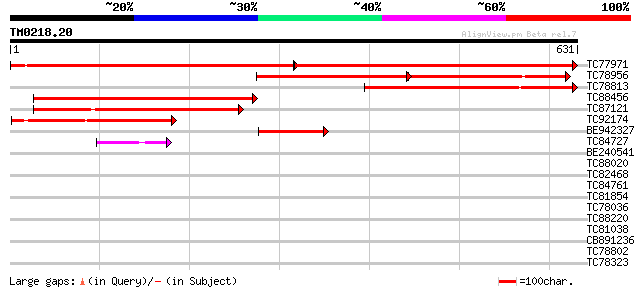
Score E
Sequences producing significant alignments: (bits) Value
TC77971 similar to PIR|T06794|T06794 vacuolar sorting receptor p... 613 0.0
TC78956 similar to PIR|G86434|G86434 protein F17F8.23 [imported]... 267 e-139
TC78813 homologue to PIR|T06794|T06794 vacuolar sorting receptor... 462 e-130
TC88456 similar to GP|15487292|dbj|BAB64531. vacuolar sorting re... 385 e-107
TC87121 weakly similar to PIR|T06794|T06794 vacuolar sorting rec... 303 1e-82
TC92174 homologue to PIR|T06794|T06794 vacuolar sorting receptor... 280 2e-75
BE942327 similar to GP|15487292|dbj vacuolar sorting receptor {V... 119 3e-27
TC84727 similar to GP|16118856|gb|AAL14629.1 growth-on protein G... 47 2e-05
BE240541 homologue to GP|15487292|dbj vacuolar sorting receptor ... 39 0.008
TC88020 similar to GP|21537054|gb|AAM61395.1 putative P-protein:... 36 0.039
TC82468 similar to GP|8920639|gb|AAF81361.1| Identical to wall-a... 32 0.57
TC84761 32 0.74
TC81854 weakly similar to PIR|T50012|T50012 hypothetical protein... 31 1.3
TC78036 homologue to PIR|T06824|T06824 translation initiation fa... 30 2.8
TC88220 similar to GP|15293201|gb|AAK93711.1 unknown protein {Ar... 30 2.8
TC81038 weakly similar to GP|15864565|emb|CAC80703. SIAH1 protei... 30 3.7
CB891236 weakly similar to PIR|T06757|T06 hypothetical protein F... 28 8.2
TC78802 similar to GP|19424009|gb|AAL87286.1 unknown protein {Ar... 28 8.2
TC78323 similar to GP|17529172|gb|AAL38812.1 unknown protein {Ar... 28 8.2
>TC77971 similar to PIR|T06794|T06794 vacuolar sorting receptor protein BP-80
- garden pea, partial (97%)
Length = 2578
Score = 613 bits (1580), Expect(2) = 0.0
Identities = 286/315 (90%), Positives = 299/315 (94%), Gaps = 1/315 (0%)
Frame = +3
Query: 318 KEKKYNKKCADAVIESLGLDNKKIERCMGDPDADSDNPVLKEEQDAQVGKGSRGDVTILP 377
+ K KCADAVIESLGLD KKIERCMGDPDADS+N VLKEEQDAQVGKGSRGDVTILP
Sbjct: 1293 RRKNTIMKCADAVIESLGLDIKKIERCMGDPDADSENSVLKEEQDAQVGKGSRGDVTILP 1472
Query: 378 TLVVNNRQYRGKLEKGAVMKAICSGFEETTEPAVCLSSDVETNECLENNGGCWKDKAANI 437
TLVVN+RQYRGKLEKGAVMKAICSGFEETTEPAVCLSS+VETNECLENNGGCWKDKAANI
Sbjct: 1473 TLVVNSRQYRGKLEKGAVMKAICSGFEETTEPAVCLSSEVETNECLENNGGCWKDKAANI 1652
Query: 438 TACKDTFRGRVCECPLVDGVQFKGDGYTTCEASGPGRCKIKNGGCWHEARNGHAFSACVD 497
TACKDTFRGRVCECPLVDGVQFKGDGYTTCEASGPGRCKI NGGCWHEARNGHAFSAC D
Sbjct: 1653 TACKDTFRGRVCECPLVDGVQFKGDGYTTCEASGPGRCKINNGGCWHEARNGHAFSACSD 1832
Query: 498 DGEVKCQCPAGFKGDGVKNCQDVDECKEKKACQCPECSCKNTWGSYDCSCSGDLLYIRDH 557
+G VKC+CPAGFKGDGVKNC D+DECKEKKACQCPECSCKNTWGSY+CSCSGDLLYIRDH
Sbjct: 1833 NGAVKCECPAGFKGDGVKNCADIDECKEKKACQCPECSCKNTWGSYNCSCSGDLLYIRDH 2012
Query: 558 DTCISKTASQ-GGRSAWAAFWVIIAGLVLSAGGGYLIYKYRIRSYMDSEIRAIMAQYMPL 616
D CISKTASQ GG+SAWAAFWVI+ GLVL+A G YL+YKYRIRSYMDSEIRAIMAQYMPL
Sbjct: 2013 DACISKTASQEGGKSAWAAFWVIVVGLVLAASGAYLVYKYRIRSYMDSEIRAIMAQYMPL 2192
Query: 617 DSQAEVVNHVNDERA 631
DSQ+EVVNHVNDERA
Sbjct: 2193 DSQSEVVNHVNDERA 2237
Score = 486 bits (1250), Expect(2) = 0.0
Identities = 248/324 (76%), Positives = 273/324 (83%), Gaps = 3/324 (0%)
Frame = +1
Query: 1 MKLRRFS-LGFLLGFLLLLSVTPSSMARFVVEKNSLTVTSPEDIKGTHDSAIGNFGIPQY 59
MK+R S +G LLGFLLL S+TPSSMA+FVVEKNSL VTSP+ IKGT+DSAIGNFGIPQY
Sbjct: 337 MKIRSSSSMGLLLGFLLL-SLTPSSMAKFVVEKNSLRVTSPDSIKGTYDSAIGNFGIPQY 513
Query: 60 GGSMAGNVVYPKDNHKGCKEFDESGISFKSKPGALPTIVLLDRGNCFFALKVWNAQKAGA 119
GGSMAGNVVYPKDN KGCKEFDESGISFKSKPGALPTIVLLDRG+CFFALKVWNAQKAGA
Sbjct: 514 GGSMAGNVVYPKDNQKGCKEFDESGISFKSKPGALPTIVLLDRGSCFFALKVWNAQKAGA 693
Query: 120 SAVLVADDIEEKLITMDTPEEDGSSAKYIENITIPSALIEKSFGEKLKKSISGGEMVNVN 179
S+VLVADDIEEKLITMDTPEEDGSSAKYIENITIPSALIEK+FGEKLKK+ISGG+MVNVN
Sbjct: 694 SSVLVADDIEEKLITMDTPEEDGSSAKYIENITIPSALIEKNFGEKLKKAISGGDMVNVN 873
Query: 180 LDWREAVPHPDDRVEYELWTNSNDECGVKCDML--MEFVKDFKGAAQILEKGGYTQFTPH 237
LDWREAVPHPDDRVEYEL + V+ + + + + + K P
Sbjct: 874 LDWREAVPHPDDRVEYEL-SGPTAMMSVELNAIC*WNL*RILRVRHKY*RKVVMPSLHPT 1050
Query: 238 YITWYCPKAFTLSKQCKSQCINHGRYCAPDPEQDFSTGYDGKDVVIENLRQLCVYKVTSE 297
+ P+AFTLSKQCKSQCINHGRYCAPDPEQDFSTGYDGKDVVIENLRQLCV+KV +E
Sbjct: 1051ILLGTVPQAFTLSKQCKSQCINHGRYCAPDPEQDFSTGYDGKDVVIENLRQLCVFKVANE 1230
Query: 298 AKKPWLWWDYVTDFQIRCPMKEKK 321
KKPW+WWDYVTDFQI P + +K
Sbjct: 1231TKKPWVWWDYVTDFQIMVPNEGEK 1302
Score = 28.5 bits (62), Expect = 8.2
Identities = 12/33 (36%), Positives = 19/33 (57%)
Frame = +2
Query: 293 KVTSEAKKPWLWWDYVTDFQIRCPMKEKKYNKK 325
++ + K P + F++ CPMKEKKYN +
Sbjct: 1217 RLQMKPKSPGCGGTMLLIFKLWCPMKEKKYNNE 1315
>TC78956 similar to PIR|G86434|G86434 protein F17F8.23 [imported] -
Arabidopsis thaliana, partial (47%)
Length = 1744
Score = 267 bits (683), Expect(2) = e-139
Identities = 118/173 (68%), Positives = 147/173 (84%)
Frame = +1
Query: 275 GYDGKDVVIENLRQLCVYKVTSEAKKPWLWWDYVTDFQIRCPMKEKKYNKKCADAVIESL 334
GYDGKD+V ENLRQLCV+ V +E + W+WWDYVTDF +RC MKEKKY+K CA+ V++SL
Sbjct: 4 GYDGKDIVYENLRQLCVHLVANETGRSWVWWDYVTDFHVRCSMKEKKYSKDCAEVVMKSL 183
Query: 335 GLDNKKIERCMGDPDADSDNPVLKEEQDAQVGKGSRGDVTILPTLVVNNRQYRGKLEKGA 394
L KI++C+GDP+AD +N VLK EQ +Q+G GSRGDVTILPTLV+NN QYRGKLE+ +
Sbjct: 184 DLPLDKIKKCVGDPEADVENEVLKIEQTSQIGGGSRGDVTILPTLVLNNVQYRGKLERTS 363
Query: 395 VMKAICSGFEETTEPAVCLSSDVETNECLENNGGCWKDKAANITACKDTFRGR 447
V+KAIC+GF+ETTEP VCLS D+ETNECLE NGGCW+D+ N+TACKDTF+G+
Sbjct: 364 VLKAICAGFKETTEPQVCLSGDIETNECLERNGGCWEDRNVNVTACKDTFQGK 522
Score = 246 bits (628), Expect(2) = e-139
Identities = 109/181 (60%), Positives = 133/181 (73%)
Frame = +2
Query: 444 FRGRVCECPLVDGVQFKGDGYTTCEASGPGRCKIKNGGCWHEARNGHAFSACVDDGEVKC 503
FRGRVCECP+V GVQ++GDGYT+CEA GP RC + NGGCW E R+ FSAC + C
Sbjct: 512 FRGRVCECPVVKGVQYRGDGYTSCEAFGPARCSLNNGGCWSETRDKITFSACSESKVNGC 691
Query: 504 QCPAGFKGDGVKNCQDVDECKEKKACQCPECSCKNTWGSYDCSCSGDLLYIRDHDTCISK 563
CP GF GDG C+DVDECKEK ACQC +CSCKNTWGSYDC C G LLY+++ D CI +
Sbjct: 692 HCPVGFDGDGNNKCEDVDECKEKSACQCDDCSCKNTWGSYDCKCKGGLLYMKEQDMCIER 871
Query: 564 TASQGGRSAWAAFWVIIAGLVLSAGGGYLIYKYRIRSYMDSEIRAIMAQYMPLDSQAEVV 623
+ S+ G+ +V+IA +V + GY+ YKYR+RSYMDSEI AIM+QYMPLD Q VV
Sbjct: 872 SGSKFGK---VVAFVVIAVVVGAGLAGYVFYKYRLRSYMDSEIMAIMSQYMPLDQQNNVV 1042
Query: 624 N 624
+
Sbjct: 1043H 1045
>TC78813 homologue to PIR|T06794|T06794 vacuolar sorting receptor protein
BP-80 - garden pea, partial (37%)
Length = 1043
Score = 462 bits (1190), Expect = e-130
Identities = 206/236 (87%), Positives = 221/236 (93%)
Frame = +2
Query: 396 MKAICSGFEETTEPAVCLSSDVETNECLENNGGCWKDKAANITACKDTFRGRVCECPLVD 455
+KAICSGFEETT+PAVCLS+DVETNECL NNGGCW+DKAANITACKDTFRGRVCECPLVD
Sbjct: 2 LKAICSGFEETTDPAVCLSNDVETNECLTNNGGCWQDKAANITACKDTFRGRVCECPLVD 181
Query: 456 GVQFKGDGYTTCEASGPGRCKIKNGGCWHEARNGHAFSACVDDGEVKCQCPAGFKGDGVK 515
GVQFKGDGYTTCE GPGRCKI NGGCWH+ARNGHAFSAC+DDG VKCQCP GFKGDGVK
Sbjct: 182 GVQFKGDGYTTCEVGGPGRCKINNGGCWHDARNGHAFSACLDDGGVKCQCPTGFKGDGVK 361
Query: 516 NCQDVDECKEKKACQCPECSCKNTWGSYDCSCSGDLLYIRDHDTCISKTASQGGRSAWAA 575
NC+D+DECKEKKACQCPECSCKNTWGSYDCSCSGDLLYI+DHDTCISKTASQ +S WAA
Sbjct: 362 NCEDIDECKEKKACQCPECSCKNTWGSYDCSCSGDLLYIKDHDTCISKTASQ-AKSTWAA 538
Query: 576 FWVIIAGLVLSAGGGYLIYKYRIRSYMDSEIRAIMAQYMPLDSQAEVVNHVNDERA 631
FWVI+ GL + AGGG+L+YKYRIR YMDSEIRAIMAQYMPLDSQAEV NHVN +RA
Sbjct: 539 FWVILIGLGMIAGGGFLVYKYRIRQYMDSEIRAIMAQYMPLDSQAEVPNHVNHQRA 706
>TC88456 similar to GP|15487292|dbj|BAB64531. vacuolar sorting receptor
{Vigna mungo}, partial (41%)
Length = 980
Score = 385 bits (988), Expect = e-107
Identities = 175/250 (70%), Positives = 207/250 (82%), Gaps = 1/250 (0%)
Frame = +2
Query: 27 RFVVEKNSLTVTSPEDIKGTHDSAIGNFGIPQYGGSMAGNVVYPKDNHKGCKEFDESGIS 86
RF+VEKNSL +TSP+ +KG+++ AIGNFG+PQYGG++ G+VVYP N KGCK F + S
Sbjct: 230 RFLVEKNSLRITSPKSLKGSYECAIGNFGVPQYGGTLVGSVVYPNVNQKGCKNFTDFSAS 409
Query: 87 FKSKPGALPTIVLLDRGNCFFALKVWNAQKAGASAVLVADDIEEKLITMDTPEEDGS-SA 145
F S PG PT VL+DRG+C+F LK WNAQ GA+A+LVADD EE LITMDTPEE +
Sbjct: 410 FHSMPGNFPTFVLVDRGDCYFTLKAWNAQNGGAAAILVADDREETLITMDTPEEGNVVND 589
Query: 146 KYIENITIPSALIEKSFGEKLKKSISGGEMVNVNLDWREAVPHPDDRVEYELWTNSNDEC 205
YIE I IPSALI KS G+++KK++S GEMV++NLDWREA+PHPDDRVEYELWTNSNDEC
Sbjct: 590 DYIEKINIPSALISKSLGDRIKKALSDGEMVHINLDWREALPHPDDRVEYELWTNSNDEC 769
Query: 206 GVKCDMLMEFVKDFKGAAQILEKGGYTQFTPHYITWYCPKAFTLSKQCKSQCINHGRYCA 265
G KCD + FVK FKGAAQ+LEK G+TQFTPHYITWYCPK F LS++CKSQCINHGRYCA
Sbjct: 770 GPKCDNQINFVKSFKGAAQLLEKKGFTQFTPHYITWYCPKEFLLSRRCKSQCINHGRYCA 949
Query: 266 PDPEQDFSTG 275
PDPEQDF+ G
Sbjct: 950 PDPEQDFNKG 979
>TC87121 weakly similar to PIR|T06794|T06794 vacuolar sorting receptor
protein BP-80 - garden pea, partial (30%)
Length = 810
Score = 303 bits (777), Expect = 1e-82
Identities = 145/235 (61%), Positives = 178/235 (75%), Gaps = 1/235 (0%)
Frame = +2
Query: 27 RFVVEKNSLTVTSPEDIKGTHDSAIGNFGIPQYGGSMAGNVVYPKDNHKGCKEFD-ESGI 85
RFVVEKNS+TV SP ++G +D AIGNFGIP YGG + G++VYP+ GC+ F+ +
Sbjct: 110 RFVVEKNSITVLSPHKLRGKNDGAIGNFGIPNYGGYIVGSLVYPEKGSHGCQVFEGDKPF 289
Query: 86 SFKSKPGALPTIVLLDRGNCFFALKVWNAQKAGASAVLVADDIEEKLITMDTPEEDGSSA 145
F+S PTIVLLDRG C+FALKVW+AQ AGA+AVLVAD I+E LITMD+PEE +
Sbjct: 290 KFQSHR---PTIVLLDRGECYFALKVWHAQLAGAAAVLVADSIDESLITMDSPEESTDAE 460
Query: 146 KYIENITIPSALIEKSFGEKLKKSISGGEMVNVNLDWREAVPHPDDRVEYELWTNSNDEC 205
YIE I IPS L+EKSFG+ LK++++ + V + +DWRE+VPHPD+RVEYE WTNSNDEC
Sbjct: 461 GYIEKIVIPSVLVEKSFGDSLKEALNNKDEVLLRIDWRESVPHPDNRVEYEFWTNSNDEC 640
Query: 206 GVKCDMLMEFVKDFKGAAQILEKGGYTQFTPHYITWYCPKAFTLSKQCKSQCINH 260
G +CD M FVK+FKG AQILE+ GYT FTPHYITW KA + QCKS CINH
Sbjct: 641 GARCDEQMNFVKNFKGHAQILERSGYTLFTPHYITWVLSKAIC*TSQCKSXCINH 805
>TC92174 homologue to PIR|T06794|T06794 vacuolar sorting receptor protein
BP-80 - garden pea, partial (28%)
Length = 669
Score = 280 bits (715), Expect = 2e-75
Identities = 145/183 (79%), Positives = 158/183 (86%)
Frame = +3
Query: 3 LRRFSLGFLLGFLLLLSVTPSSMARFVVEKNSLTVTSPEDIKGTHDSAIGNFGIPQYGGS 62
L R S+ L+GF+L T S ARFVVEKNSL+VTSP+ IKG HDSAIGNFGIPQYGGS
Sbjct: 135 LWRLSVFMLVGFML----TGMSTARFVVEKNSLSVTSPDKIKGKHDSAIGNFGIPQYGGS 302
Query: 63 MAGNVVYPKDNHKGCKEFDESGISFKSKPGALPTIVLLDRGNCFFALKVWNAQKAGASAV 122
MAGNVVYPKDN+KGCK+FD+S SFKSKPGALPTI+LLDRG+CFFALKVWNA KAGASAV
Sbjct: 303 MAGNVVYPKDNNKGCKDFDDSS-SFKSKPGALPTILLLDRGSCFFALKVWNAXKAGASAV 479
Query: 123 LVADDIEEKLITMDTPEEDGSSAKYIENITIPSALIEKSFGEKLKKSISGGEMVNVNLDW 182
LVADDIEE LITMDTPEED SSAKYIENITIPSALI K+FG L +I GG+M NVNLDW
Sbjct: 480 LVADDIEEPLITMDTPEEDVSSAKYIENITIPSALIGKTFGXXLXDAIXGGDMXNVNLDW 659
Query: 183 REA 185
REA
Sbjct: 660 REA 668
>BE942327 similar to GP|15487292|dbj vacuolar sorting receptor {Vigna mungo},
partial (12%)
Length = 235
Score = 119 bits (299), Expect = 3e-27
Identities = 50/78 (64%), Positives = 64/78 (81%)
Frame = +2
Query: 278 GKDVVIENLRQLCVYKVTSEAKKPWLWWDYVTDFQIRCPMKEKKYNKKCADAVIESLGLD 337
GKDV ++N RQ C +KV +E+ +PW WWDYVTDF IRCPMKEKKY ++C+D VI+SLG+D
Sbjct: 2 GKDVEVQNSRQACFFKVANESGRPWQWWDYVTDFSIRCPMKEKKYTQECSDEVIKSLGVD 181
Query: 338 NKKIERCMGDPDADSDNP 355
KKI+ C+GDP AD +NP
Sbjct: 182 LKKIKDCVGDPIADVENP 235
>TC84727 similar to GP|16118856|gb|AAL14629.1 growth-on protein GRO10
{Euphorbia esula}, partial (47%)
Length = 936
Score = 47.0 bits (110), Expect = 2e-05
Identities = 27/84 (32%), Positives = 45/84 (53%)
Frame = +2
Query: 97 IVLLDRGNCFFALKVWNAQKAGASAVLVADDIEEKLITMDTPEEDGSSAKYIENITIPSA 156
++++DRGNC F K +AQ A ASA+L+ ++ +E + P+E S I IP+
Sbjct: 377 VIMVDRGNCTFTKKANSAQNANASAILIINNQKELYKMVCDPDETDLS------IHIPAG 538
Query: 157 LIEKSFGEKLKKSISGGEMVNVNL 180
++ G KL+ + V+V L
Sbjct: 539 MLPLDAGTKLENMLKSTSSVSVQL 610
>BE240541 homologue to GP|15487292|dbj vacuolar sorting receptor {Vigna
mungo}, partial (4%)
Length = 294
Score = 38.5 bits (88), Expect = 0.008
Identities = 17/26 (65%), Positives = 22/26 (84%)
Frame = +2
Query: 602 MDSEIRAIMAQYMPLDSQAEVVNHVN 627
MD+EIRAIMAQYMPLD+Q ++ + N
Sbjct: 2 MDTEIRAIMAQYMPLDNQPLIIPNPN 79
>TC88020 similar to GP|21537054|gb|AAM61395.1 putative P-protein: chorismate
mutase prephenate dehydratase {Arabidopsis thaliana},
partial (77%)
Length = 1686
Score = 36.2 bits (82), Expect = 0.039
Identities = 29/105 (27%), Positives = 40/105 (37%), Gaps = 3/105 (2%)
Frame = +1
Query: 445 RGRVCECPLVDGVQFK-GDGYTTCEASGPGRCKIKN--GGCWHEARNGHAFSACVDDGEV 501
R ++C L DG + G+ C S G C + G W R+G +S C
Sbjct: 562 RSKICSTSLFDGQPWC*ASGFEACS*SSTGSCTM*EHLDGVWLSQRSG**YSRCC----- 726
Query: 502 KCQCPAGFKGDGVKNCQDVDECKEKKACQCPECSCKNTWGSYDCS 546
K CP Q C+ +C C C C++ W Y CS
Sbjct: 727 KACCP-----------QKTTRCR---SC-C*LCCCRDLWLKYTCS 816
>TC82468 similar to GP|8920639|gb|AAF81361.1| Identical to wall-associated
kinase 4 from Arabidopsis thaliana gb|AJ009695 and
contains Eukaryotic, partial (22%)
Length = 937
Score = 32.3 bits (72), Expect = 0.57
Identities = 13/29 (44%), Positives = 17/29 (57%)
Frame = +2
Query: 519 DVDECKEKKACQCPECSCKNTWGSYDCSC 547
D+DECK E C+NT GS++C C
Sbjct: 5 DIDECKTGNNTCISEKHCRNTDGSHECFC 91
>TC84761
Length = 620
Score = 32.0 bits (71), Expect = 0.74
Identities = 21/73 (28%), Positives = 32/73 (43%)
Frame = -1
Query: 245 KAFTLSKQCKSQCINHGRYCAPDPEQDFSTGYDGKDVVIENLRQLCVYKVTSEAKKPWLW 304
K F S KSQCINH C + F + + + NL L + + E+K
Sbjct: 191 KRFLKSTNSKSQCINHHSRCNESELKKFEENFQDRRI---NLINLKIETLKGESK----- 36
Query: 305 WDYVTDFQIRCPM 317
V +F+ +CP+
Sbjct: 35 ---VLEFRQKCPI 6
>TC81854 weakly similar to PIR|T50012|T50012 hypothetical protein T31P16.70
- Arabidopsis thaliana, partial (29%)
Length = 1124
Score = 31.2 bits (69), Expect = 1.3
Identities = 16/48 (33%), Positives = 24/48 (49%)
Frame = +1
Query: 506 PAGFKGDGVKNCQDVDECKEKKACQCPECSCKNTWGSYDCSCSGDLLY 553
P + D VK+ K ++AC C C CK + ++ C S D+LY
Sbjct: 664 PLPSEWDIVKSAACR*AAKNQRACSCSSCCCKGSSKTFSCFIS-DILY 804
>TC78036 homologue to PIR|T06824|T06824 translation initiation factor -
garden pea, complete
Length = 1571
Score = 30.0 bits (66), Expect = 2.8
Identities = 14/37 (37%), Positives = 19/37 (50%)
Frame = +2
Query: 525 EKKACQCPECSCKNTWGSYDCSCSGDLLYIRDHDTCI 561
+ AC CP+ S K C CSG + Y +D D C+
Sbjct: 260 DSTACGCPDYSGKG------CYCSGAIWYWKDLDDCL 352
>TC88220 similar to GP|15293201|gb|AAK93711.1 unknown protein {Arabidopsis
thaliana}, partial (26%)
Length = 1388
Score = 30.0 bits (66), Expect = 2.8
Identities = 16/40 (40%), Positives = 20/40 (50%)
Frame = +1
Query: 497 DDGEVKCQCPAGFKGDGVKNCQDVDECKEKKACQCPECSC 536
D EV C GDG +C+D +C+EK AC C C
Sbjct: 841 DGEEVSCYG----SGDGETSCRDC-KCREKSACYCCRHRC 945
>TC81038 weakly similar to GP|15864565|emb|CAC80703. SIAH1 protein {Brassica
napus}, partial (21%)
Length = 1016
Score = 29.6 bits (65), Expect = 3.7
Identities = 22/75 (29%), Positives = 34/75 (45%), Gaps = 4/75 (5%)
Frame = +2
Query: 484 HEARNGH-AFSACVDDGEVKCQCPAGFKGDGVKNCQDVDECKEKKACQCPEC--SCKNTW 540
++ NGH A S C GE++ +CP G C+ V++ E CP CK+ +
Sbjct: 329 YQCENGHIACSKCC--GELRNKCPMCSMPIGYNRCRAVEKLLESIKISCPNAKYGCKDMF 502
Query: 541 G-SYDCSCSGDLLYI 554
S S + +YI
Sbjct: 503 SCSMKSSHEKECIYI 547
>CB891236 weakly similar to PIR|T06757|T06 hypothetical protein F15B8.180 -
Arabidopsis thaliana, partial (29%)
Length = 709
Score = 28.5 bits (62), Expect = 8.2
Identities = 11/37 (29%), Positives = 17/37 (45%)
Frame = +2
Query: 520 VDECKEKKACQCPECSCKNTWGSYDCSCSGDLLYIRD 556
+++C C+ C C N W DCS + IR+
Sbjct: 344 INQCSRHGHCRGGFCQCANGWYGADCSTPSVISSIRE 454
>TC78802 similar to GP|19424009|gb|AAL87286.1 unknown protein {Arabidopsis
thaliana}, complete
Length = 1151
Score = 28.5 bits (62), Expect = 8.2
Identities = 11/29 (37%), Positives = 15/29 (50%)
Frame = +2
Query: 236 PHYITWYCPKAFTLSKQCKSQCINHGRYC 264
P+ I W C K +QC + C N R+C
Sbjct: 701 PNKI*WSCKKDNVWRRQCSNSCSNWTRHC 787
>TC78323 similar to GP|17529172|gb|AAL38812.1 unknown protein {Arabidopsis
thaliana}, partial (86%)
Length = 1006
Score = 28.5 bits (62), Expect = 8.2
Identities = 15/47 (31%), Positives = 20/47 (41%), Gaps = 2/47 (4%)
Frame = +1
Query: 509 FKGDGVKNCQDVDE--CKEKKACQCPECSCKNTWGSYDCSCSGDLLY 553
+K DG C + C + C P SCK G+ DCS L +
Sbjct: 517 WKRDGCSKCPKNSKAVCLNGQDCALPTSSCKTHSGTVDCSLGIQLAF 657
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.136 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,116,383
Number of Sequences: 36976
Number of extensions: 366904
Number of successful extensions: 1907
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 1854
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1892
length of query: 631
length of database: 9,014,727
effective HSP length: 102
effective length of query: 529
effective length of database: 5,243,175
effective search space: 2773639575
effective search space used: 2773639575
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0218.20


